
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 20,278,566 – 20,278,686 |
| Length | 120 |
| Max. P | 0.842082 |

| Location | 20,278,566 – 20,278,686 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 90.73 |
| Mean single sequence MFE | -37.19 |
| Consensus MFE | -28.65 |
| Energy contribution | -30.45 |
| Covariance contribution | 1.80 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.93 |
| Structure conservation index | 0.77 |
| SVM decision value | 0.75 |
| SVM RNA-class probability | 0.842082 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
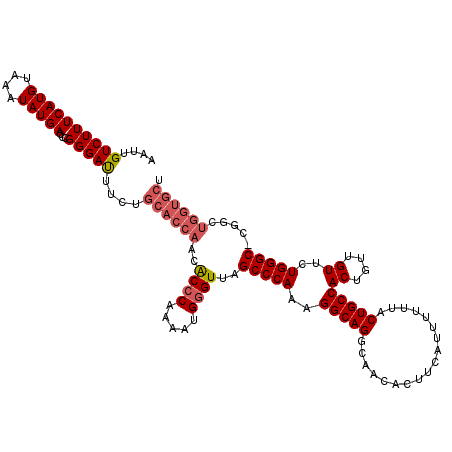
>2L_DroMel_CAF1 20278566 120 + 22407834 AGCACCAGCCCGGCCCAGAACAACAGUGGCAGUAAAAAAUGAAGUGUUGCCUGCCUUUGGGCUAACCCAUUUUGGGUGUUGGUGCAGAAAUCCCGGGUCAUAUUUACAUGAAAGACAAUU .((((((((..((((((((((....))((((((((...........))).))))))))))))).((((.....))))))))))))........(...((((......))))..)...... ( -38.30) >DroSec_CAF1 29400 119 + 1 AGCACCAGCCG-GCCCAGAACAACAGUGGCAGUAAAAAAUGAAGUGUGGCCUGCCUUUGGGCUAACCCAUUUUGGGUGUUGGUGCAGAAAUCCCGGGUCAUAUUUACAUGAAAGACAAUU .((((((((.(-(((((((((....))(((((..................))))))))))))).((((.....))))))))))))........(...((((......))))..)...... ( -37.87) >DroSim_CAF1 28261 120 + 1 AGCACCAGCCGGGCCCAGAACAACAGUGGCAGUAAAAAAUGAAGUGUUGCCUGCCUGUGGGCUAACCCAUUUUGGGUGUUGGUGCAGAAAUCCCGGGUCAUAUUUACAUGAAAGACAAUU .((((((((..((((((.(((....))((((((((...........))).)))))).)))))).((((.....))))))))))))........(...((((......))))..)...... ( -36.90) >DroEre_CAF1 14505 119 + 1 AGCACCAUCCC-GCCCAGAACAACAGUGGCAGUUAAAAAUGUAGUGCUGCCUGCCUUUGGGCUAACCCAUUCUGGGCGUUGGUGCAGAAGUCCCGGCUCAUAUUUACAUGAAAGACAAUU .((((((...(-((((((((.......((((((............))))))......((((....))))))))))))).))))))....(((.....((((......))))..))).... ( -43.70) >DroYak_CAF1 9827 109 + 1 ----------G-GCCCAGAACAACAGUGGCAGUAAAAAAUGUAGAGUUGCCUGCCUUUGGGCUAACACAUCCUGGGUGUUGGUGCGGAAAUCCCGGGUCAUAUUUACAUGAAAGACAAUU ----------(-((((...........((((((((...........))).)))))((((.((((((((.......)))))))).))))......)))))..................... ( -29.20) >consensus AGCACCAGCCG_GCCCAGAACAACAGUGGCAGUAAAAAAUGAAGUGUUGCCUGCCUUUGGGCUAACCCAUUUUGGGUGUUGGUGCAGAAAUCCCGGGUCAUAUUUACAUGAAAGACAAUU .((((((((...((((((((.......(((((..................)))))..((((....))))))))))))))))))))............((((......))))......... (-28.65 = -30.45 + 1.80)


