
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 19,731,732 – 19,731,913 |
| Length | 181 |
| Max. P | 0.997461 |

| Location | 19,731,732 – 19,731,840 |
|---|---|
| Length | 108 |
| Sequences | 6 |
| Columns | 115 |
| Reading direction | forward |
| Mean pairwise identity | 70.47 |
| Mean single sequence MFE | -24.92 |
| Consensus MFE | -7.26 |
| Energy contribution | -9.52 |
| Covariance contribution | 2.25 |
| Combinations/Pair | 1.24 |
| Mean z-score | -2.03 |
| Structure conservation index | 0.29 |
| SVM decision value | 0.14 |
| SVM RNA-class probability | 0.604866 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
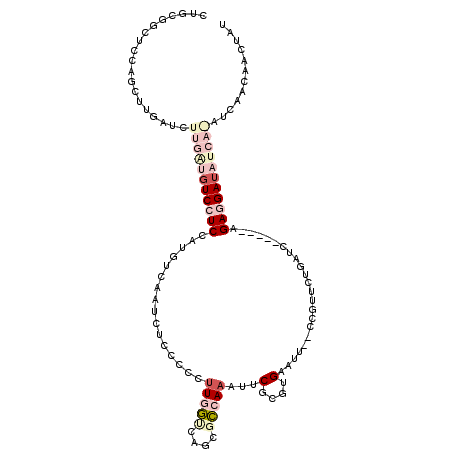
>2L_DroMel_CAF1 19731732 108 + 22407834 CUGCGGCUCCAUCUUGAUCUUGAUGUCCUCCAUGUCAAUUUCCCCCUUGGCCAGCGCCAAAUUCGCGAGAAUU--CCGUUCUGAUC-----AGAGGAUAUCAGAUCAACAACUAU .............(((((((.(((((((((...(((((........)))))(((.((..(((((....)))))--..)).)))...-----.))))))))))))))))....... ( -34.00) >DroPse_CAF1 5529 98 + 1 UUCCCCCUCCAGCUUGAUCGUGGUGUCAUCAAUGUCGAUUUCCUCUUUAAUUAUACUCAAAUCCACAUGCAUUUUCCGUUCUUUUUGCUUUUGAGGAU----------------- .....((((.(((..((.((.(((((.....((((.(((((.................))))).)))))))))...)).)).....)))...))))..----------------- ( -14.23) >DroSec_CAF1 2920 108 + 1 CUGCGGCUCCAACUUGAUCUUGAUGUCCUCCAUGUCAAUCUCACCCUUGGUCAGCGCCAAAUUCGAGUGAAUU--CCGUUCUGAUC-----AGAGGAUAUCAAAUCAACAACUAU .............(((((.(((((((((((..((.......))....(((((((.((..(((((....)))))--..)).))))))-----)))))))))))))))))....... ( -29.70) >DroSim_CAF1 2892 108 + 1 CUGCGGCUCCAACUUGAUCUUGAUGUCCUCCAUGUCAAUCUCACCCGUGGUCAGCGCCAAAUUCGCGUGAAUU--CCGUUCUGAUC-----AGAGGAUAUCAAAUCAACAACUAU .............(((((.(((((((((((..((.......))....(((((((.((..(((((....)))))--..)).))))))-----)))))))))))))))))....... ( -29.70) >DroYak_CAF1 4129 108 + 1 CUGCGGCUCUAGCUUGAUCUUGGUGUCCUCCAUGUCUAUCUCUCCCUUGGCCAGCGCCAAAUUCGCCUGCAUU--UCGGUCCGAUC-----AGAGGAAAUCAGAUCAACAACUAU ....(((....)))((((((.(((.(((((...(((..........(((((....)))))....(((......--..)))..))).-----.))))).)))))))))........ ( -26.40) >DroPer_CAF1 6053 98 + 1 UUCCCCCUCCAGCUUGAUCGUGGUGUCAUCAAUGUCGAUUUCCUCUUUAAUUAUACUCAAAUCCACAUGCAUUUCCCGUUCUUUUUGCUUUUGAGGAU----------------- .....((((.(((..((.((.((........((((.(((((.................))))).))))......)))).)).....)))...))))..----------------- ( -15.47) >consensus CUGCGGCUCCAGCUUGAUCUUGAUGUCCUCCAUGUCAAUCUCCCCCUUGGUCAGCGCCAAAUUCGCGUGAAUU__CCGUUCUGAUC_____AGAGGAUAUCA_AUCAACAACUAU ...................(((((((((((................(((((....)))))...(....).......................)))))))))))............ ( -7.26 = -9.52 + 2.25)


