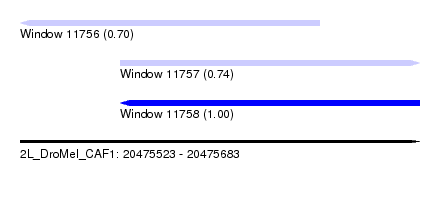
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 20,475,523 – 20,475,683 |
| Length | 160 |
| Max. P | 0.998646 |

| Location | 20,475,523 – 20,475,643 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 98.67 |
| Mean single sequence MFE | -25.46 |
| Consensus MFE | -24.14 |
| Energy contribution | -23.98 |
| Covariance contribution | -0.16 |
| Combinations/Pair | 1.02 |
| Mean z-score | -1.31 |
| Structure conservation index | 0.95 |
| SVM decision value | 0.35 |
| SVM RNA-class probability | 0.700607 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 20475523 120 - 22407834 GAAGGCGAAAUGCGAUCCGUAUUACACAAAAUCCCAUUUGUAUAUUUAUACACCAGGGCCCACAUGUGUGUGCUCUUCAAAUACACGAAAAUAUUUUUGAAAAUAUCUCGAAUGUUUGUU (((((((.((((((...))))))(((((....(((...(((((....)))))...)))......))))).))).))))..........((((((((..((.....))..))))))))... ( -24.20) >DroSec_CAF1 28533 120 - 1 GAAGGCGAAAUGCGAUCCGUAUUACACAAAAUCCCAUUUGUAUAUUUAUACACCAGGGCCCACAUGUGUGUGCUCUUCAAAUACACGAAAAUAUUUUUGAAAAUAUCUCGAAUGUUUGUU (((((((.((((((...))))))(((((....(((...(((((....)))))...)))......))))).))).))))..........((((((((..((.....))..))))))))... ( -24.20) >DroSim_CAF1 32308 120 - 1 GAAGGCGAAAUGCGAUCCGUAUUACACAAAAUCCCAUUUGUAUAUUUAUACACCAGGGCCCACAUGUGUGUGCUCUUCAAAUACACGAAAAUAUUUUUGAAAAUAUCUCGAAUGUUUGUU (((((((.((((((...))))))(((((....(((...(((((....)))))...)))......))))).))).))))..........((((((((..((.....))..))))))))... ( -24.20) >DroEre_CAF1 30720 120 - 1 GAAGGCGAAAUGCGAUCCGUAUUACACAAAAUCCCAUUUGUGUAUUUAUACACCGGGGCCCACAUGUGUGUGCUCUUCAAAUACACGAGAAUAUUUUUGAAAAUAUCUCGAAUGUUUGUU ...((.(.((((((...))))))(((((((......))))))).......).))(((((.(((....))).))))).((((((..(((((.(((((....))))))))))..)))))).. ( -30.60) >DroYak_CAF1 27929 120 - 1 GAAGGCGAAAUGCGAUCCGUAUUACACAAAAUCCCAUUUGUAUAUUUAUACACCAGGGCCCACAUGUGUGUGCUCUUCAAAUACACGAAAAUAUUUUUCAAAAUAUCUCGAAUGUUUGUU (((((((.((((((...))))))(((((....(((...(((((....)))))...)))......))))).))).))))(((((..(((..((((((....)))))).)))..)))))... ( -24.10) >consensus GAAGGCGAAAUGCGAUCCGUAUUACACAAAAUCCCAUUUGUAUAUUUAUACACCAGGGCCCACAUGUGUGUGCUCUUCAAAUACACGAAAAUAUUUUUGAAAAUAUCUCGAAUGUUUGUU (((((((.((((((...))))))(((((....(((...(((((....)))))...)))......))))).))).))))(((((..(((..((((((....)))))).)))..)))))... (-24.14 = -23.98 + -0.16)

