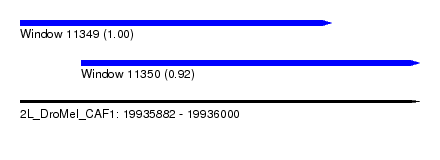

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 19,935,882 – 19,936,000 |
| Length | 118 |
| Max. P | 0.997949 |

| Location | 19,935,882 – 19,935,974 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 97 |
| Reading direction | forward |
| Mean pairwise identity | 87.24 |
| Mean single sequence MFE | -37.44 |
| Consensus MFE | -30.16 |
| Energy contribution | -30.68 |
| Covariance contribution | 0.52 |
| Combinations/Pair | 1.09 |
| Mean z-score | -3.16 |
| Structure conservation index | 0.81 |
| SVM decision value | 2.97 |
| SVM RNA-class probability | 0.997949 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
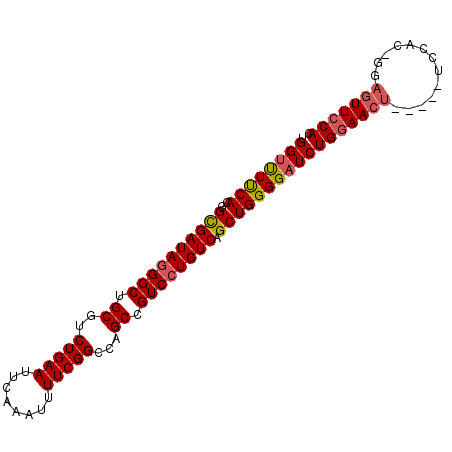
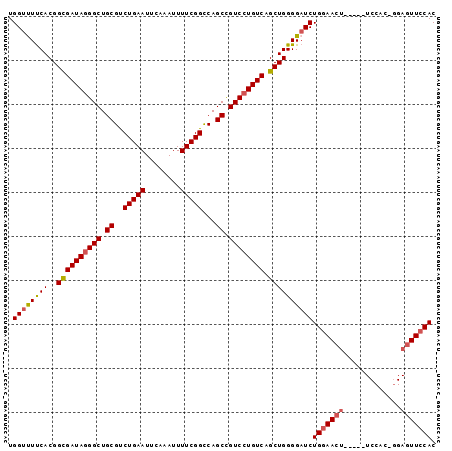
>2L_DroMel_CAF1 19935882 92 + 22407834 UGGUUUUCACGGUGAUAGGGCUGCGCCUGAAUUCAAAUUUUCGGCCAGCCGUCCUGUCAGCUGGGGAUCUGGAACU-----UCCACCGGAGUUCCAC .((((..((.(.(((((((((.(((((.(((........))))))..)).))))))))).)))..))))(((((((-----((....))))))))). ( -42.10) >DroSec_CAF1 111769 92 + 1 UGGUUUUCACGGCGAUAGGGCUGCGUCUGAAUUCAAAUUUUCGGCCAGCCGUCCUGUCAGCUGGGGAUCUGGAACU-----UCCACUGGAGUUCCAC .((((..((..((((((((((.((..(((((........)))))...)).)))))))).))))..))))(((((((-----((....))))))))). ( -39.10) >DroSim_CAF1 123397 92 + 1 UGGUUUUCACGGCGAUAGGGCUGCGUCUGAAUUCAAAUUUUCGGCCAGCCGUCCUGUCAGCUGGGGAUCUGGAACU-----UCCACUGGAGUUCCAC .((((..((..((((((((((.((..(((((........)))))...)).)))))))).))))..))))(((((((-----((....))))))))). ( -39.10) >DroEre_CAF1 117403 93 + 1 UGGUUUCCACGGCGAUAGGGCUGCGUCUGAAUUCAAAUUUUCGGUCAGCUGUCCUGUCAGCUGGGGUUCUGCAACUAGAGUUCC----GAGUUCCAC (((.(((...(((((((((((.((..(((((........)))))...)).)))))))).)))((..(((((....)))))..))----)))..))). ( -35.30) >DroYak_CAF1 117863 86 + 1 UGGUCUUCACGGCGAUAUGGCUGCGCCUGAAUUCAAAUUUUCGGUCAGCUGUCCUGUCAGCUGGGGAUCUGGAACUGGAG-----------UUCCAC .(((((((((((((..(((((((.(((.(((........)))))))))))))..)))).).))))))))((((((....)-----------))))). ( -31.60) >consensus UGGUUUUCACGGCGAUAGGGCUGCGUCUGAAUUCAAAUUUUCGGCCAGCCGUCCUGUCAGCUGGGGAUCUGGAACU_____UCCAC_GGAGUUCCAC .((((((((..((((((((((.((..(((((........)))))...)).)))))))).))))))))))(((((((.............))))))). (-30.16 = -30.68 + 0.52)

