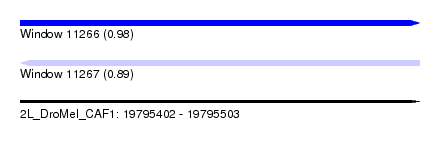

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 19,795,402 – 19,795,503 |
| Length | 101 |
| Max. P | 0.978482 |

| Location | 19,795,402 – 19,795,503 |
|---|---|
| Length | 101 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 88.16 |
| Mean single sequence MFE | -23.78 |
| Consensus MFE | -19.63 |
| Energy contribution | -20.23 |
| Covariance contribution | 0.60 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.21 |
| Structure conservation index | 0.83 |
| SVM decision value | 1.81 |
| SVM RNA-class probability | 0.978482 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 19795402 101 + 22407834 GCUGUCGGCCACAAUACCAGCGUUAUUGGGCAUCCAGUAUUAUCCUGCAUUAUCCUGUAUAUUCCACUUCGAUAAUACAGUCGUUGCACCCUUCGACACAA ..((((((((.(((((.......))))))))..............((((.....((((((..((......))..))))))....)))).....)))))... ( -22.40) >DroSec_CAF1 61915 101 + 1 GCUGUCGGCCACAAUACCAGCGUUAUUGGGCUUCCUGUAUUAUCCUGCAUUAUCCUGUAUAUUCCACUUCGAUAAUACAGUCGUUGCACCCUUCGACACAA ..((((((((.(((((.......))))))))...((((((((((.((((......))))...........)))))))))).............)))))... ( -22.60) >DroSim_CAF1 63199 91 + 1 GCUGUCGGCCACAAUACCAGCGUUAUUGGGCUUCCUGUAUUA----------UCCUGUAUAUUCCACUUCGAUAAUACAGUCGUUGCACCCUUCGACACAA ..((((((((.(((((.......))))))))...((((((((----------((..((.......))...)))))))))).............)))))... ( -23.50) >DroEre_CAF1 63331 89 + 1 GCUGUCGGCCACAAUACCAGCGUUAUUGGGCUUCCUGUAUUAUCCCG------------UAUUCCAGUGCGAUAAUACAGUCGUUGCACCCUUCGACACAA ..((((((((.(((((.......))))))))...((((((((((..(------------(((....)))))))))))))).............)))))... ( -25.60) >DroYak_CAF1 64360 89 + 1 GCUGUCGGCCACAAUACCAGCGAUAUUGGGCUUCCUGUAUUAUCCUG------------UAUUCCAGUUCGAUAAUACAGUCGUUGCACCCUUCGACACAA ..((((((((.(((((.......))))))))...(((((((((((((------------.....)))...)))))))))).............)))))... ( -24.80) >consensus GCUGUCGGCCACAAUACCAGCGUUAUUGGGCUUCCUGUAUUAUCCUG_____UCCUGUAUAUUCCACUUCGAUAAUACAGUCGUUGCACCCUUCGACACAA ..((((((((.(((((.......))))))))...((((((((((..........................)))))))))).............)))))... (-19.63 = -20.23 + 0.60)


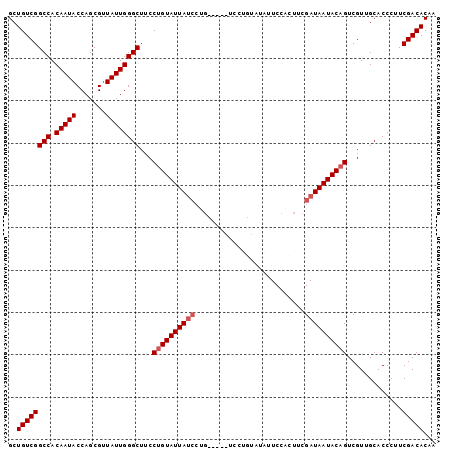
| Location | 19,795,402 – 19,795,503 |
|---|---|
| Length | 101 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 88.16 |
| Mean single sequence MFE | -28.54 |
| Consensus MFE | -22.23 |
| Energy contribution | -22.83 |
| Covariance contribution | 0.60 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.01 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.98 |
| SVM RNA-class probability | 0.894549 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 19795402 101 - 22407834 UUGUGUCGAAGGGUGCAACGACUGUAUUAUCGAAGUGGAAUAUACAGGAUAAUGCAGGAUAAUACUGGAUGCCCAAUAACGCUGGUAUUGUGGCCGACAGC (((..((....))..)))...((((((((((...(((.....)))..)))))))))).......(((...((((((((.......))))).)))...))). ( -29.20) >DroSec_CAF1 61915 101 - 1 UUGUGUCGAAGGGUGCAACGACUGUAUUAUCGAAGUGGAAUAUACAGGAUAAUGCAGGAUAAUACAGGAAGCCCAAUAACGCUGGUAUUGUGGCCGACAGC (((..((....))..)))((.((((((((((...(((.....)))..)))))))))).(((((((....(((........))).)))))))...))..... ( -27.30) >DroSim_CAF1 63199 91 - 1 UUGUGUCGAAGGGUGCAACGACUGUAUUAUCGAAGUGGAAUAUACAGGA----------UAAUACAGGAAGCCCAAUAACGCUGGUAUUGUGGCCGACAGC ...(((((..((((.......((((((((((...(((.....)))..))----------))))))))...)))).....(((.......)))..))))).. ( -26.90) >DroEre_CAF1 63331 89 - 1 UUGUGUCGAAGGGUGCAACGACUGUAUUAUCGCACUGGAAUA------------CGGGAUAAUACAGGAAGCCCAAUAACGCUGGUAUUGUGGCCGACAGC ...(((((..((((.......((((((((((...(((.....------------)))))))))))))...)))).....(((.......)))..))))).. ( -27.80) >DroYak_CAF1 64360 89 - 1 UUGUGUCGAAGGGUGCAACGACUGUAUUAUCGAACUGGAAUA------------CAGGAUAAUACAGGAAGCCCAAUAUCGCUGGUAUUGUGGCCGACAGC (((..((....))..)))((.((((((((((...(((.....------------)))))))))))))...(((((((((.....)))))).)))))..... ( -31.50) >consensus UUGUGUCGAAGGGUGCAACGACUGUAUUAUCGAAGUGGAAUAUACAGGA_____CAGGAUAAUACAGGAAGCCCAAUAACGCUGGUAUUGUGGCCGACAGC (((..((....))..)))((.((((((((((..........................))))))))))...((((((((.......))))).)))))..... (-22.23 = -22.83 + 0.60)

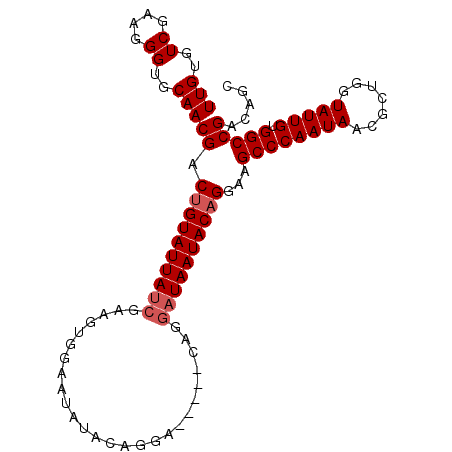

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:26:28 2006