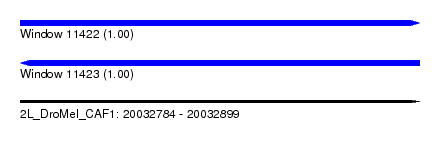

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 20,032,784 – 20,032,899 |
| Length | 115 |
| Max. P | 0.999962 |

| Location | 20,032,784 – 20,032,899 |
|---|---|
| Length | 115 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 94.80 |
| Mean single sequence MFE | -33.23 |
| Consensus MFE | -29.72 |
| Energy contribution | -29.61 |
| Covariance contribution | -0.11 |
| Combinations/Pair | 1.07 |
| Mean z-score | -3.33 |
| Structure conservation index | 0.89 |
| SVM decision value | 3.82 |
| SVM RNA-class probability | 0.999636 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
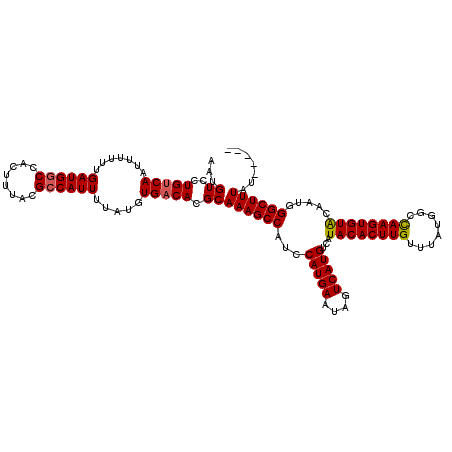
>2L_DroMel_CAF1 20032784 115 + 22407834 -----AUAAAGCCCAAUGUACACUUGGUCAUAAACAAGUGUAUGACAUGACUAUUCAUGGAUGGCUUUGCGUGUCACAUAAAAUGGCGUAAAGUGGCCAUCAAAAAAUUGACAGAACAUU -----.(((((((....(((((((((........)))))))))..(((((....)))))...)))))))..(((((......(((((........)))))........)))))....... ( -33.64) >DroSec_CAF1 215103 115 + 1 -----AUAAAGCCCAAUGUACACUUGGCCAUAAACAAGUGUAUGACAUGACUAUUCAUGGAUGGCUUUGCGUGUCACAUAAAAUGGCGUAAAGUGGCCAUCAAAAAAUUGACAGGACAUU -----.(((((((....(((((((((........)))))))))..(((((....)))))...)))))))(.(((((......(((((........)))))........))))).)..... ( -35.44) >DroSim_CAF1 212506 115 + 1 -----AUAAAGCCCAAUGUACACUUGGCCAUAAACAAGUGUAUGACAUGACUAUUCAUGGAUGGCUUUGCGUGUCACAUAAAAUGGCGUAAAGUGGCCAUCAAAAAAUUGACAGGACAUU -----.(((((((....(((((((((........)))))))))..(((((....)))))...)))))))(.(((((......(((((........)))))........))))).)..... ( -35.44) >DroEre_CAF1 209778 115 + 1 -----AUAAAGCCCAUUGCACACUUGGCCAUAAACAAGUGUAUGCCAUGACUAUUCAUGGAUGGCUUUGCGUGUCACAUAAAAUGGCGUAAAGUGGCCAUCAAAAAAUUGACAGGACAUU -----.(((((((......(((((((........)))))))...((((((....))))))..)))))))(.(((((......(((((........)))))........))))).)..... ( -35.64) >DroYak_CAF1 208575 115 + 1 -----AUAAAGCCCAUUGCACACUUGGCCAUAAACAAGUGUAUGACAUGACUAUUCAUGGAUGGCUUUGCGUGUCACAUAAAAUGGCGUAAAGUGGUCAUCAAAAAAUUGACAGGACAUU -----.(((((((.(((..(((((((........)))))))....(((((....)))))))))))))))(.(((((......(((((........)))))........))))).)..... ( -30.64) >DroAna_CAF1 225636 120 + 1 GGAAUUUAAAGCCCAUUGCACACUUAGCCAUAAACAAGUGUAUGACAUGACUAUUCAUGAAUGGCUUUGCGUGUCACAUAAAAUGGCGUAAAGUGGCCAUCAAAAAAUUGGCAGAACAUC ..........(((.(((((((.(..((((((..((....))....(((((....))))).))))))..).))))........(((((........))))).....))).)))........ ( -28.60) >consensus _____AUAAAGCCCAAUGCACACUUGGCCAUAAACAAGUGUAUGACAUGACUAUUCAUGGAUGGCUUUGCGUGUCACAUAAAAUGGCGUAAAGUGGCCAUCAAAAAAUUGACAGGACAUU ......(((((((......(((((((........)))))))....(((((....)))))...)))))))..(((((......(((((........)))))........)))))....... (-29.72 = -29.61 + -0.11)



