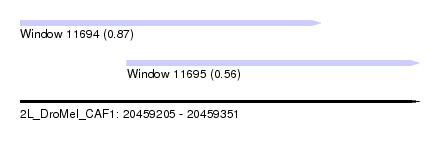
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 20,459,205 – 20,459,351 |
| Length | 146 |
| Max. P | 0.865824 |

| Location | 20,459,205 – 20,459,315 |
|---|---|
| Length | 110 |
| Sequences | 4 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 84.93 |
| Mean single sequence MFE | -23.73 |
| Consensus MFE | -17.05 |
| Energy contribution | -17.68 |
| Covariance contribution | 0.63 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.07 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.85 |
| SVM RNA-class probability | 0.865824 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 20459205 110 + 22407834 CUAAAAUGCUGUAAGAGGCAGUGACGAAAAAAGCGAGGAAAGCUGGGAAAGCUGGGAAAAGGGAAAAUUCCUACCGGGCUCUUCACGUAAUGAAAAA--------AUAAGGGAAACCG .......(((((.....))))).(((.....(((.......))).(((.((((.((...(((((...))))).)).)))).))).))).........--------....((....)). ( -27.40) >DroSec_CAF1 12244 101 + 1 CUAAAAUGCUGUAAGAGGCAGUGACGAAAAAA---------GCUGGGAAAGCUGGGAAAAGGGAAAAUUCCUACCGGGCUCUUCACGUAAUGAAAAA--------AUAAGGGAAACCG .......(((((.....))))).(((..((.(---------(((.((.....((((((.........)))))))).)))).))..))).........--------....((....)). ( -23.30) >DroSim_CAF1 14067 94 + 1 CUAAAAUGCUGUAAGAGGCAGUGACGAAAAAA---------GCUGGGAAAGCUGGGA-------AAAUUCCUACCGGGCUCUUCACGUAAUGAAAAA--------AUAAGGGAAACCG .......(((((.....))))).(((..((.(---------(((.((.....(((((-------....))))))).)))).))..))).........--------....((....)). ( -23.20) >DroEre_CAF1 14675 108 + 1 CUAAAAUGCUGUAAGCGGCAGUGACGAAA-AA---------GCAGGGAAAGCUGGGCAAAGGGAAAAUUCCUGUUGGGCUCUUCACGUAAUGAAAAAAUAAAAAAAUAAAGGAAGCCG ......(((((....)))))((((.((..-.(---------(((((((...((........))....))))))))....)).))))................................ ( -21.00) >consensus CUAAAAUGCUGUAAGAGGCAGUGACGAAAAAA_________GCUGGGAAAGCUGGGAAAAGGGAAAAUUCCUACCGGGCUCUUCACGUAAUGAAAAA________AUAAGGGAAACCG .......(((((.....))))).......................(((.((((.((....(((......))).)).)))).))).........................((....)). (-17.05 = -17.68 + 0.63)

