
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 20,994,761 – 20,994,855 |
| Length | 94 |
| Max. P | 0.998329 |

| Location | 20,994,761 – 20,994,855 |
|---|---|
| Length | 94 |
| Sequences | 6 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 95.90 |
| Mean single sequence MFE | -25.87 |
| Consensus MFE | -21.84 |
| Energy contribution | -22.62 |
| Covariance contribution | 0.78 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.36 |
| Structure conservation index | 0.84 |
| SVM decision value | 1.26 |
| SVM RNA-class probability | 0.937036 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 20994761 94 + 22407834 CCACAAAAACCAGAGCUGGCAUAUUGCCGCUAUUGUCUUUUAUAGUUUCCUGGAACAUUUGCCGCUUUGGCU-UCCAUCUCGGGCCACAGCAGCA .........((((....(((.....)))(((((........)))))...))))......(((.(((.(((((-(.......)))))).))).))) ( -26.80) >DroSec_CAF1 43512 95 + 1 CCACAAAAACCAGAGCUGGCAUAUUGCCGCUAUUGUCUUUUAUAGUUUCCUGGAACAUUUGCCGCUUUGGCUUUCCAUCUCGGGCCACAGCAGCA .........((((....(((.....)))(((((........)))))...))))......(((.(((.((((((........)))))).))).))) ( -27.10) >DroSim_CAF1 43940 94 + 1 CCACAAAAACCAGAGCUGGCAUAUUGCCGCUAUUGUCUUUUAUAGUUUCCUGGAACAUUUGCCGCUUUGGCU-UCCAUCUCGGGCCACAGCAGCA .........((((....(((.....)))(((((........)))))...))))......(((.(((.(((((-(.......)))))).))).))) ( -26.80) >DroEre_CAF1 49304 94 + 1 CCACAAAAACCAGAGCUGGCAUAUUGCCGCUAUUGUCUUUUAUAGUUUCCUGGAACAUUUGCCGCUUUGGCU-UCCAUCUCGGGCCACAGCAGCA .........((((....(((.....)))(((((........)))))...))))......(((.(((.(((((-(.......)))))).))).))) ( -26.80) >DroYak_CAF1 53853 94 + 1 CCACAAAAACCAGAGCUGGCAUAUUGCCGCUAUUGUCUUUUAUAGUUUCCUGGAACAUUUGCCGCUUUGGCU-UCCAUCUCGAGCCACAGCAGCA .........((((....(((.....)))(((((........)))))...))))......(((.(((.(((((-(.......)))))).))).))) ( -27.60) >DroAna_CAF1 75268 87 + 1 CCACAAAAACCAGAGCUGGCAUAUUGCCGCUAUUGUCUUUUAUAGUUUCCUGGAACAUUUGCCGCUUUGGCU-UCCAUCUUGAACCAC------- .........(((((((.((((.....(((((((........))))).....))......)))))))))))..-...............------- ( -20.10) >consensus CCACAAAAACCAGAGCUGGCAUAUUGCCGCUAUUGUCUUUUAUAGUUUCCUGGAACAUUUGCCGCUUUGGCU_UCCAUCUCGGGCCACAGCAGCA .........((((....(((.....)))(((((........)))))...)))).......((.(((.(((((..........))))).))).)). (-21.84 = -22.62 + 0.78)


