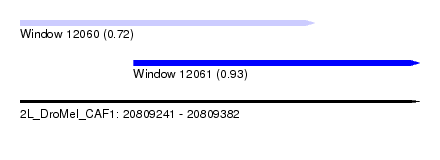
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 20,809,241 – 20,809,382 |
| Length | 141 |
| Max. P | 0.929534 |

| Location | 20,809,241 – 20,809,345 |
|---|---|
| Length | 104 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 79.91 |
| Mean single sequence MFE | -28.67 |
| Consensus MFE | -23.05 |
| Energy contribution | -23.36 |
| Covariance contribution | 0.31 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.58 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.39 |
| SVM RNA-class probability | 0.717348 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
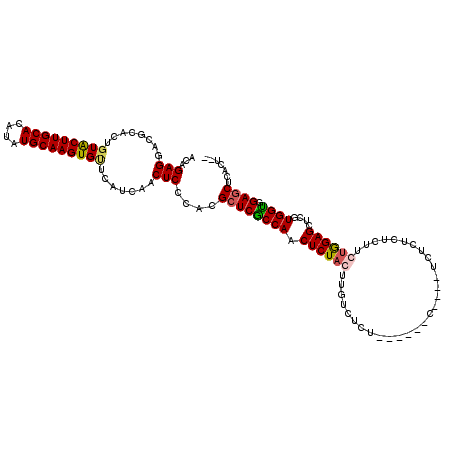
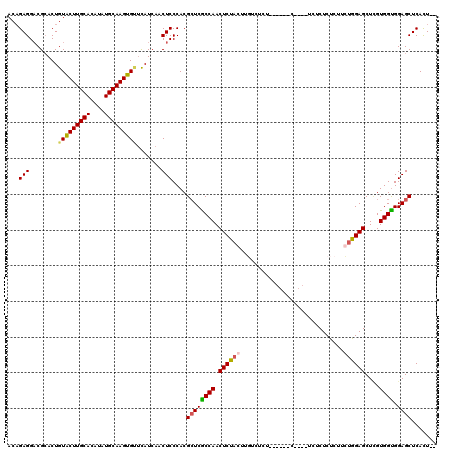
>2L_DroMel_CAF1 20809241 104 + 22407834 ACAGAGGACGCACUGUACUUGCACAUAUGCAAGUGCACAUCAACUCCCACGCUCACCAACUCUACUUGUCUCU--------------CUCUCUUCUAGAGCUCGUGGUGGAGCUCACU-- ...(((.......((((((((((....))))))))))......)))....((((((((.(((((.........--------------........)))))....)))).)))).....-- ( -28.95) >DroSim_CAF1 36629 108 + 1 ACAGAGGACGCACUGUACUUGCACAUAUGCAAGUGUUCAUCAACUCCCACGCUCGCCAACUCUAUUUGUCUCU------C----UCUCUCUCUUCUGGAGCUCGUGGUGGAGCUCACU-- .(((((((.((.....(((((((....)))))))(((....)))......))..(.(((......))).)...------.----......)))))))((((((......))))))...-- ( -27.00) >DroEre_CAF1 34763 107 + 1 ACAGAGGCCGCACUGUGCUUGCACAUAUGCAAGUGAUUAUCAACUCCCACGCUCGCCAACUCUACU-------------GUCUCUCUCUCUUUUGCGGAGCUGGUGGUGGAGCUCACUCC ...(((.((((.....(((((((....)))))))...................(((((.(((..(.-------------...............)..))).)))))))))..)))..... ( -28.59) >DroYak_CAF1 33205 117 + 1 ACUGAGGACGGACUGUGCUUGCACAUAUGCAAGUGUUCAUAAACUCCCACG-UCGCCAACUCUACUUCUCUCUUCAUCUCUAUCUCUCUCUUUUGUGGAGCUUGUGGCGGAGCUCAUU-- ..((((...((((...(((((((....))))))))))).....((((...(-((((...((((((.............................))))))...)))))))))))))..-- ( -30.15) >consensus ACAGAGGACGCACUGUACUUGCACAUAUGCAAGUGUUCAUCAACUCCCACGCUCGCCAACUCUACUUGUCUCU______C____UCUCUCUCUUCUGGAGCUCGUGGUGGAGCUCACU__ ...(((........(((((((((....))))))))).......)))....((((((((.((((((.............................))))))....)))).))))....... (-23.05 = -23.36 + 0.31)