| Location | 20,032,784 – 20,032,899 |
|---|---|
| Length | 115 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 94.80 |
| Mean single sequence MFE | -31.04 |
| Consensus MFE | -30.31 |
| Energy contribution | -30.25 |
| Covariance contribution | -0.06 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.61 |
| Structure conservation index | 0.98 |
| SVM decision value | 4.93 |
| SVM RNA-class probability | 0.999962 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 20032784 115 - 22407834 AAUGUUCUGUCAAUUUUUUGAUGGCCACUUUACGCCAUUUUAUGUGACACGCAAAGCCAUCCAUGAAUAGUCAUGUCAUACACUUGUUUAUGACCAAGUGUACAUUGGGCUUUAU----- ...((..(((((.......((((((........)))))).....))))).))((((((...(((((....)))))...((((((((........)))))))).....))))))..----- ( -32.90) >DroSec_CAF1 215103 115 - 1 AAUGUCCUGUCAAUUUUUUGAUGGCCACUUUACGCCAUUUUAUGUGACACGCAAAGCCAUCCAUGAAUAGUCAUGUCAUACACUUGUUUAUGGCCAAGUGUACAUUGGGCUUUAU----- ...((..(((((.......((((((........)))))).....))))).))((((((...(((((....)))))...((((((((........)))))))).....))))))..----- ( -32.30) >DroSim_CAF1 212506 115 - 1 AAUGUCCUGUCAAUUUUUUGAUGGCCACUUUACGCCAUUUUAUGUGACACGCAAAGCCAUCCAUGAAUAGUCAUGUCAUACACUUGUUUAUGGCCAAGUGUACAUUGGGCUUUAU----- ...((..(((((.......((((((........)))))).....))))).))((((((...(((((....)))))...((((((((........)))))))).....))))))..----- ( -32.30) >DroEre_CAF1 209778 115 - 1 AAUGUCCUGUCAAUUUUUUGAUGGCCACUUUACGCCAUUUUAUGUGACACGCAAAGCCAUCCAUGAAUAGUCAUGGCAUACACUUGUUUAUGGCCAAGUGUGCAAUGGGCUUUAU----- ...((..(((((.......((((((........)))))).....))))).))((((((..((((((....))))))..((((((((........)))))))).....))))))..----- ( -34.90) >DroYak_CAF1 208575 115 - 1 AAUGUCCUGUCAAUUUUUUGAUGACCACUUUACGCCAUUUUAUGUGACACGCAAAGCCAUCCAUGAAUAGUCAUGUCAUACACUUGUUUAUGGCCAAGUGUGCAAUGGGCUUUAU----- ...(((..(((((....))))))))(((...............)))......((((((...(((((....)))))...((((((((........)))))))).....))))))..----- ( -24.96) >DroAna_CAF1 225636 120 - 1 GAUGUUCUGCCAAUUUUUUGAUGGCCACUUUACGCCAUUUUAUGUGACACGCAAAGCCAUUCAUGAAUAGUCAUGUCAUACACUUGUUUAUGGCUAAGUGUGCAAUGGGCUUUAAAUUCC ........(((........((((((........)))))).....((.(((((..((((((.(((((....)))))..............))))))..)))))))...))).......... ( -28.86) >consensus AAUGUCCUGUCAAUUUUUUGAUGGCCACUUUACGCCAUUUUAUGUGACACGCAAAGCCAUCCAUGAAUAGUCAUGUCAUACACUUGUUUAUGGCCAAGUGUACAAUGGGCUUUAU_____ ...((..(((((.......((((((........)))))).....))))).))((((((...(((((....)))))...((((((((........)))))))).....))))))....... (-30.31 = -30.25 + -0.06)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:28:48 2006