
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 19,353,941 – 19,354,034 |
| Length | 93 |
| Max. P | 0.747852 |

| Location | 19,353,941 – 19,354,034 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 93 |
| Reading direction | forward |
| Mean pairwise identity | 79.35 |
| Mean single sequence MFE | -21.00 |
| Consensus MFE | -15.24 |
| Energy contribution | -15.84 |
| Covariance contribution | 0.60 |
| Combinations/Pair | 1.13 |
| Mean z-score | -1.63 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.47 |
| SVM RNA-class probability | 0.747852 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
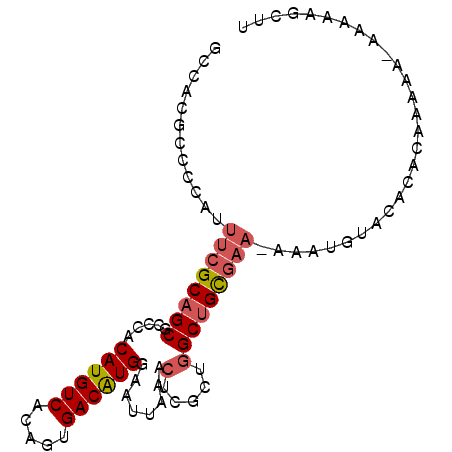
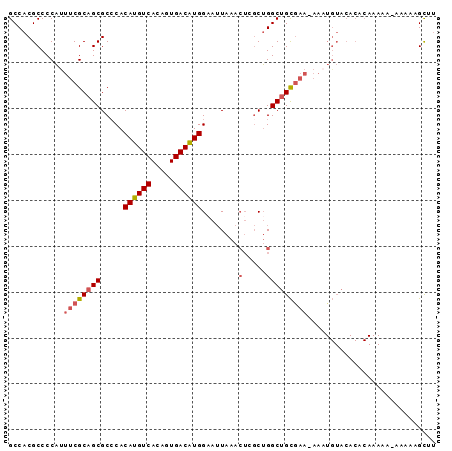
>2L_DroMel_CAF1 19353941 93 + 22407834 GCCACGCCCCAUUUCGCAGCGCACACAUGUCACAGUGACGUGGUAUUAAACUGGCUGGCUGCAAAAAAAUGUACACACAAAAAAAUAAAGUUU ((((.(((.......((...))((.((((((.....))))))))........)))))))((((......)))).................... ( -19.40) >DroSec_CAF1 17013 90 + 1 GCCACGCCCCAUUUCGCAGCGCCCACAUGUCACAGUGACAUGGAAUUAAACUCGCUGGCUGCGAA-AAAUGUACACUCAAAAA--AAAAGCUU .....((....(((((((((((...((((((.....))))))((.......))))..))))))))-)..((......))....--....)).. ( -22.00) >DroSim_CAF1 17238 92 + 1 GCCACGCCCCAUUUCGCAGCGCCCACAUGUCACAGUGACAUGGAAUUAAACUGGCUGGCUGCGAA-AAAUGUACACUCAAAAAAAAAAAGCUU .....((....((((((((((((..((((((.....))))))..........)))..))))))))-)..((......))..........)).. ( -23.70) >DroYak_CAF1 19881 91 + 1 GCCACGCCCCACUUCGCAGCGCCCACAUGUCACAGUGACGUGGAAUUAAACUCGCUGGCUGCGAA-AAUUGUACACACAAAAA-AGACUGCUU .....((.....((((((((((......))..((((((.((........))))))))))))))))-..((((....))))...-.....)).. ( -22.20) >DroAna_CAF1 16136 71 + 1 GCCACGCCCCUUAACGCAGCGCCCACAUGUCACAGUGACAUGGAAAUAC-----CAGGCAGUGGA-A--------------GA--AAAAGUUA .((((((........))...(((..((((((.....))))))(......-----).))).)))).-.--------------..--........ ( -17.70) >consensus GCCACGCCCCAUUUCGCAGCGCCCACAUGUCACAGUGACAUGGAAUUAAACUCGCUGGCUGCGAA_AAAUGUACACACAAAAA_AAAAAGCUU ............((((((((.....((((((.....))))))........(.....)))))))))............................ (-15.24 = -15.84 + 0.60)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:24:11 2006