
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 20,251,101 – 20,251,220 |
| Length | 119 |
| Max. P | 0.717785 |

| Location | 20,251,101 – 20,251,220 |
|---|---|
| Length | 119 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 88.00 |
| Mean single sequence MFE | -40.33 |
| Consensus MFE | -30.35 |
| Energy contribution | -31.80 |
| Covariance contribution | 1.45 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.38 |
| Structure conservation index | 0.75 |
| SVM decision value | 0.39 |
| SVM RNA-class probability | 0.717785 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
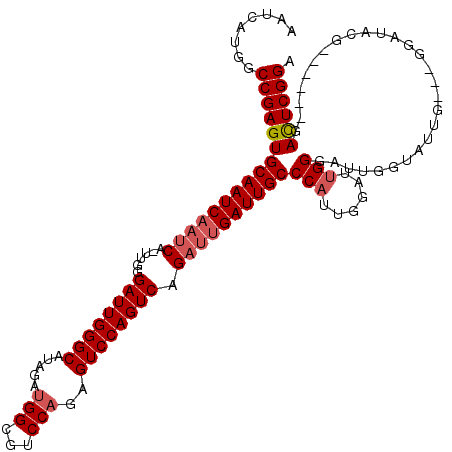
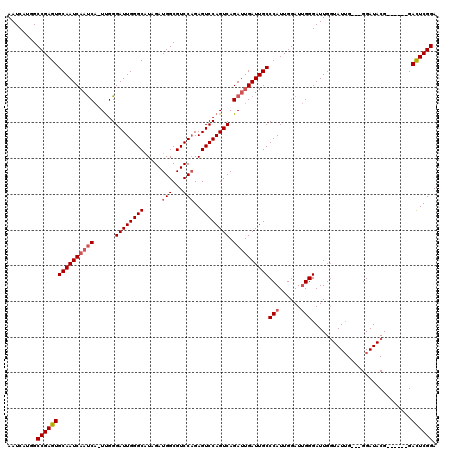
>2L_DroMel_CAF1 20251101 119 + 22407834 AAUCAUGGCCGAGUGCAAUCAAUCA-UUGGGAUUGGGCAUAGAUGGCGUCCAGAGUCCAGUCAGAUUGAUUGCCCAUUGGAUUGGGAUUGUUAUUGGAAGGAUACGGAUUCGGAUUCGGA ........(((((((((((((((((-(((((.((((((.........))))))..))))))..))))))))))...(((((((.(.(((.((.....)).))).).))))))))))))). ( -38.50) >DroSec_CAF1 12672 110 + 1 AAUCAUGGCCGAGUGCAAUCAAUCA-UUGGGAUUGGGCGUAGAUGGCGUCCAGAGUCCAGUCAGAUUGAUUGCCCAUUGGAUUGGGAUUGGUAUUG---GGAUACG------GACUCGGA ........(((((((((((((((((-(((((.((((((((.....))))))))..))))))..))))))))))(((......))).....((((..---..)))).------.)))))). ( -42.80) >DroSim_CAF1 11081 109 + 1 AAUCAUGGCCGAGUGCAAUCCUCCUCUCGGGAUUGGGCAUAGAUGGCGUCCCGAGUCCAGUCAGAUUGAUUGCCCAUUGGAU-GGGAUUGGUAU-G---GGAUACG------GACUCGGA .......((((.((.(((((((......))))))).)).....))))...((((((((.(((..((..((..(((((...))-)))))..))..-.---.)))..)------))))))). ( -39.70) >consensus AAUCAUGGCCGAGUGCAAUCAAUCA_UUGGGAUUGGGCAUAGAUGGCGUCCAGAGUCCAGUCAGAUUGAUUGCCCAUUGGAUUGGGAUUGGUAUUG___GGAUACG______GACUCGGA ........((((((((((((((((......((((((((.....(((...)))..)))))))).))))))))))(((......)))............................)))))). (-30.35 = -31.80 + 1.45)

| Location | 20,251,101 – 20,251,220 |
|---|---|
| Length | 119 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 88.00 |
| Mean single sequence MFE | -28.87 |
| Consensus MFE | -23.14 |
| Energy contribution | -24.37 |
| Covariance contribution | 1.23 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.12 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.33 |
| SVM RNA-class probability | 0.689998 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 20251101 119 - 22407834 UCCGAAUCCGAAUCCGUAUCCUUCCAAUAACAAUCCCAAUCCAAUGGGCAAUCAAUCUGACUGGACUCUGGACGCCAUCUAUGCCCAAUCCCAA-UGAUUGAUUGCACUCGGCCAUGAUU .......((((........................(((......)))((((((((((.....(((...(((..((.......))))).)))...-.))))))))))..))))........ ( -24.70) >DroSec_CAF1 12672 110 - 1 UCCGAGUC------CGUAUCC---CAAUACCAAUCCCAAUCCAAUGGGCAAUCAAUCUGACUGGACUCUGGACGCCAUCUACGCCCAAUCCCAA-UGAUUGAUUGCACUCGGCCAUGAUU .((((((.------.......---...........(((......)))((((((((((.....(((...(((.((.......)).))).)))...-.))))))))))))))))........ ( -30.20) >DroSim_CAF1 11081 109 - 1 UCCGAGUC------CGUAUCC---C-AUACCAAUCCC-AUCCAAUGGGCAAUCAAUCUGACUGGACUCGGGACGCCAUCUAUGCCCAAUCCCGAGAGGAGGAUUGCACUCGGCCAUGAUU .((((((.------((.((((---(-...(((..(((-((...)))))...((.....)).))).(((((((.((.......))....)))))))..).)))))).))))))........ ( -31.70) >consensus UCCGAGUC______CGUAUCC___CAAUACCAAUCCCAAUCCAAUGGGCAAUCAAUCUGACUGGACUCUGGACGCCAUCUAUGCCCAAUCCCAA_UGAUUGAUUGCACUCGGCCAUGAUU .((((((............................(((......)))((((((((((.....(((...(((.((.......)).))).))).....))))))))))))))))........ (-23.14 = -24.37 + 1.23)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:30:27 2006