
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 19,598,524 – 19,598,615 |
| Length | 91 |
| Max. P | 0.652040 |

| Location | 19,598,524 – 19,598,615 |
|---|---|
| Length | 91 |
| Sequences | 6 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 79.96 |
| Mean single sequence MFE | -25.72 |
| Consensus MFE | -16.58 |
| Energy contribution | -17.25 |
| Covariance contribution | 0.67 |
| Combinations/Pair | 1.13 |
| Mean z-score | -1.54 |
| Structure conservation index | 0.64 |
| SVM decision value | 0.25 |
| SVM RNA-class probability | 0.652040 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
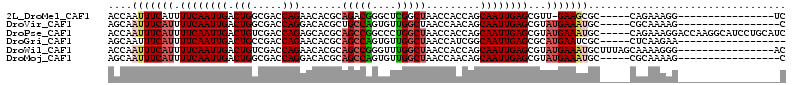
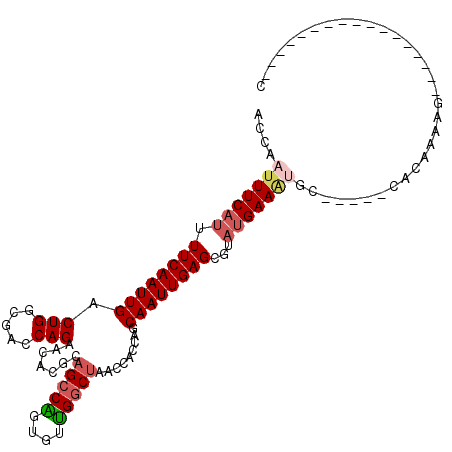
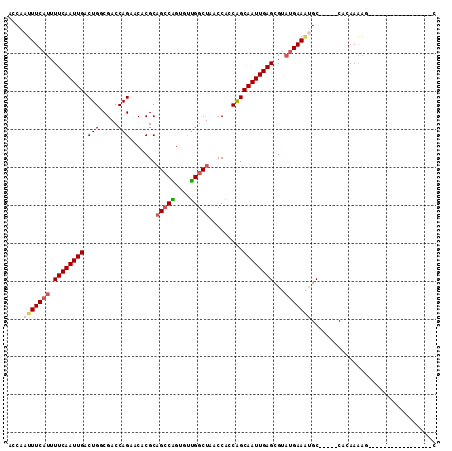
>2L_DroMel_CAF1 19598524 91 - 22407834 ACCAAUUUCAUUUUCAAUUGACUGGCGACCAGAACACGCAGACGGGCUCGGCUAACCACCAGCAAUUGAGCGUU-GAAGCGC-----CAGAAAGG----------------UC (((..((((....(((((((.((((............((......))..((....)).)))))))))))((((.-...))))-----..))))))----------------). ( -22.70) >DroVir_CAF1 25046 91 - 1 AGCAAUUUCAUUUUCAAUUGACUGGCGACCAGGACACGCUGCCAGUGUUGGCUAACCAACAGCAAUUGAGCGUAUGAAAUGC-----CGCAAAAG-----------------C .((.(((((((.(((((((((((((((.(........).)))))))(((((....)))))..))))))))...)))))))..-----.)).....-----------------. ( -29.90) >DroPse_CAF1 26537 108 - 1 ACCAAUUUCAUUUUCAAUUGACUGUCGACCAGAGCACGCAGCCGGCCCUGGCUAACCACCAGCAAUUGAGCGUAUGAAAUGC-----CAGAAAGGACCAAGGCAUCCUGCAUC ....(((((((.((((((((.(((..(.((((.((.((....)))).)))).....)..)))))))))))...)))))))((-----(............))).......... ( -25.80) >DroGri_CAF1 26156 90 - 1 AGCAAUUUCAUUUUCAAUUGACUGCCGACCAGAACACGCAGCCAGUGUUGGCUAACCAUCGGCAAUUGAGCGCAUGAAUCGC-----CUCAAGAA------------------ ..(((((........)))))..((((((....(((((.......)))))((....)).)))))).(((((.((.......))-----)))))...------------------ ( -21.90) >DroWil_CAF1 36870 97 - 1 ACCAAUUUCAUUUUCAAUUGACUGUCGACCAGAACACGCAGCCGGGUUUGGCUAACCACCAGCAAUUGAGCGUAUGAAAUGCUUUAGCAAAAGGG----------------AC .((.(((((((.((((((((.(((((.....))......(((((....)))))......)))))))))))...)))))))((....))....)).----------------.. ( -24.00) >DroMoj_CAF1 33620 91 - 1 AGCAAUUUCAUUUUCAAUUGACUGGCGACCAGGACACGCAGCCAGUGUUGGCUAACCAACAGCAAUUGAGCGUAUGAAAUGC-----CGCAAAAG-----------------C .((.(((((((.((((((((((((((..(........)..))))))(((((....)))))..))))))))...)))))))..-----.)).....-----------------. ( -30.00) >consensus ACCAAUUUCAUUUUCAAUUGACUGGCGACCAGAACACGCAGCCAGUGUUGGCUAACCACCAGCAAUUGAGCGUAUGAAAUGC_____CACAAAAG_________________C ....(((((((.((((((((.(((.....))).......(((((....))))).........))))))))...)))))))................................. (-16.58 = -17.25 + 0.67)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:25:34 2006