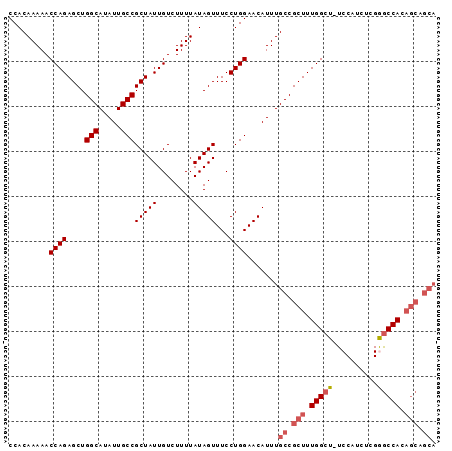
| Location | 20,994,761 – 20,994,855 |
|---|---|
| Length | 94 |
| Sequences | 6 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 95.90 |
| Mean single sequence MFE | -31.25 |
| Consensus MFE | -27.96 |
| Energy contribution | -28.77 |
| Covariance contribution | 0.81 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.66 |
| Structure conservation index | 0.89 |
| SVM decision value | 3.07 |
| SVM RNA-class probability | 0.998329 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 20994761 94 - 22407834 UGCUGCUGUGGCCCGAGAUGGA-AGCCAAAGCGGCAAAUGUUCCAGGAAACUAUAAAAGACAAUAGCGGCAAUAUGCCAGCUCUGGUUUUUGUGG (((((((.((((((.....)).-.)))).)))))))......((((....))...((((((...((((((.....))).)))...))))))..)) ( -32.60) >DroSec_CAF1 43512 95 - 1 UGCUGCUGUGGCCCGAGAUGGAAAGCCAAAGCGGCAAAUGUUCCAGGAAACUAUAAAAGACAAUAGCGGCAAUAUGCCAGCUCUGGUUUUUGUGG (((((((.((((((.....))...)))).)))))))......((((....))...((((((...((((((.....))).)))...))))))..)) ( -32.20) >DroSim_CAF1 43940 94 - 1 UGCUGCUGUGGCCCGAGAUGGA-AGCCAAAGCGGCAAAUGUUCCAGGAAACUAUAAAAGACAAUAGCGGCAAUAUGCCAGCUCUGGUUUUUGUGG (((((((.((((((.....)).-.)))).)))))))......((((....))...((((((...((((((.....))).)))...))))))..)) ( -32.60) >DroEre_CAF1 49304 94 - 1 UGCUGCUGUGGCCCGAGAUGGA-AGCCAAAGCGGCAAAUGUUCCAGGAAACUAUAAAAGACAAUAGCGGCAAUAUGCCAGCUCUGGUUUUUGUGG (((((((.((((((.....)).-.)))).)))))))......((((....))...((((((...((((((.....))).)))...))))))..)) ( -32.60) >DroYak_CAF1 53853 94 - 1 UGCUGCUGUGGCUCGAGAUGGA-AGCCAAAGCGGCAAAUGUUCCAGGAAACUAUAAAAGACAAUAGCGGCAAUAUGCCAGCUCUGGUUUUUGUGG (((((((.((((((......).-))))).)))))))......((((....))...((((((...((((((.....))).)))...))))))..)) ( -33.50) >DroAna_CAF1 75268 87 - 1 -------GUGGUUCAAGAUGGA-AGCCAAAGCGGCAAAUGUUCCAGGAAACUAUAAAAGACAAUAGCGGCAAUAUGCCAGCUCUGGUUUUUGUGG -------..........(..((-(((((.(((.((...((((..((....))......))))...))(((.....))).))).)))))))..).. ( -24.00) >consensus UGCUGCUGUGGCCCGAGAUGGA_AGCCAAAGCGGCAAAUGUUCCAGGAAACUAUAAAAGACAAUAGCGGCAAUAUGCCAGCUCUGGUUUUUGUGG (((((((.((((((.....))...)))).)))))))......((((....))...((((((...((((((.....))).)))...))))))..)) (-27.96 = -28.77 + 0.81)

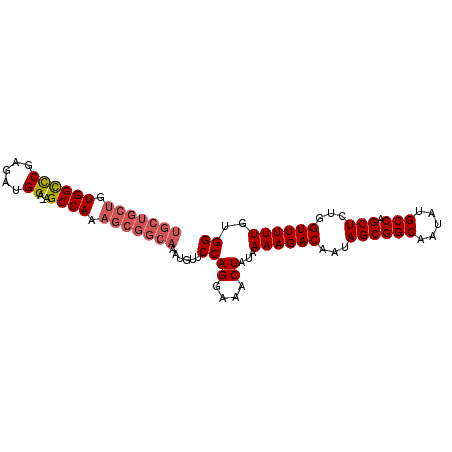
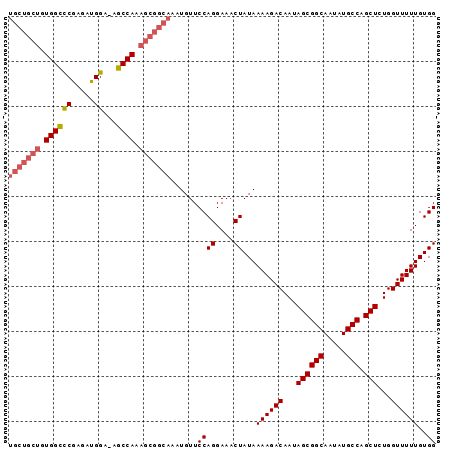
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:40:36 2006