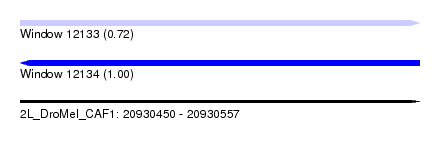

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 20,930,450 – 20,930,557 |
| Length | 107 |
| Max. P | 0.999722 |

| Location | 20,930,450 – 20,930,557 |
|---|---|
| Length | 107 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 76.22 |
| Mean single sequence MFE | -34.29 |
| Consensus MFE | -25.99 |
| Energy contribution | -27.00 |
| Covariance contribution | 1.01 |
| Combinations/Pair | 1.15 |
| Mean z-score | -1.30 |
| Structure conservation index | 0.76 |
| SVM decision value | 0.39 |
| SVM RNA-class probability | 0.717696 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 20930450 107 + 22407834 UGCAUUGAGAUGCCGAGUGCAUAUGCAUAUGCAAGCGGUGGCUGUCGUGUGGAGAAAGGGGUGAGUGGGGCUAUCCCCGGAAAAUGGGGCCGUUGUGAUUGGUGUUG------------- .((((..(...((((..((((((....))))))..))))((((.((((.........((((..((.....))..)))).....)))))))).......)..))))..------------- ( -33.34) >DroSec_CAF1 16857 109 + 1 UGCAUUGGAAAGCCGAGUGCAUAUGCAAAUGCAAACGGUGGCUGGCGUGUGAA---GGGGAUGAGUGGGGCUAUCCCCGGAAAAUGGGGCCGUUGUGAUUGGUAUGGUGUGU-------- ((((((((....)).)))))).(((((.((((..((..(((((..(((.....---((((((.((.....)))))))).....))).)))))..)).....))))..)))))-------- ( -35.30) >DroEre_CAF1 14324 114 + 1 UGCAUUGGGACGCCGAGUGCAUAUGCAUAUGCAAGCGGCGGGUAGCGUGUGGA---GGGG---GGUGAGGCUAUCCCCGGAAAAGGGGGCCGUUGUGAUUGGUNNNNNNNNNNNNNNNNN .(((.(((..(((((..((((((....))))))..))))).(((((.(.....---....---....).)))))((((......)))).))).)))........................ ( -34.22) >consensus UGCAUUGGGAAGCCGAGUGCAUAUGCAUAUGCAAGCGGUGGCUGGCGUGUGGA___GGGG_UGAGUGGGGCUAUCCCCGGAAAAUGGGGCCGUUGUGAUUGGU_U_G_____________ .((((.....(((((..((((((....))))))..)))))......))))......((((((.((.....))))))))..........((((.......))))................. (-25.99 = -27.00 + 1.01)


| Location | 20,930,450 – 20,930,557 |
|---|---|
| Length | 107 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 76.22 |
| Mean single sequence MFE | -23.87 |
| Consensus MFE | -17.19 |
| Energy contribution | -16.97 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.81 |
| Structure conservation index | 0.72 |
| SVM decision value | 3.95 |
| SVM RNA-class probability | 0.999722 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 20930450 107 - 22407834 -------------CAACACCAAUCACAACGGCCCCAUUUUCCGGGGAUAGCCCCACUCACCCCUUUCUCCACACGACAGCCACCGCUUGCAUAUGCAUAUGCACUCGGCAUCUCAAUGCA -------------................(((..........((((....))))...........((.......))..)))..((..((((((....))))))..))((((....)))). ( -20.80) >DroSec_CAF1 16857 109 - 1 --------ACACACCAUACCAAUCACAACGGCCCCAUUUUCCGGGGAUAGCCCCACUCAUCCCC---UUCACACGCCAGCCACCGUUUGCAUUUGCAUAUGCACUCGGCUUUCCAAUGCA --------.....................(((..........((((((((.....)).))))))---...........))).(((..(((((......)))))..)))............ ( -23.30) >DroEre_CAF1 14324 114 - 1 NNNNNNNNNNNNNNNNNACCAAUCACAACGGCCCCCUUUUCCGGGGAUAGCCUCACC---CCCC---UCCACACGCUACCCGCCGCUUGCAUAUGCAUAUGCACUCGGCGUCCCAAUGCA .............................(((((((......))))...))).....---....---.......((....(((((..((((((....))))))..))))).......)). ( -27.50) >consensus _____________C_A_ACCAAUCACAACGGCCCCAUUUUCCGGGGAUAGCCCCACUCA_CCCC___UCCACACGCCAGCCACCGCUUGCAUAUGCAUAUGCACUCGGCAUCCCAAUGCA ..............................((..........((((....))))............................(((..(((((......)))))..))).........)). (-17.19 = -16.97 + -0.22)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:39:33 2006