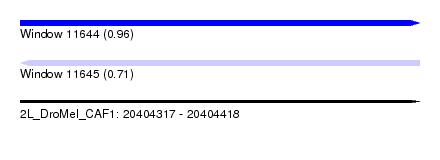

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 20,404,317 – 20,404,418 |
| Length | 101 |
| Max. P | 0.961885 |

| Location | 20,404,317 – 20,404,418 |
|---|---|
| Length | 101 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 89.59 |
| Mean single sequence MFE | -29.69 |
| Consensus MFE | -24.08 |
| Energy contribution | -23.80 |
| Covariance contribution | -0.28 |
| Combinations/Pair | 1.26 |
| Mean z-score | -2.10 |
| Structure conservation index | 0.81 |
| SVM decision value | 1.53 |
| SVM RNA-class probability | 0.961885 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 20404317 101 + 22407834 UUUUAUUUAUUCUGAACGAUCCGGCAGCCUGGCGGGAAAUAUUGAUAGUCGAUAGGCGAUGGGUCAUCGCACAAAAAGGGCUGCCAGAUUAUUCUGUUAAA .............(((..(((.((((((((.....(...(((((.....))))).((((((...)))))).).....)))))))).)))..)))....... ( -33.20) >DroSec_CAF1 55467 100 + 1 UUUUAUUUAUUCUAGACGAUCCGGCAGCCUGGCGGGAAAUAUUGAUAGUCGAUAGACGAUGGACUAUCGAA-AAAAAGGGCUGCCAGGUUAUUCUGCUUAA ............((((.((((.((((((((...........(((((((((.((.....)).))))))))).-.....)))))))).))))..))))..... ( -30.83) >DroSim_CAF1 56241 100 + 1 UUUUAUUUAAUCUAGACGAUCCGGCAGCCUGGCGGGAAAUAUUGAUAGUCGAUAGACGAUGGACUAUCGAA-AAAAAGGGCUGCCAGGUUAUUCUGUUUAA ............(((((((.((((((((((...........(((((((((.((.....)).))))))))).-.....)))))))).))....)).))))). ( -31.23) >DroEre_CAF1 54176 101 + 1 UAUUAUUUAUUCCAGACGAUCCGGCAGCCUGGCGGGAAAUAUUGAUAGUCGAUAGGCGAUGUUCCCUCGCAUAAAAAGAGUUGCCAGGUUAUUCUGUUUAA ............((((.((((.((((((((.(((((.(((((((.((.....))..))))))).)).)))......)).)))))).))))..))))..... ( -28.10) >DroYak_CAF1 55171 101 + 1 UGUUAUUUAUUCUAGACGAUCCGGCAGCCUGGCGGGAAAUAUUGAUAGUCGAUAGGUGAUGUUCCAUUGCAUAAAGAGGGUUGCCAGGUUAUUCUGUUUAA ............(((((((.((((((((((.((.((((.(((((.....))))).......))))...)).......)))))))).))....)).))))). ( -25.10) >consensus UUUUAUUUAUUCUAGACGAUCCGGCAGCCUGGCGGGAAAUAUUGAUAGUCGAUAGGCGAUGGACCAUCGCA_AAAAAGGGCUGCCAGGUUAUUCUGUUUAA ............((((.((((.((((((((.........(((((.....))))).((((((...)))))).......)))))))).))))..))))..... (-24.08 = -23.80 + -0.28)



| Location | 20,404,317 – 20,404,418 |
|---|---|
| Length | 101 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 89.59 |
| Mean single sequence MFE | -22.74 |
| Consensus MFE | -16.86 |
| Energy contribution | -17.50 |
| Covariance contribution | 0.64 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.86 |
| Structure conservation index | 0.74 |
| SVM decision value | 0.37 |
| SVM RNA-class probability | 0.708827 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 20404317 101 - 22407834 UUUAACAGAAUAAUCUGGCAGCCCUUUUUGUGCGAUGACCCAUCGCCUAUCGACUAUCAAUAUUUCCCGCCAGGCUGCCGGAUCGUUCAGAAUAAAUAAAA .......(((..(((((((((((........((((((...)))))).....(.....)..............)))))))))))..)))............. ( -31.30) >DroSec_CAF1 55467 100 - 1 UUAAGCAGAAUAACCUGGCAGCCCUUUUU-UUCGAUAGUCCAUCGUCUAUCGACUAUCAAUAUUUCCCGCCAGGCUGCCGGAUCGUCUAGAAUAAAUAAAA ......(((.....(((((((((......-.(((((((.(....).))))))).((....))..........)))))))))....)))............. ( -24.10) >DroSim_CAF1 56241 100 - 1 UUAAACAGAAUAACCUGGCAGCCCUUUUU-UUCGAUAGUCCAUCGUCUAUCGACUAUCAAUAUUUCCCGCCAGGCUGCCGGAUCGUCUAGAUUAAAUAAAA ((((.((((.....(((((((((......-.(((((((.(....).))))))).((....))..........)))))))))....))).).))))...... ( -24.20) >DroEre_CAF1 54176 101 - 1 UUAAACAGAAUAACCUGGCAACUCUUUUUAUGCGAGGGAACAUCGCCUAUCGACUAUCAAUAUUUCCCGCCAGGCUGCCGGAUCGUCUGGAAUAAAUAAUA .....((((.....((((((.(((((.......)))))......((((..((...............))..))))))))))....))))............ ( -22.16) >DroYak_CAF1 55171 101 - 1 UUAAACAGAAUAACCUGGCAACCCUCUUUAUGCAAUGGAACAUCACCUAUCGACUAUCAAUAUUUCCCGCCAGGCUGCCGGAUCGUCUAGAAUAAAUAACA ......(((.....((((((..((............)).......(((..((...............))..))).))))))....)))............. ( -11.96) >consensus UUAAACAGAAUAACCUGGCAGCCCUUUUU_UGCGAUGGACCAUCGCCUAUCGACUAUCAAUAUUUCCCGCCAGGCUGCCGGAUCGUCUAGAAUAAAUAAAA ......(((.....(((((((((..........(((((........)))))(.....)..............)))))))))....)))............. (-16.86 = -17.50 + 0.64)

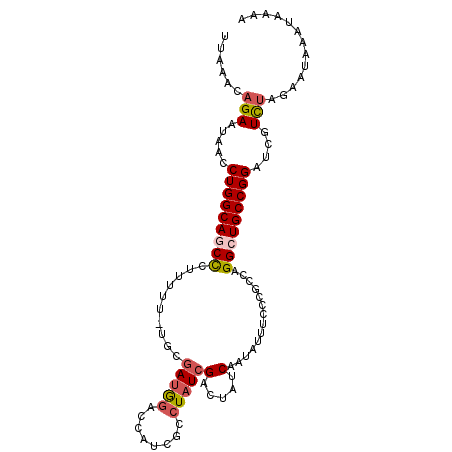

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:32:08 2006