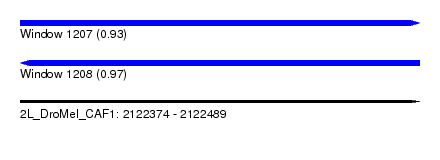

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 2,122,374 – 2,122,489 |
| Length | 115 |
| Max. P | 0.967982 |

| Location | 2,122,374 – 2,122,489 |
|---|---|
| Length | 115 |
| Sequences | 5 |
| Columns | 115 |
| Reading direction | forward |
| Mean pairwise identity | 88.89 |
| Mean single sequence MFE | -37.74 |
| Consensus MFE | -27.82 |
| Energy contribution | -30.06 |
| Covariance contribution | 2.24 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.55 |
| Structure conservation index | 0.74 |
| SVM decision value | 1.24 |
| SVM RNA-class probability | 0.934902 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
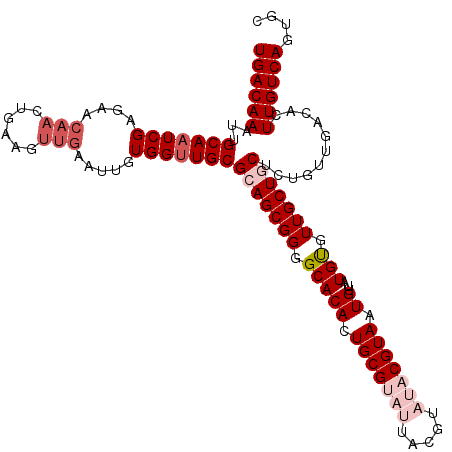
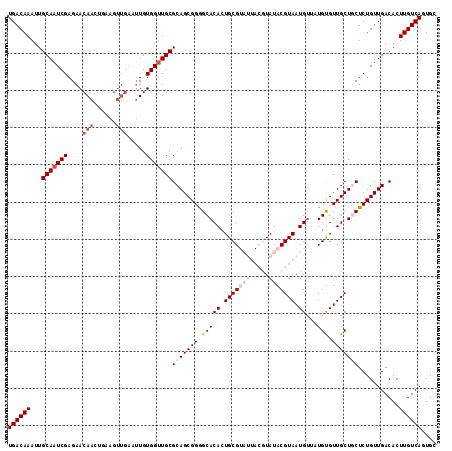
>2L_DroMel_CAF1 2122374 115 + 22407834 UGACAAAUUGCAAUCGAGAACAAGUGAAGUUGAAUUGUGGUUGCGCAGCGGAGCACACUGCGUAUUACGUAUACGUAAUGUUAUGCGUUGCUGCUCUGUUGACACUUGUCAGUGC ((((((...(((((((....(((......))).....)))))))(((((((((((((.(((((((((((....)))))...)))))).)).)))))))))).)..)))))).... ( -40.40) >DroSec_CAF1 122672 109 + 1 UGACAAAUUGCAAUCGAGAACAAAUGAAG------UGUGGUUGCGCAGCGGGGCACACUGCGUAUUACGUAUACGUAAUGUUUUGUGUUGCUGCUCUGUUGACACUUGUCAGUGC ((((((...(((((((...((.......)------).)))))))(((((((((((((..((((((((((....)))))))...)))..)).)))))))))).)..)))))).... ( -37.00) >DroSim_CAF1 122441 115 + 1 UGACAAAUUGCAAUCGAGAACAAGUGAAGUUGAAUUGUGGUUGCGCAGCGGGGCACAGUGCGUAUUACGUAUACGUAAUGUUUUGUGUUGCUGCUCUGUUGACACUUGUCAGUGC ((((((...(((((((....(((......))).....)))))))((((((((((((((..(((((((((....)))))))...))..))).)))))))))).)..)))))).... ( -39.90) >DroEre_CAF1 125369 105 + 1 UGACAAAUUGCA-UCGAGAACAACACAAGUUGAGUUGUGGUUGCGCAGCGGGGCACACUGCG-------UAUACGUAAUGC--UGUGUUGCUCCUCUGUUGACACUUGUCAAUGC ((((((...(((-......(((((.(.....).)))))...)))((((.(((((((((.(((-------(.......))))--.))).)))))).))))......)))))).... ( -34.80) >DroYak_CAF1 127720 108 + 1 UGACAAAUUGCAAUCGAGAACAACUGAAGUUGAAUUGUGGUUGCGCAGCGGGGCACACUGCG-------UAUACGUAAUGUUAUGUGUUGCUCCUCUGUUGACACUUGUCAGUGC ((((((...(((((((....((((....)))).....)))))))((((.(((((((((.(((-------(.......))))...))).)))))).))))......)))))).... ( -36.60) >consensus UGACAAAUUGCAAUCGAGAACAACUGAAGUUGAAUUGUGGUUGCGCAGCGGGGCACACUGCGUAUUACGUAUACGUAAUGUUAUGUGUUGCUGCUCUGUUGACACUUGUCAGUGC ((((((...(((((((....(((......))).....)))))))(((((((.(((((.(((((((.....))))))).))...))).)))))))...........)))))).... (-27.82 = -30.06 + 2.24)

