
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 21,021,895 – 21,021,985 |
| Length | 90 |
| Max. P | 0.611303 |

| Location | 21,021,895 – 21,021,985 |
|---|---|
| Length | 90 |
| Sequences | 4 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 68.41 |
| Mean single sequence MFE | -22.60 |
| Consensus MFE | -12.81 |
| Energy contribution | -11.93 |
| Covariance contribution | -0.88 |
| Combinations/Pair | 1.33 |
| Mean z-score | -1.40 |
| Structure conservation index | 0.57 |
| SVM decision value | 0.16 |
| SVM RNA-class probability | 0.611303 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 21021895 90 - 22407834 AGUGCGGAUCCGCAUGAAUGAA------------GUGAAGUGGCAAC---------GAAC--AAUGACAU-GUGGGAAACUGCAGGCCGAAGGGCAUUA--AAAACAGAUCGAUUG ..(((((.((((((((.((...------------((....((....)---------).))--.))..)))-)))))...))))).(((....)))....--............... ( -24.40) >DroPse_CAF1 155336 114 - 1 AGUGAGGAUUCGCAUGAAUGAAAACAGAAGUGAAGUGAAGUGUUUACAAGCAGAACGAACCCACAGACACAGAGGGAGACA--GAGCCAUGGGGCAUCAUCAAAACGAAUCAAUUU ......((((((..(((.(((........(((..(((...(((((.......)))))....)))...)))....(....).--..(((....))).))))))...))))))..... ( -22.00) >DroYak_CAF1 78100 89 - 1 A-UGCGGAUCCGCAUGAAUGAA------------GUGAAGUGGUAAC---------GAAC--AAUAACAU-GUGGGAAACUGCAGGCCGAAGGGCAUUA--AGAACAGAUCGAUAG .-(((((.((((((((......------------((...((....))---------..))--.....)))-)))))...))))).(((....)))....--............... ( -23.40) >DroPer_CAF1 145978 114 - 1 AGUGAGGAUUCGCAUGAAUGAAAACAGAAGUGAAGUGAAGUGUUUACAAGCAGAACGAACCCACAGACACAGAGGGAGAAA--AAGCCGUGUGGCAUCAUCAAAACGAAUCAAUUU ......((((((..(((.(((........(((..(((...(((((.......)))))....)))...)))...........--..(((....))).))))))...))))))..... ( -20.60) >consensus AGUGAGGAUCCGCAUGAAUGAA____________GUGAAGUGGUAAC_________GAAC__AAAGACAC_GAGGGAAACA__AAGCCGAAGGGCAUCA__AAAACAAAUCAAUUG ......((((((......(((..................((((((...................))))))...............(((....))).)))......))))))..... (-12.81 = -11.93 + -0.88)

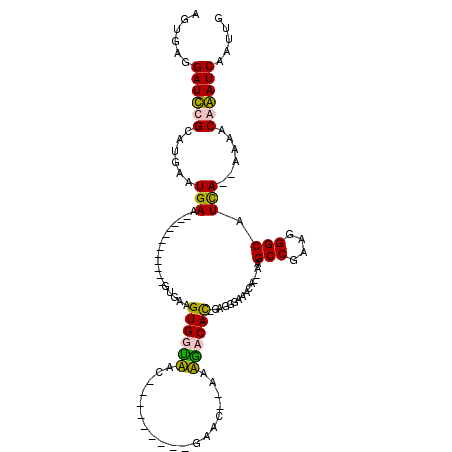
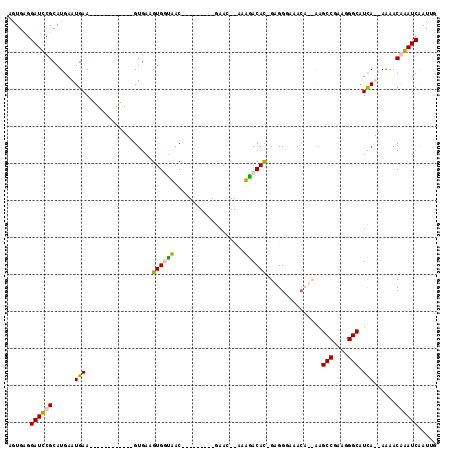
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:40:49 2006