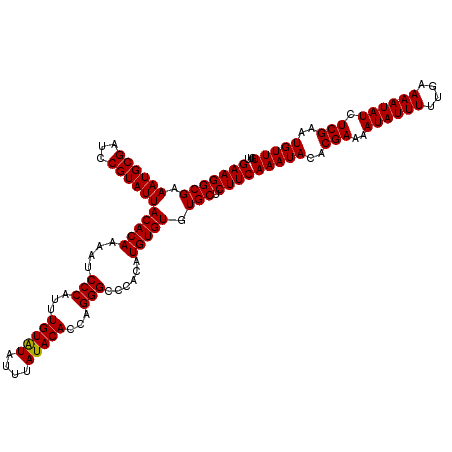
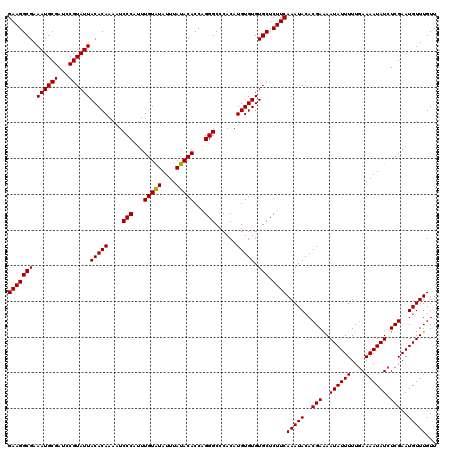
| Location | 20,475,563 – 20,475,683 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 96.99 |
| Mean single sequence MFE | -30.46 |
| Consensus MFE | -29.26 |
| Energy contribution | -29.66 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.20 |
| Structure conservation index | 0.96 |
| SVM decision value | 0.46 |
| SVM RNA-class probability | 0.743528 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 20475563 120 + 22407834 UUGAAGAGCACACACAUGUGGGCCCUGGUGUAUAAAUAUACAAAUGGGAUUUUGUGUAAUACGGAUCGCAUUUCGCCUUCUGUCUGUCCGACGUUGUUUAAAAAUAAAUCCAUUCGGACU ..((((.((..(((((......(((...(((((....)))))...)))....))))).....(((......))))))))).....((((((....(((((....)))))....)))))). ( -30.80) >DroSec_CAF1 28573 120 + 1 UUGAAGAGCACACACAUGUGGGCCCUGGUGUAUAAAUAUACAAAUGGGAUUUUGUGUAAUACGGAUCGCAUUUCGCCUUCUGUCUGUCCGACGUUGUUUAAAAAUAAAUCCAUUCGGACU ..((((.((..(((((......(((...(((((....)))))...)))....))))).....(((......))))))))).....((((((....(((((....)))))....)))))). ( -30.80) >DroSim_CAF1 32348 120 + 1 UUGAAGAGCACACACAUGUGGGCCCUGGUGUAUAAAUAUACAAAUGGGAUUUUGUGUAAUACGGAUCGCAUUUCGCCUUCUGUCUGUCCGACGUUGUUUAAAAAUAAAUCCAUUCGGACU ..((((.((..(((((......(((...(((((....)))))...)))....))))).....(((......))))))))).....((((((....(((((....)))))....)))))). ( -30.80) >DroEre_CAF1 30760 116 + 1 UUGAAGAGCACACACAUGUGGGCCCCGGUGUAUAAAUACACAAAUGGGAUUUUGUGUAAUACGGAUCGCAUUUCGCCUUCU----GUCCGCCGUUGUUUAAAAAUAAAUCCAUUCGGACU (((((.(((.(((....)))......((((......((((((((......))))))))..(((((..((.....))..)))----)).))))))).))))).......(((....))).. ( -29.10) >DroYak_CAF1 27969 116 + 1 UUGAAGAGCACACACAUGUGGGCCCUGGUGUAUAAAUAUACAAAUGGGAUUUUGUGUAAUACGGAUCGCAUUUCGCCUUCU----GUCCGACGUUGUUUAAAAAUAAAUCCAUUCGGACU ..((((.((..(((((......(((...(((((....)))))...)))....))))).....(((......))))))))).----((((((....(((((....)))))....)))))). ( -30.80) >consensus UUGAAGAGCACACACAUGUGGGCCCUGGUGUAUAAAUAUACAAAUGGGAUUUUGUGUAAUACGGAUCGCAUUUCGCCUUCUGUCUGUCCGACGUUGUUUAAAAAUAAAUCCAUUCGGACU ..((((.((..(((((......(((...(((((....)))))...)))....))))).....(((......))))))))).....((((((....(((((....)))))....)))))). (-29.26 = -29.66 + 0.40)

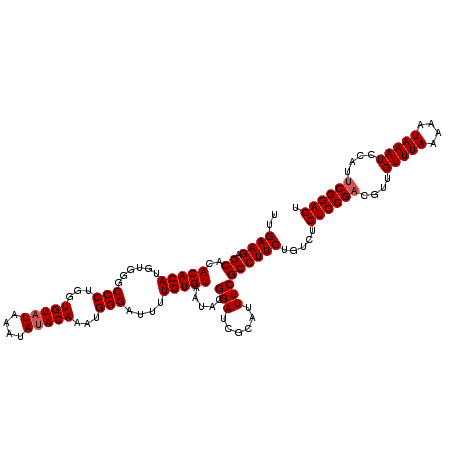
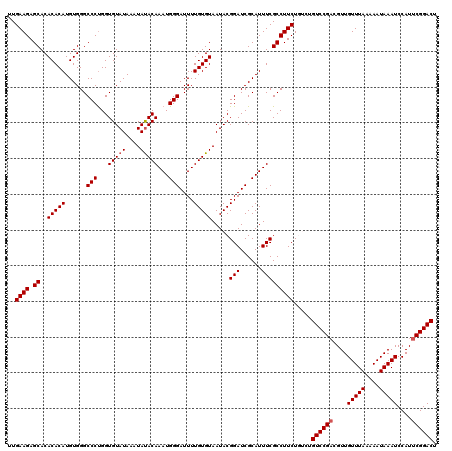
