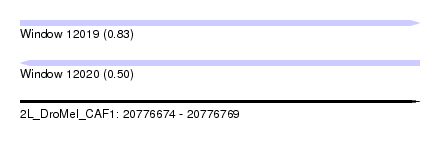

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 20,776,674 – 20,776,769 |
| Length | 95 |
| Max. P | 0.827667 |

| Location | 20,776,674 – 20,776,769 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 90.42 |
| Mean single sequence MFE | -21.32 |
| Consensus MFE | -18.64 |
| Energy contribution | -18.36 |
| Covariance contribution | -0.28 |
| Combinations/Pair | 1.18 |
| Mean z-score | -1.46 |
| Structure conservation index | 0.87 |
| SVM decision value | 0.70 |
| SVM RNA-class probability | 0.827667 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
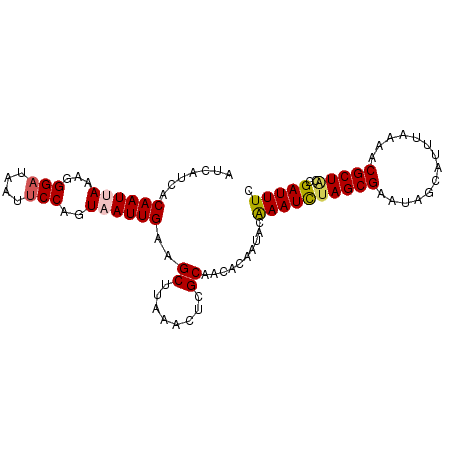
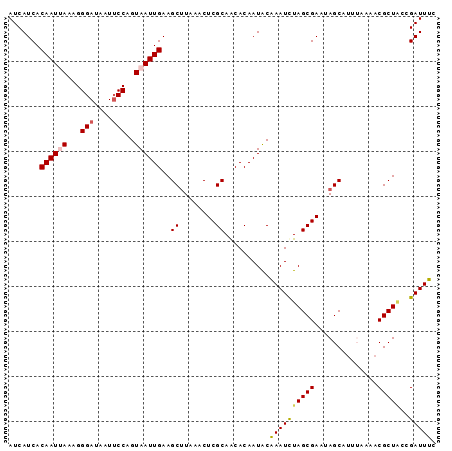
>2L_DroMel_CAF1 20776674 95 + 22407834 GAAAUCGCUAGCGUUUUAAAUGCUCUUCGCUAAAUUUGUAUUGUGUUGCGAGUUUAAGCUUCAAUUACUGGAAUUAUCCCUUUAAUUGUGAUGAU ...(((((.(((((.....)))))....((((((((((((......))))))))).)))...(((((..((.......))..))))))))))... ( -19.80) >DroSec_CAF1 28336 95 + 1 GAAAUCGGUAGCGUUUUAAAUGCUAUUCGCUAGAUUUGCAUUGUGUUGCGAGUUUAAGCUUCAAUUACUGGCAUUAUCCCUUUAAUUGUGAUGAU ...(((((((((((.....)))))))).((((((((((((......))))))))).)))..((((((..((.......))..)))))).)))... ( -23.30) >DroSim_CAF1 28392 95 + 1 GAAAUCGGUAGCGUUUUAAAUGCUAUUCGCUAGAUUUGCAUUGUGUUGCGAGUUUAAGCUUCAAUUACUGGAAUUAUCCCUUUAAUUGUGAUGAU ...(((((((((((.....)))))))).((((((((((((......))))))))).)))..((((((..((.......))..)))))).)))... ( -24.10) >DroEre_CAF1 27264 94 + 1 GAAAUCUGCAGCGUUUCUUAUGCUUUUCGCUAGAUUCGUAUUGUGCUGCGAGUUUAGGCUUCAAUUACUGGAAUUAUCCCU-UUAUUGUGAUGAU ...(((.(((((((.....))))).....(((((((((((......)))))))))))))..((((....(((....)))..-..)))).)))... ( -20.60) >DroYak_CAF1 29605 92 + 1 GAAAUCU--AGCGUUUCUAUUGCU-UUCGCUAGAUUUGUAUUGUGCUGCGAGUCUAAGCUUCAAUUACUGGAAUUAUCCCUUUUAUUGUGAUGAU .((((((--((((...........-..))))))))))(((...((..((........))..))..)))....((((((.(.......).)))))) ( -18.82) >consensus GAAAUCGGUAGCGUUUUAAAUGCUAUUCGCUAGAUUUGUAUUGUGUUGCGAGUUUAAGCUUCAAUUACUGGAAUUAUCCCUUUAAUUGUGAUGAU ...(((...(((((.....)))))....((((((((((((......))))))))).)))..((((((..((.......))..)))))).)))... (-18.64 = -18.36 + -0.28)



| Location | 20,776,674 – 20,776,769 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 90.42 |
| Mean single sequence MFE | -15.75 |
| Consensus MFE | -12.06 |
| Energy contribution | -12.42 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.46 |
| Structure conservation index | 0.77 |
| SVM decision value | -0.06 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 20776674 95 - 22407834 AUCAUCACAAUUAAAGGGAUAAUUCCAGUAAUUGAAGCUUAAACUCGCAACACAAUACAAAUUUAGCGAAGAGCAUUUAAAACGCUAGCGAUUUC ...(((.((((((...(((....)))..))))))..(((.....((((.................))))..(((.........)))))))))... ( -15.93) >DroSec_CAF1 28336 95 - 1 AUCAUCACAAUUAAAGGGAUAAUGCCAGUAAUUGAAGCUUAAACUCGCAACACAAUGCAAAUCUAGCGAAUAGCAUUUAAAACGCUACCGAUUUC ...(((.((((((...((......))..)))))).(((........(((......))).......((.....)).........)))...)))... ( -14.30) >DroSim_CAF1 28392 95 - 1 AUCAUCACAAUUAAAGGGAUAAUUCCAGUAAUUGAAGCUUAAACUCGCAACACAAUGCAAAUCUAGCGAAUAGCAUUUAAAACGCUACCGAUUUC ...(((.((((((...(((....)))..)))))).(((........(((......))).......((.....)).........)))...)))... ( -16.30) >DroEre_CAF1 27264 94 - 1 AUCAUCACAAUAA-AGGGAUAAUUCCAGUAAUUGAAGCCUAAACUCGCAGCACAAUACGAAUCUAGCGAAAAGCAUAAGAAACGCUGCAGAUUUC ...(((.((((.(-..(((....)))..).))))............(((((.(.....)......((.....)).........))))).)))... ( -15.90) >DroYak_CAF1 29605 92 - 1 AUCAUCACAAUAAAAGGGAUAAUUCCAGUAAUUGAAGCUUAGACUCGCAGCACAAUACAAAUCUAGCGAA-AGCAAUAGAAACGCU--AGAUUUC .......((((.(...(((....)))..).))))..((........))..........((((((((((..-...........))))--)))))). ( -16.32) >consensus AUCAUCACAAUUAAAGGGAUAAUUCCAGUAAUUGAAGCUUAAACUCGCAACACAAUACAAAUCUAGCGAAUAGCAUUUAAAACGCUACCGAUUUC .......((((((...(((....)))..))))))..((........))..........((((((((((..............)))))..))))). (-12.06 = -12.42 + 0.36)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:37:48 2006