| Location | 20,809,281 – 20,809,382 |
|---|---|
| Length | 101 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 78.48 |
| Mean single sequence MFE | -27.29 |
| Consensus MFE | -16.41 |
| Energy contribution | -16.47 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.23 |
| Mean z-score | -2.25 |
| Structure conservation index | 0.60 |
| SVM decision value | 1.20 |
| SVM RNA-class probability | 0.929534 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
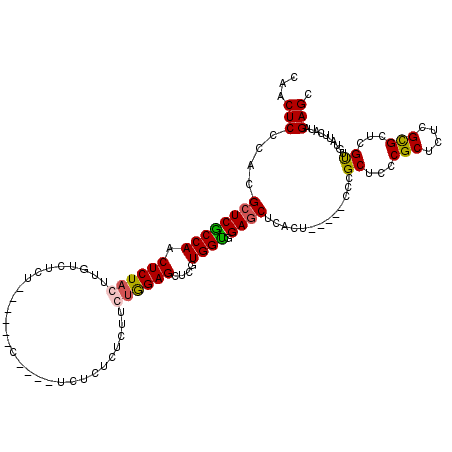
>2L_DroMel_CAF1 20809281 101 + 22407834 CAACUCCCACGCUCACCAACUCUACUUGUCUCU--------------CUCUCUUCUAGAGCUCGUGGUGGAGCUCACU-----CCCGCUCCCGCUCUCGUGCUCGUUAUAUUCAUAGAGC ..........((((..(((......))).....--------------..........((((.((.((((((((.....-----...))).)))))..)).))))............)))) ( -23.80) >DroSim_CAF1 36669 105 + 1 CAACUCCCACGCUCGCCAACUCUAUUUGUCUCU------C----UCUCUCUCUUCUGGAGCUCGUGGUGGAGCUCACU-----CCCGCUACCGCUCUCGCGCUCGUUGUAUUCAUAGAGC ..........(((((.(((......))).)...------.----.............((((.((.((((((((.....-----...))).)))))..)).))))............)))) ( -23.50) >DroEre_CAF1 34803 107 + 1 CAACUCCCACGCUCGCCAACUCUACU-------------GUCUCUCUCUCUUUUGCGGAGCUGGUGGUGGAGCUCACUCCGCUCCCGCUCCCGCUCUCGCGCUCGCUGUAUUCGUAGAGC ...................((((((.-------------...............(((((((.((..((((((....))))))..))))).))))....((....)).......)))))). ( -34.30) >DroYak_CAF1 33245 112 + 1 AAACUCCCACG-UCGCCAACUCUACUUCUCUCUUCAUCUCUAUCUCUCUCUUUUGUGGAGCUUGUGGCGGAGCUCAUU-----CCCGCUC-CGCUCUCGCGC-CGUUUUAUUCAUAGAGC ........(((-.((((((((((((.............................)))))).))).((((((((.....-----...))))-))))...))).-))).............. ( -27.55) >consensus CAACUCCCACGCUCGCCAACUCUACUUGUCUCU______C____UCUCUCUCUUCUGGAGCUCGUGGUGGAGCUCACU_____CCCGCUCCCGCUCUCGCGCUCGUUGUAUUCAUAGAGC ...(((....((((((((.((((((.............................))))))....)))).)))).............((...(((....)))...))..........))). (-16.41 = -16.47 + 0.06)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:38:26 2006