| Location | 20,475,563 – 20,475,683 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 96.99 |
| Mean single sequence MFE | -30.86 |
| Consensus MFE | -29.66 |
| Energy contribution | -29.70 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.03 |
| Structure conservation index | 0.96 |
| SVM decision value | 3.17 |
| SVM RNA-class probability | 0.998646 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
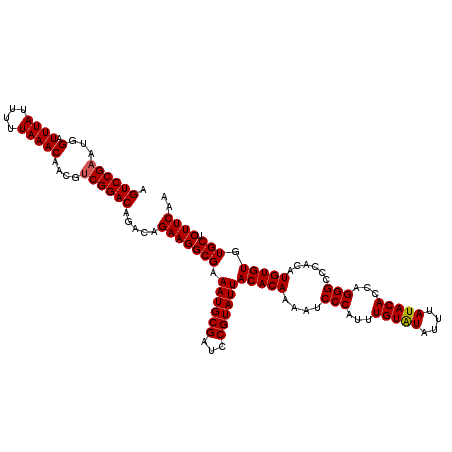
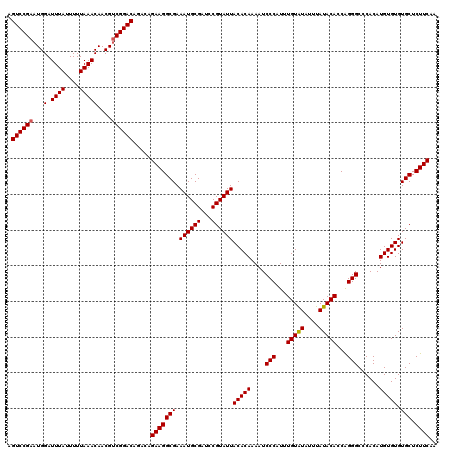
>2L_DroMel_CAF1 20475563 120 - 22407834 AGUCCGAAUGGAUUUAUUUUUAAACAACGUCGGACAGACAGAAGGCGAAAUGCGAUCCGUAUUACACAAAAUCCCAUUUGUAUAUUUAUACACCAGGGCCCACAUGUGUGUGCUCUUCAA .((((((...(.((((....)))))....)))))).....(((((((.((((((...))))))(((((....(((...(((((....)))))...)))......))))).))).)))).. ( -30.90) >DroSec_CAF1 28573 120 - 1 AGUCCGAAUGGAUUUAUUUUUAAACAACGUCGGACAGACAGAAGGCGAAAUGCGAUCCGUAUUACACAAAAUCCCAUUUGUAUAUUUAUACACCAGGGCCCACAUGUGUGUGCUCUUCAA .((((((...(.((((....)))))....)))))).....(((((((.((((((...))))))(((((....(((...(((((....)))))...)))......))))).))).)))).. ( -30.90) >DroSim_CAF1 32348 120 - 1 AGUCCGAAUGGAUUUAUUUUUAAACAACGUCGGACAGACAGAAGGCGAAAUGCGAUCCGUAUUACACAAAAUCCCAUUUGUAUAUUUAUACACCAGGGCCCACAUGUGUGUGCUCUUCAA .((((((...(.((((....)))))....)))))).....(((((((.((((((...))))))(((((....(((...(((((....)))))...)))......))))).))).)))).. ( -30.90) >DroEre_CAF1 30760 116 - 1 AGUCCGAAUGGAUUUAUUUUUAAACAACGGCGGAC----AGAAGGCGAAAUGCGAUCCGUAUUACACAAAAUCCCAUUUGUGUAUUUAUACACCGGGGCCCACAUGUGUGUGCUCUUCAA .(((((....(.((((....))))).....)))))----.(((((((.((((((...))))))(((((....(((....(((((....)))))..)))......))))).))).)))).. ( -30.70) >DroYak_CAF1 27969 116 - 1 AGUCCGAAUGGAUUUAUUUUUAAACAACGUCGGAC----AGAAGGCGAAAUGCGAUCCGUAUUACACAAAAUCCCAUUUGUAUAUUUAUACACCAGGGCCCACAUGUGUGUGCUCUUCAA .((((((...(.((((....)))))....))))))----.(((((((.((((((...))))))(((((....(((...(((((....)))))...)))......))))).))).)))).. ( -30.90) >consensus AGUCCGAAUGGAUUUAUUUUUAAACAACGUCGGACAGACAGAAGGCGAAAUGCGAUCCGUAUUACACAAAAUCCCAUUUGUAUAUUUAUACACCAGGGCCCACAUGUGUGUGCUCUUCAA .((((((...(.((((....)))))....)))))).....(((((((.((((((...))))))(((((....(((...(((((....)))))...)))......))))).))).)))).. (-29.66 = -29.70 + 0.04)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:33:51 2006