
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 19,762,765 – 19,762,856 |
| Length | 91 |
| Max. P | 0.523327 |

| Location | 19,762,765 – 19,762,856 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 91 |
| Reading direction | forward |
| Mean pairwise identity | 82.75 |
| Mean single sequence MFE | -16.77 |
| Consensus MFE | -11.76 |
| Energy contribution | -11.04 |
| Covariance contribution | -0.72 |
| Combinations/Pair | 1.42 |
| Mean z-score | -1.54 |
| Structure conservation index | 0.70 |
| SVM decision value | -0.06 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 19762765 91 + 22407834 CCUGCUCGAAAUGCCUUUUGACAAUUUAAUUGCUAACUCAAAUAUUGCAUUGUCUAAAAGCUGGAAAACUUGAGCAACUCUUUUUCGCUCU ..(((((((....((((((((((((.((((.............)))).)))))).))))...)).....)))))))............... ( -15.22) >DroSec_CAF1 29095 91 + 1 UCUGCUCAAAAUGCCUUUUGACAAUUUAGUUGCUAACUCAAAUAUUUCAUUGUCUAAAAGUUGGAAAACUUGAGCAGCUCUUUGUCGUUCU ...((.......)).....(((((...(((((((....(((........))).....(((((....))))).)))))))..)))))..... ( -17.70) >DroSim_CAF1 28193 91 + 1 UCUGCUCAAAAUGCCUUUUGACAAUUUAGUUGCUAACUCAAAUAUUUCAUUGUCUAAAAGUUGGAAAACUUGAGCAGCUCUUUUUCUUUCU .((((((............((((((..(((.............)))..))))))...(((((....))))))))))).............. ( -17.62) >DroEre_CAF1 30582 91 + 1 UCUGUUCGAAAUGCCUUUUGACAAUUUAAUUACUAACUCAAAUAUUUCAUUGUUUCAAGGCUGGAGAACUUGAAUAGUUCUUUUUCAUUCU .......((((.(((((..((((((..(((.............)))..))))))..)))))..(((((((.....)))))))))))..... ( -18.82) >DroYak_CAF1 32618 88 + 1 UCCCUUUGAAAUGCCUUUUGGCAAUUUAAUUACUAACAAAAAGA---CAUUGUUUUAACGCUGGAGAACUCGAACAGAAAUUUGUUGUUCU (((..(((((.((((....)))).))))).....(((((.....---..)))))........)))......((((((.......)))))). ( -14.50) >consensus UCUGCUCGAAAUGCCUUUUGACAAUUUAAUUGCUAACUCAAAUAUUUCAUUGUCUAAAAGCUGGAAAACUUGAGCAGCUCUUUUUCGUUCU .((((((((....((....((((((..(((.............)))..))))))........)).....)))))))).............. (-11.76 = -11.04 + -0.72)

| Location | 19,762,765 – 19,762,856 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 91 |
| Reading direction | reverse |
| Mean pairwise identity | 82.75 |
| Mean single sequence MFE | -18.40 |
| Consensus MFE | -11.46 |
| Energy contribution | -12.98 |
| Covariance contribution | 1.52 |
| Combinations/Pair | 1.24 |
| Mean z-score | -1.76 |
| Structure conservation index | 0.62 |
| SVM decision value | -0.02 |
| SVM RNA-class probability | 0.523327 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
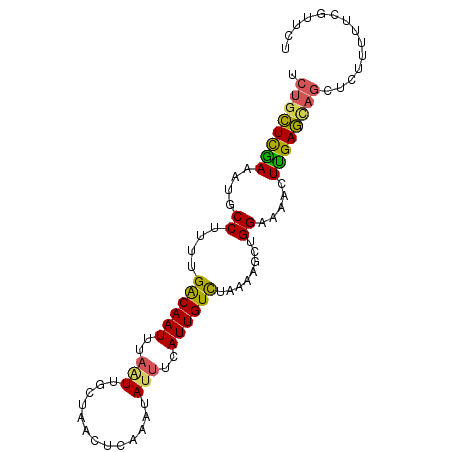
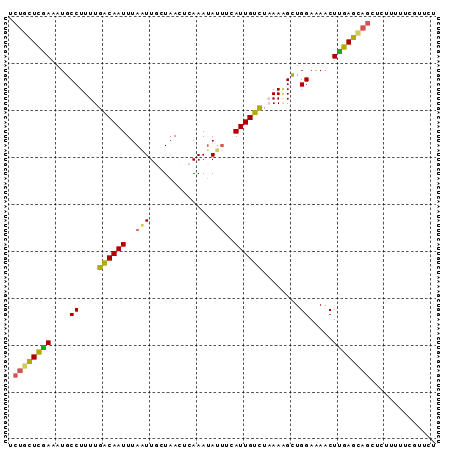
>2L_DroMel_CAF1 19762765 91 - 22407834 AGAGCGAAAAAGAGUUGCUCAAGUUUUCCAGCUUUUAGACAAUGCAAUAUUUGAGUUAGCAAUUAAAUUGUCAAAAGGCAUUUCGAGCAGG .((((((.......))))))..((((....((((((.((((((....((.(((......))).)).)))))).)))))).....))))... ( -20.50) >DroSec_CAF1 29095 91 - 1 AGAACGACAAAGAGCUGCUCAAGUUUUCCAACUUUUAGACAAUGAAAUAUUUGAGUUAGCAACUAAAUUGUCAAAAGGCAUUUUGAGCAGA ..............(((((((((........(((((.((((((....((.(((......))).)).)))))))))))....))))))))). ( -20.10) >DroSim_CAF1 28193 91 - 1 AGAAAGAAAAAGAGCUGCUCAAGUUUUCCAACUUUUAGACAAUGAAAUAUUUGAGUUAGCAACUAAAUUGUCAAAAGGCAUUUUGAGCAGA ..............(((((((((........(((((.((((((....((.(((......))).)).)))))))))))....))))))))). ( -20.10) >DroEre_CAF1 30582 91 - 1 AGAAUGAAAAAGAACUAUUCAAGUUCUCCAGCCUUGAAACAAUGAAAUAUUUGAGUUAGUAAUUAAAUUGUCAAAAGGCAUUUCGAACAGA ....(((((.((((((.....))))))...(((((...(((((....((((......)))).....)))))...))))).)))))...... ( -17.80) >DroYak_CAF1 32618 88 - 1 AGAACAACAAAUUUCUGUUCGAGUUCUCCAGCGUUAAAACAAUG---UCUUUUUGUUAGUAAUUAAAUUGCCAAAAGGCAUUUCAAAGGGA .(((((.........)))))......(((...((....))((((---((((((.((.(((......))))).)))))))))).....))). ( -13.50) >consensus AGAACGAAAAAGAGCUGCUCAAGUUUUCCAGCUUUUAGACAAUGAAAUAUUUGAGUUAGCAAUUAAAUUGUCAAAAGGCAUUUCGAGCAGA ..............(((((((((.......((((((.((((((.......................)))))).))))))..))))))))). (-11.46 = -12.98 + 1.52)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:26:19 2006