
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 2,015,486 – 2,015,576 |
| Length | 90 |
| Max. P | 0.895325 |

| Location | 2,015,486 – 2,015,576 |
|---|---|
| Length | 90 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 78.18 |
| Mean single sequence MFE | -26.05 |
| Consensus MFE | -18.17 |
| Energy contribution | -17.45 |
| Covariance contribution | -0.72 |
| Combinations/Pair | 1.26 |
| Mean z-score | -1.69 |
| Structure conservation index | 0.70 |
| SVM decision value | 0.99 |
| SVM RNA-class probability | 0.895325 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
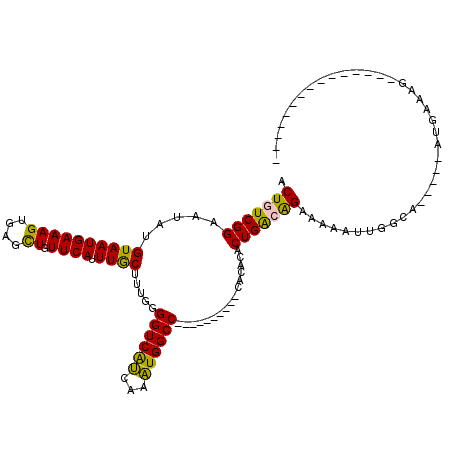
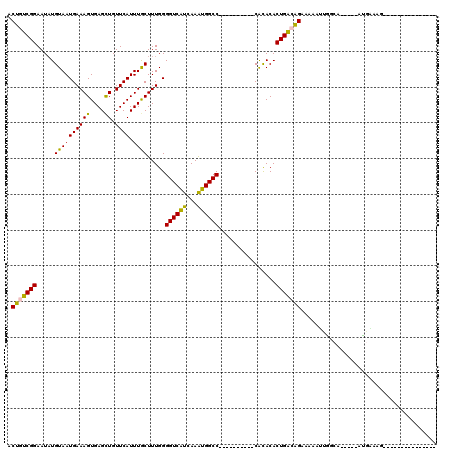
>2L_DroMel_CAF1 2015486 90 - 22407834 ACUGUCGGAAUAUGUAAUGAAAGUGAGCUGUUCAUUUGCUUUGGGGUCAUCAAAUGGCC----------CACACACUGAGAGAAAAAUUGGCA-----UUGAAAU--------------- .((.((((.....((((((((((....)).)))).))))..(((((((((...))))))----------).))..)))).))...........-----.......--------------- ( -20.50) >DroSec_CAF1 32940 90 - 1 ACUGUCGGAAUAUGUAAUGAAAGUGAGUUGUUCAUUUGCUUUGGGGUCAUCAAAUGGCC----------CACACACUGACAGAAAAAUUGGCA-----UUAAAAG--------------- .(((((((...........((((..(((.....)))..)))).(((((((...))))))----------).....)))))))...........-----.......--------------- ( -25.10) >DroSim_CAF1 31418 90 - 1 ACUGUCGGAAUAUGUAAUGAAAGUGAGCUGUUCAUUUGCUUUGGGGUCAUCAAAUGGCC----------CACACACUGACAGAAAAAUUAGCA-----UUGAAAG--------------- .(((((((.....((((((((((....)).)))).))))..(((((((((...))))))----------).))..)))))))...........-----.......--------------- ( -24.40) >DroEre_CAF1 30332 105 - 1 ACUGUCGGAAUAUGUAAUGAAAGUGAGCUGUUCAUUUGCUUUGGGGUCAUCAAAUGGCC----------CACACACUGGCAGAAAAAUUGGCA-----AUGGUUGGUUAUGAUGAUCAAG .(((((((.....((((((((((....)).)))).))))..(((((((((...))))))----------).))..)))))))...(((..((.-----...))..)))............ ( -27.80) >DroYak_CAF1 34545 105 - 1 ACUGUCGGAAUAUGUAAUGAAAGUGAGCUGUUCAUUUGCUUUGGGGUCAUCAAAUGGCC----------CGCACACUGGCAGAAAAACUGGCAGGCAGACGGUUGGU-----UGAUUAAA ((((((.(((((.((.((....))..)))))))...(((..(((((((((...))))))----------).))..(((.(((.....))).))))))))))))....-----........ ( -31.30) >DroAna_CAF1 30291 98 - 1 ACUGUCGGAAUUUGUAAUGAAAGUGAGCUGUUCAUUUACUUUGGGGUCGCCAAGUGGCCCCACACAUCCCAUACACUGAGGGAAAUCUCGUUA-----AUU--AG--------------- .((.((((....((..(((((((((((.......))))))))((((((((...))))))))...)))..))....)))).))...........-----...--..--------------- ( -27.20) >consensus ACUGUCGGAAUAUGUAAUGAAAGUGAGCUGUUCAUUUGCUUUGGGGUCAUCAAAUGGCC__________CACACACUGACAGAAAAAUUGGCA_____AUGAAAG_______________ .(((((((.....((((((((((....)).)))).)))).....((((((...))))))................)))))))...................................... (-18.17 = -17.45 + -0.72)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:43:12 2006