| Location | 2,122,374 – 2,122,489 |
|---|---|
| Length | 115 |
| Sequences | 5 |
| Columns | 115 |
| Reading direction | reverse |
| Mean pairwise identity | 88.89 |
| Mean single sequence MFE | -30.10 |
| Consensus MFE | -23.12 |
| Energy contribution | -23.36 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.50 |
| Structure conservation index | 0.77 |
| SVM decision value | 1.62 |
| SVM RNA-class probability | 0.967982 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
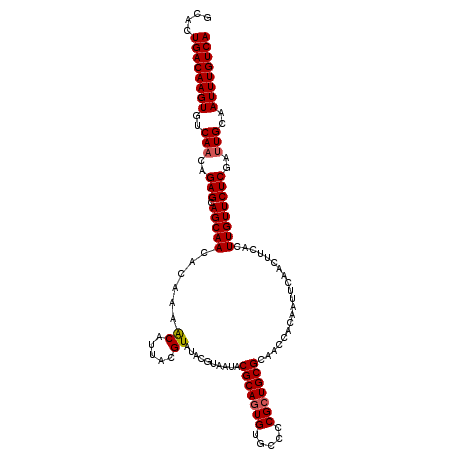
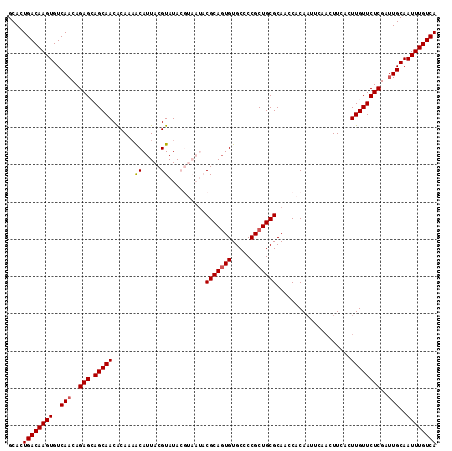
>2L_DroMel_CAF1 2122374 115 - 22407834 GCACUGACAAGUGUCAACAGAGCAGCAACGCAUAACAUUACGUAUACGUAAUACGCAGUGUGCUCCGCUGCGCAACCACAAUUCAACUUCACUUGUUCUCGAUUGCAAUUUGUCA .....(((((((..(((..(((.(((((........((((((....)))))).(((((((.....)))))))....................))))))))..)))..))))))). ( -31.20) >DroSec_CAF1 122672 109 - 1 GCACUGACAAGUGUCAACAGAGCAGCAACACAAAACAUUACGUAUACGUAAUACGCAGUGUGCCCCGCUGCGCAACCACA------CUUCAUUUGUUCUCGAUUGCAAUUUGUCA .....(((((((..(((..(((.(((((........((((((....)))))).(((((((.....)))))))........------......))))))))..)))..))))))). ( -31.30) >DroSim_CAF1 122441 115 - 1 GCACUGACAAGUGUCAACAGAGCAGCAACACAAAACAUUACGUAUACGUAAUACGCACUGUGCCCCGCUGCGCAACCACAAUUCAACUUCACUUGUUCUCGAUUGCAAUUUGUCA .....(((((((..(((..(((.(((((........((((((....))))))......(((((......)))))..................))))))))..)))..))))))). ( -26.20) >DroEre_CAF1 125369 105 - 1 GCAUUGACAAGUGUCAACAGAGGAGCAACACA--GCAUUACGUAUA-------CGCAGUGUGCCCCGCUGCGCAACCACAACUCAACUUGUGUUGUUCUCGA-UGCAAUUUGUCA ((((((((....)))....(((.(((((((((--((.....))...-------(((((((.....)))))))................))))))))))))))-)))......... ( -32.70) >DroYak_CAF1 127720 108 - 1 GCACUGACAAGUGUCAACAGAGGAGCAACACAUAACAUUACGUAUA-------CGCAGUGUGCCCCGCUGCGCAACCACAAUUCAACUUCAGUUGUUCUCGAUUGCAAUUUGUCA .....(((((((..(((..(((.((((((.....((.....))...-------(((((((.....)))))))...................)))))))))..)))..))))))). ( -29.10) >consensus GCACUGACAAGUGUCAACAGAGCAGCAACACAAAACAUUACGUAUACGUAAUACGCAGUGUGCCCCGCUGCGCAACCACAAUUCAACUUCACUUGUUCUCGAUUGCAAUUUGUCA ....((((((((..(((..(((.(((((......((.....))..........(((((((.....)))))))....................))))))))..)))..)))))))) (-23.12 = -23.36 + 0.24)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:44:17 2006