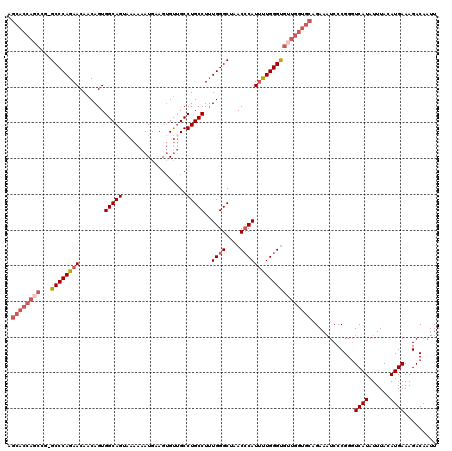
| Location | 20,278,566 – 20,278,686 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 90.73 |
| Mean single sequence MFE | -35.09 |
| Consensus MFE | -28.23 |
| Energy contribution | -29.31 |
| Covariance contribution | 1.08 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.67 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.65 |
| SVM RNA-class probability | 0.813092 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 20278566 120 - 22407834 AAUUGUCUUUCAUGUAAAUAUGACCCGGGAUUUCUGCACCAACACCCAAAAUGGGUUAGCCCAAAGGCAGGCAACACUUCAUUUUUUACUGCCACUGUUGUUCUGGGCCGGGCUGGUGCU ....((((((((((....)))))...)))))....(((((...((((.....)))).(((((...(((((..................))))).((........))...)))))))))). ( -34.47) >DroSec_CAF1 29400 119 - 1 AAUUGUCUUUCAUGUAAAUAUGACCCGGGAUUUCUGCACCAACACCCAAAAUGGGUUAGCCCAAAGGCAGGCCACACUUCAUUUUUUACUGCCACUGUUGUUCUGGGC-CGGCUGGUGCU ....((((((((((....)))))...)))))....((((((.(((((.....))))..(((((..(((((..................)))))((....))..)))))-..).)))))). ( -33.97) >DroSim_CAF1 28261 120 - 1 AAUUGUCUUUCAUGUAAAUAUGACCCGGGAUUUCUGCACCAACACCCAAAAUGGGUUAGCCCACAGGCAGGCAACACUUCAUUUUUUACUGCCACUGUUGUUCUGGGCCCGGCUGGUGCU ....((((((((((....)))))...)))))....((((((.(((((.....))))..(((((..(((((..................)))))((....))..)))))...).)))))). ( -33.67) >DroEre_CAF1 14505 119 - 1 AAUUGUCUUUCAUGUAAAUAUGAGCCGGGACUUCUGCACCAACGCCCAGAAUGGGUUAGCCCAAAGGCAGGCAGCACUACAUUUUUAACUGCCACUGUUGUUCUGGGC-GGGAUGGUGCU ....((((((((((....))))))...))))....((((((.(((((((((((((....)))...(((((..................)))))......)))))))))-)...)))))). ( -47.67) >DroYak_CAF1 9827 109 - 1 AAUUGUCUUUCAUGUAAAUAUGACCCGGGAUUUCCGCACCAACACCCAGGAUGUGUUAGCCCAAAGGCAGGCAACUCUACAUUUUUUACUGCCACUGUUGUUCUGGGC-C---------- .........(((((....)))))((((((.((..((((((........)).))))..)))))...(((((..................)))))...........))).-.---------- ( -25.67) >consensus AAUUGUCUUUCAUGUAAAUAUGACCCGGGAUUUCUGCACCAACACCCAAAAUGGGUUAGCCCAAAGGCAGGCAACACUUCAUUUUUUACUGCCACUGUUGUUCUGGGC_CGGCUGGUGCU ....((((((((((....)))))...)))))....((((((..((((.....))))..(((((..(((((..................)))))((....))..))))).....)))))). (-28.23 = -29.31 + 1.08)

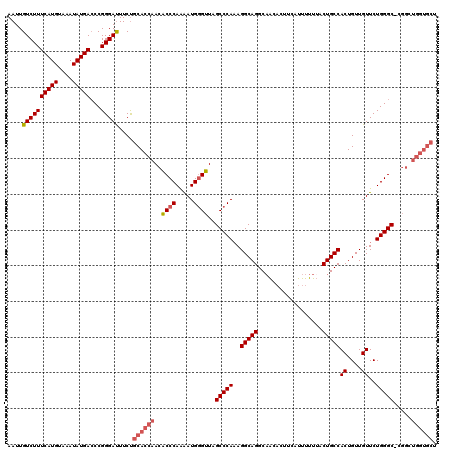
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:31:28 2006