
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 20,572,086 – 20,572,189 |
| Length | 103 |
| Max. P | 0.996055 |

| Location | 20,572,086 – 20,572,189 |
|---|---|
| Length | 103 |
| Sequences | 5 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 68.42 |
| Mean single sequence MFE | -28.33 |
| Consensus MFE | -16.78 |
| Energy contribution | -17.82 |
| Covariance contribution | 1.04 |
| Combinations/Pair | 1.30 |
| Mean z-score | -1.91 |
| Structure conservation index | 0.59 |
| SVM decision value | 1.66 |
| SVM RNA-class probability | 0.970774 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 20572086 103 + 22407834 UCUGUUGCAUUCGCAAUUCCGCAUUGGAUACGGAAAUGCCACGCACAGUGGUGUUCUCUGUUUCCAGAGCUAUUAGUUUUGAGUGGGAUCGUAUUGGAGUAUC-------------- .....(((.(((((.(((((((...(((.(((((((((((((.....)))))))).))))).)))(((((.....)))))..))))))).))...))))))..-------------- ( -32.40) >DroSec_CAF1 86891 94 + 1 UCUGUUGCAUUCGCAAUUCCGCAUUGGAUACGGAAAUACCACGCACAGUGGUGUUCUCUGUUUCUAAAUUUAUUAGUUUUGAGUGGUCUCAUAU----------------------- ...(((((....))))).((((.(((((.(((((((((((((.....)))))))).))))).)))))..(((.......)))))))........----------------------- ( -27.10) >DroSim_CAF1 87594 91 + 1 UCUGUUGCAUUCGCAAUUCCGCAUUGGAUACGGAAAUGCCACGCACAGUGGUGUUCUCUGUUUCUAAAUUUAUUAGGUGUGAGUGGUCUCA-------------------------- .......((((((((........(((((.(((((((((((((.....)))))))).))))).)))))..........))))))))......-------------------------- ( -28.37) >DroEre_CAF1 78916 105 + 1 UCUGCUGCACUCGCGAUUCCGCAUCAGACACGGAAAUACCUCGCACACUGGUGC-UUCUGU-UUAGGAGGUUUUAGAUUU----------AUAUGUGGGAAACAGCAAAAGUAUCUG ..(((((..((((((.(((((.........)))))..(((((((((....))))-..((..-..))))))).........----------...))))))...))))).......... ( -27.00) >DroYak_CAF1 81193 105 + 1 UCUGUUGCAUUCGCGUAUCUAAUGCGGAUACGGAAAUACCACGCACACUGGUAUUUUCUGUUUUAAGAGGUAUUAGUUUUAAGUGAGCUCAUAUAUGAGAAACAA------------ ......(.((((((((.....)))))))).)((((((((((.......))))))))))((((((.....((((.(((((.....))))).))))....)))))).------------ ( -26.80) >consensus UCUGUUGCAUUCGCAAUUCCGCAUUGGAUACGGAAAUACCACGCACAGUGGUGUUCUCUGUUUCAAGAGGUAUUAGUUUUGAGUGGGCUCAUAU__G_G_A_C______________ ...(((((....)))))......(((((.(((((((((((((.....)))))))).))))).))))).................................................. (-16.78 = -17.82 + 1.04)


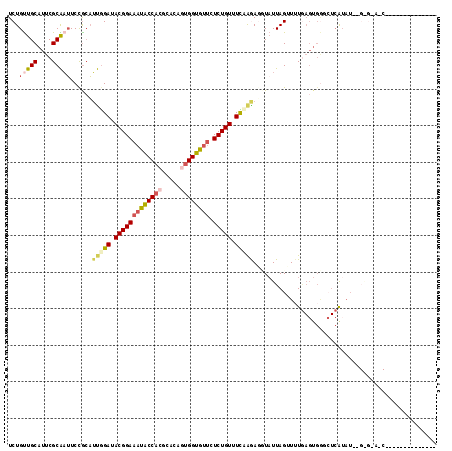
| Location | 20,572,086 – 20,572,189 |
|---|---|
| Length | 103 |
| Sequences | 5 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 68.42 |
| Mean single sequence MFE | -25.55 |
| Consensus MFE | -14.32 |
| Energy contribution | -15.44 |
| Covariance contribution | 1.12 |
| Combinations/Pair | 1.22 |
| Mean z-score | -2.36 |
| Structure conservation index | 0.56 |
| SVM decision value | 2.65 |
| SVM RNA-class probability | 0.996055 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 20572086 103 - 22407834 --------------GAUACUCCAAUACGAUCCCACUCAAAACUAAUAGCUCUGGAAACAGAGAACACCACUGUGCGUGGCAUUUCCGUAUCCAAUGCGGAAUUGCGAAUGCAACAGA --------------..................................(((((....))))).......((((((((.(((.((((((((...)))))))).)))..)))).)))). ( -32.40) >DroSec_CAF1 86891 94 - 1 -----------------------AUAUGAGACCACUCAAAACUAAUAAAUUUAGAAACAGAGAACACCACUGUGCGUGGUAUUUCCGUAUCCAAUGCGGAAUUGCGAAUGCAACAGA -----------------------...((((....))))...((((.....))))...............((((((((.(((.((((((((...)))))))).)))..)))).)))). ( -24.70) >DroSim_CAF1 87594 91 - 1 --------------------------UGAGACCACUCACACCUAAUAAAUUUAGAAACAGAGAACACCACUGUGCGUGGCAUUUCCGUAUCCAAUGCGGAAUUGCGAAUGCAACAGA --------------------------((((....))))...((((.....))))...............((((((((.(((.((((((((...)))))))).)))..)))).)))). ( -25.90) >DroEre_CAF1 78916 105 - 1 CAGAUACUUUUGCUGUUUCCCACAUAU----------AAAUCUAAAACCUCCUAA-ACAGAA-GCACCAGUGUGCGAGGUAUUUCCGUGUCUGAUGCGGAAUCGCGAGUGCAGCAGA ........(((((((((((.(......----------.........(((((((..-..))..-((((....)))))))))..((((((((...))))))))..).))).)))))))) ( -27.10) >DroYak_CAF1 81193 105 - 1 ------------UUGUUUCUCAUAUAUGAGCUCACUUAAAACUAAUACCUCUUAAAACAGAAAAUACCAGUGUGCGUGGUAUUUCCGUAUCCGCAUUAGAUACGCGAAUGCAACAGA ------------.................((.((((................................)))).))((.((((((.((((((.......)))))).)))))).))... ( -17.65) >consensus ______________G_U_C_C__AUAUGAGACCACUCAAAACUAAUAACUCUAGAAACAGAGAACACCACUGUGCGUGGUAUUUCCGUAUCCAAUGCGGAAUUGCGAAUGCAACAGA .....................................................................((((((((.(((.(((((((.....))))))).)))..)))).)))). (-14.32 = -15.44 + 1.12)

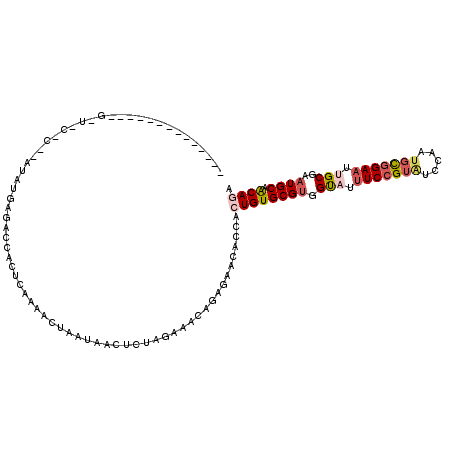

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:35:39 2006