| Location | 19,731,805 – 19,731,913 |
|---|---|
| Length | 108 |
| Sequences | 5 |
| Columns | 111 |
| Reading direction | reverse |
| Mean pairwise identity | 93.13 |
| Mean single sequence MFE | -29.48 |
| Consensus MFE | -28.39 |
| Energy contribution | -28.19 |
| Covariance contribution | -0.20 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.79 |
| Structure conservation index | 0.96 |
| SVM decision value | 2.86 |
| SVM RNA-class probability | 0.997461 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 19731805 108 - 22407834 UAUCGAUAGUCGUCUUCUGAUCGCAGCGGUAUACAGCGGUAUUUAUUGCGCUGCUGCUCUCU---UCUUUAUAUUGAUAGUUGUUGAUCUGAUAUCCUCUGAUCAGAACGG ((((((((...(((....))).((((((((.....(((((....))))))))))))).....---......))))))))(((.((((((.((.....)).))))))))).. ( -29.70) >DroSec_CAF1 2993 108 - 1 UAUCGAUAUACGUCUUCUGAUCGCAGCGGUAUACAGCGGUAUUUAUUGCGCUGCUGCUCUCU---UUUAUAUAUUGAUAGUUGUUGAUUUGAUAUCCUCUGAUCAGAACGG ((((((((((.(((....))).((((((((.....(((((....))))))))))))).....---....))))))))))(((.((((((.((.....)).))))))))).. ( -30.00) >DroSim_CAF1 2965 108 - 1 UAUCGAUAGUCGUCUUCUGAUCGCAGCGGUAUACAGCGGUAUUUAUUGCGCUGCUGCUCUCU---UUUAUAUAUUGAUAGUUGUUGAUUUGAUAUCCUCUGAUCAGAACGG ..........(((..(((((((((((((((.....(((((....))))))))))))).....---..(((..((..((....))..))...)))......)))))))))). ( -28.50) >DroEre_CAF1 4209 111 - 1 UAUCGAUAGUCGUCUUCUGAUCGCAGCGGUAUACAGCGGUAUUUAUUGCGCUGCUGCUCUCUUCUUCUUGAUAUUGAUAGUUGUUGAUCUGAUAUCCUCUGAUCGGAGCGA .........((((..(((((((((((((((.....(((((....)))))))))))))............(((((((((........))).))))))....))))))))))) ( -31.80) >DroYak_CAF1 4202 108 - 1 UUUCGAUAGUCCUCUUCUGAUCGCAGCGGUAUACAGCGGUAUUUAUUGCGCUGCUGCUCUCU---UCUUGAUAUUGAUAGUUGUUGAUCUGAUUUCCUCUGAUCGGACCGA ..(((...((((......((((((((((((.....(((((....))))))))))))).....---....((((........)))))))).((((......))))))))))) ( -27.40) >consensus UAUCGAUAGUCGUCUUCUGAUCGCAGCGGUAUACAGCGGUAUUUAUUGCGCUGCUGCUCUCU___UCUUUAUAUUGAUAGUUGUUGAUCUGAUAUCCUCUGAUCAGAACGG ((((((((..........((..((((((((.....(((((....)))))))))))))..))..........))))))))(((.((((((.((.....)).))))))))).. (-28.39 = -28.19 + -0.20)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:26:15 2006