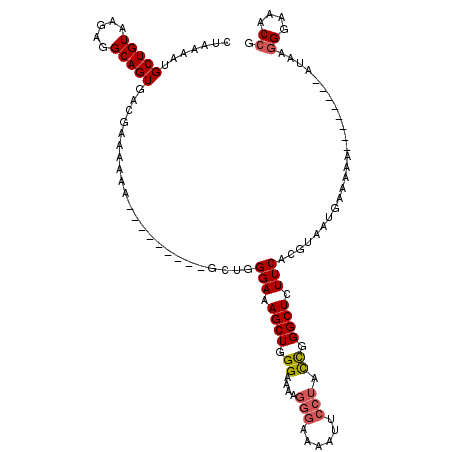
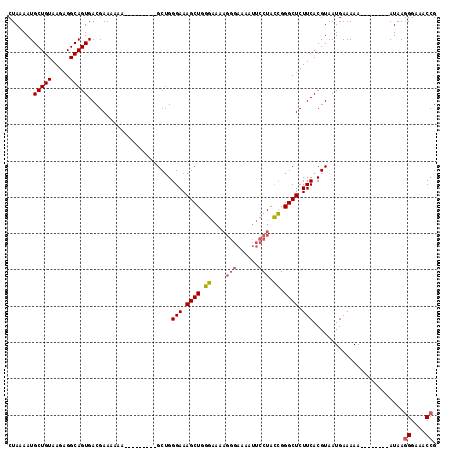
| Location | 20,459,244 – 20,459,351 |
|---|---|
| Length | 107 |
| Sequences | 4 |
| Columns | 112 |
| Reading direction | forward |
| Mean pairwise identity | 87.93 |
| Mean single sequence MFE | -24.06 |
| Consensus MFE | -16.50 |
| Energy contribution | -17.32 |
| Covariance contribution | 0.81 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.64 |
| Structure conservation index | 0.69 |
| SVM decision value | 0.06 |
| SVM RNA-class probability | 0.564457 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 20459244 107 + 22407834 AAGCUGGGAA-AGCUGGGAAAAGGGAAAAUUCCUACCGGGCUCUUCACGUAAUGAAAAAAUAAGGGAAACCGAAAAUCCAUUGAAAUGAAGUUCACUGAAGCAAAU----GA ..((((((((-((((.((...(((((...))))).)).))))(((((..(((((.........((....)).......)))))...)))))))).))..)))....----.. ( -25.69) >DroSec_CAF1 12276 109 + 1 --GCUGGGAA-AGCUGGGAAAAGGGAAAAUUCCUACCGGGCUCUUCACGUAAUGAAAAAAUAAGGGAAACCGAAAAUCCAUUGAAAUGAAGUUCACUGAAGCAUUUUUUGGU --((..((((-((((.......((((....))))..((((..(((((..(((((.........((....)).......)))))...)))))..).))).))).)))))..)) ( -26.09) >DroSim_CAF1 14099 102 + 1 --GCUGGGAA-AGCUGGGA-------AAAUUCCUACCGGGCUCUUCACGUAAUGAAAAAAUAAGGGAAACCGAAAAUCCAUUGAAAUGAAGUUCACUGAAGCAUUUUUUGGU --((..((((-((((((((-------....))))..((((..(((((..(((((.........((....)).......)))))...)))))..).))).))).)))))..)) ( -25.99) >DroYak_CAF1 12562 110 + 1 --GCAAGGAAAAAAAGGGAAAAGCGAAAAUUCCUGCCGGGCUCUUCACGUAAUGAAAAAAUAAAGGAACCCGAAAAUCCAUUGAAAUGAAGUUCAUUGAAGCAUUUUUUGGU --.(((((((.....(((((.........)))))((((((.((((..................)))).))))......((.((((......)))).))..)).))))))).. ( -18.47) >consensus __GCUGGGAA_AGCUGGGAAAAGGGAAAAUUCCUACCGGGCUCUUCACGUAAUGAAAAAAUAAGGGAAACCGAAAAUCCAUUGAAAUGAAGUUCACUGAAGCAUUUUUUGGU ..(((.........((((((.........)))))).(((...(((((..(((((.........((....)).......)))))...)))))....))).))).......... (-16.50 = -17.32 + 0.81)

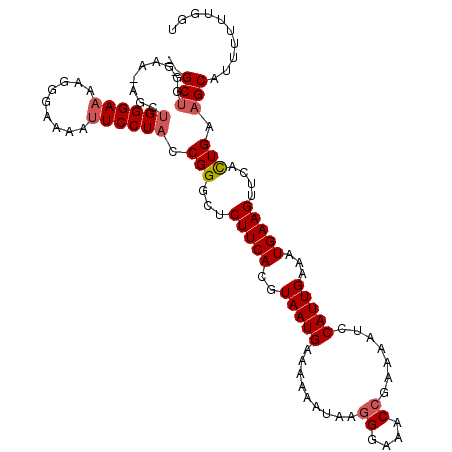
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:32:53 2006