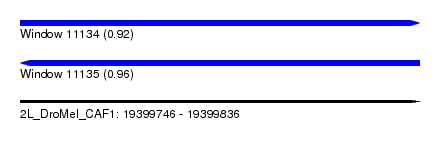
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 19,399,746 – 19,399,836 |
| Length | 90 |
| Max. P | 0.955988 |

| Location | 19,399,746 – 19,399,836 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 74.34 |
| Mean single sequence MFE | -18.02 |
| Consensus MFE | -8.34 |
| Energy contribution | -8.38 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.46 |
| SVM decision value | 1.15 |
| SVM RNA-class probability | 0.922864 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 19399746 90 + 22407834 ACAUUUAGAUUCCGAAUGCUAAAUGAAUGCUAGAUGUCUAGUC-----AAGAUGUGUAAAAUUCAUAUAGAGGAAUCAAAUGUGUCAGUGGCAAA ((((((.((((((.....(((.((((((..((.((((((....-----.)))))).))..)))))).))).))))))))))))((.....))... ( -25.60) >DroSec_CAF1 2743 83 + 1 ACAUUUCGAUUCCGAAUGGUAAAUGAAG-------GUCUAGUC-----UAGAUGUGUGAAAUGCAUAUAGAGGAAUCAAAUGUGUCAGUGGCAAA ((((((.((((((((.(((.........-------..))).))-----...((((((.....))))))...))))))))))))((.....))... ( -17.40) >DroSim_CAF1 2742 83 + 1 ACAUUUCGAUUCCGAAUGGUAAAUGAAG-------GUGUAGUC-----UAGAGGUGUGAAAUACAUAUAGAGGAAUCAAAUGUGUCAGUGGCAAA ((((((.((((((..((.....))....-------......((-----((...(((((...))))).))))))))))))))))((.....))... ( -15.70) >DroEre_CAF1 2685 73 + 1 GCAUUUAGAUUCCCAAUGGUAA-UAGAG-------GUAUAAAU---------UA-AUUAAAUCCAUUUAAAGGAAUCAAAUGUGCCAGUAG---- ((((((.((((((.((((((((-(.((.-------.......)---------).-))))...)))))....))))))))))))........---- ( -14.20) >DroYak_CAF1 2698 87 + 1 ACAUUUAGAUUCCCAAUGGUAUGUGUAA-------GUAUAGAUAGGCUUAGAUA-UAGAAACCCAUAUAAAAGAAUCAAAUGUGUCAGUGGCAAA ((((((.(((((...(((((((((.(((-------((........))))).)))-)).....))))......)))))))))))((.....))... ( -17.20) >consensus ACAUUUAGAUUCCGAAUGGUAAAUGAAG_______GUAUAGUC_____UAGAUGUGUGAAAUCCAUAUAGAGGAAUCAAAUGUGUCAGUGGCAAA ((((((.((((((..(((.............................................))).....))))))))))))............ ( -8.34 = -8.38 + 0.04)


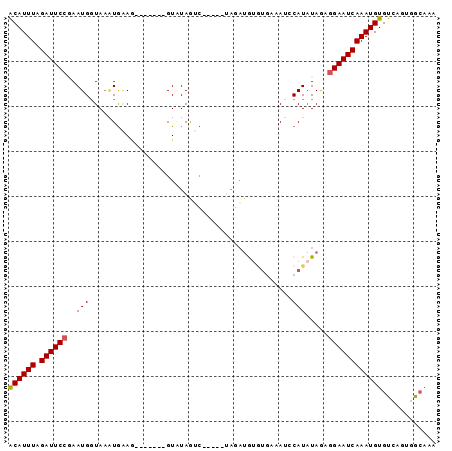
| Location | 19,399,746 – 19,399,836 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 74.34 |
| Mean single sequence MFE | -13.80 |
| Consensus MFE | -7.54 |
| Energy contribution | -7.58 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.09 |
| Structure conservation index | 0.55 |
| SVM decision value | 1.46 |
| SVM RNA-class probability | 0.955988 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 19399746 90 - 22407834 UUUGCCACUGACACAUUUGAUUCCUCUAUAUGAAUUUUACACAUCUU-----GACUAGACAUCUAGCAUUCAUUUAGCAUUCGGAAUCUAAAUGU ............((((((((((((.(((.((((((....((.....)-----).((((....)))).)))))).))).....)))))).)))))) ( -17.40) >DroSec_CAF1 2743 83 - 1 UUUGCCACUGACACAUUUGAUUCCUCUAUAUGCAUUUCACACAUCUA-----GACUAGAC-------CUUCAUUUACCAUUCGGAAUCGAAAUGU ............((((((((((((((((.(((.........))).))-----))...(..-------...)...........)))))).)))))) ( -12.80) >DroSim_CAF1 2742 83 - 1 UUUGCCACUGACACAUUUGAUUCCUCUAUAUGUAUUUCACACCUCUA-----GACUACAC-------CUUCAUUUACCAUUCGGAAUCGAAAUGU ............((((((((((((......(((.....)))......-----((......-------..))...........)))))).)))))) ( -10.50) >DroEre_CAF1 2685 73 - 1 ----CUACUGGCACAUUUGAUUCCUUUAAAUGGAUUUAAU-UA---------AUUUAUAC-------CUCUA-UUACCAUUGGGAAUCUAAAUGC ----.........((((((((((((...((((((..(((.-..---------..)))...-------.))))-))......))))))).))))). ( -13.30) >DroYak_CAF1 2698 87 - 1 UUUGCCACUGACACAUUUGAUUCUUUUAUAUGGGUUUCUA-UAUCUAAGCCUAUCUAUAC-------UUACACAUACCAUUGGGAAUCUAAAUGU ............((((((((((((..((.((((((((...-.....)))))))).))...-------.....((......)))))))).)))))) ( -15.00) >consensus UUUGCCACUGACACAUUUGAUUCCUCUAUAUGGAUUUCACACAUCUA_____GACUAGAC_______CUUCAUUUACCAUUCGGAAUCUAAAUGU ............((((((((((((..........................................................)))))).)))))) ( -7.54 = -7.58 + 0.04)


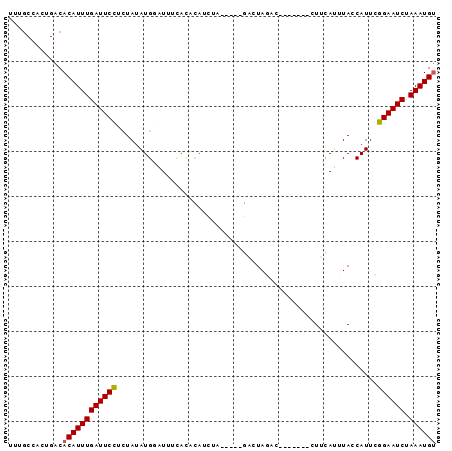
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:24:29 2006