| Location | 19,935,900 – 19,936,000 |
|---|---|
| Length | 100 |
| Sequences | 5 |
| Columns | 105 |
| Reading direction | forward |
| Mean pairwise identity | 89.01 |
| Mean single sequence MFE | -35.62 |
| Consensus MFE | -27.68 |
| Energy contribution | -28.16 |
| Covariance contribution | 0.48 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.10 |
| Structure conservation index | 0.78 |
| SVM decision value | 1.12 |
| SVM RNA-class probability | 0.917794 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
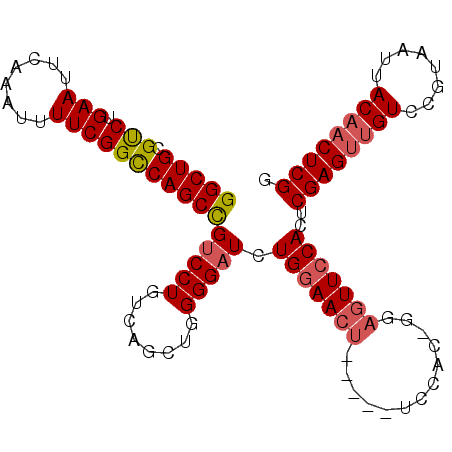
>2L_DroMel_CAF1 19935900 100 + 22407834 GGCUGCGCCUGAAUUCAAAUUUUCGGCCAGCCGUCCUGUCAGCUGGGGAUCUGGAACU-----UCCACCGGAGUUCCACUCGAGUUGUCCGUAAUUACAACUCGG (((((.(((.(((........)))))))))))(((((........))))).(((((((-----((....)))))))))..((((((((........)))))))). ( -41.30) >DroSec_CAF1 111787 100 + 1 GGCUGCGUCUGAAUUCAAAUUUUCGGCCAGCCGUCCUGUCAGCUGGGGAUCUGGAACU-----UCCACUGGAGUUCCACUCGAGUUGUCCGUAAUUACAACUCGG (((((.(.(((((........)))))))))))(((((........))))).(((((((-----((....)))))))))..((((((((........)))))))). ( -39.00) >DroSim_CAF1 123415 100 + 1 GGCUGCGUCUGAAUUCAAAUUUUCGGCCAGCCGUCCUGUCAGCUGGGGAUCUGGAACU-----UCCACUGGAGUUCCACUCGAGUUGUCCGUAAUUACAACUCGG (((((.(.(((((........)))))))))))(((((........))))).(((((((-----((....)))))))))..((((((((........)))))))). ( -39.00) >DroEre_CAF1 117421 101 + 1 GGCUGCGUCUGAAUUCAAAUUUUCGGUCAGCUGUCCUGUCAGCUGGGGUUCUGCAACUAGAGUUCC----GAGUUCCACUCGAGUGGUCCGUAAUUACAACUCGG ((.(((..(((((........)))))(((((((......)))))))......))).))......((----(((((((((....))))...((....))))))))) ( -28.90) >DroYak_CAF1 117881 94 + 1 GGCUGCGCCUGAAUUCAAAUUUUCGGUCAGCUGUCCUGUCAGCUGGGGAUCUGGAACUGGAG-----------UUCCACUCGAGCUGUCCGUAAUUACAACUCGG (((((.(((.(((........))))))))))).....(.(((((.(((...((((((....)-----------)))))))).))))).)................ ( -29.90) >consensus GGCUGCGUCUGAAUUCAAAUUUUCGGCCAGCCGUCCUGUCAGCUGGGGAUCUGGAACU_____UCCAC_GGAGUUCCACUCGAGUUGUCCGUAAUUACAACUCGG (((((.(((.(((........)))))))))))(((((........))))).(((((((.............)))))))..((((((((........)))))))). (-27.68 = -28.16 + 0.48)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:27:42 2006