| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 19,318,749 – 19,318,869 |
| Length | 120 |
| Max. P | 0.896451 |

| Location | 19,318,749 – 19,318,869 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 86.18 |
| Mean single sequence MFE | -39.88 |
| Consensus MFE | -28.26 |
| Energy contribution | -29.10 |
| Covariance contribution | 0.84 |
| Combinations/Pair | 1.16 |
| Mean z-score | -2.40 |
| Structure conservation index | 0.71 |
| SVM decision value | 0.99 |
| SVM RNA-class probability | 0.896451 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 19318749 120 + 22407834 AUCAGGAGAUGUGACAAAUGAACUGUCACAAUAAUGCCAGCGACGAAAGGGAAUGACGACACAUGUGUUGGCACCGUUUCGGUUCUGGAGGAGUUUUCGGAUCUCCGGAUACGACUUCCC ....(((..(((((((.......)))))))......((((...(((((.((..((.(((((....))))).)))).)))))...))))((..((.(((((....))))).))..))))). ( -40.10) >DroSec_CAF1 11333 119 + 1 AUCAGGAGAUGUGACAAAUGAACUGUCACAAUAAUGCCAGCGACGAAAGGGAAUGACGACACAUGUGUUGGCACUGUUUCGGCUCUGGAGGAGUCUCCGGAUCUCUGGUUACGAG-UCCC ....((((((((((((.......)))))).......((((.(.(((((.((..((.(((((....))))).)))).))))).).))))....))))))(((.(((.......)))-))). ( -40.70) >DroSim_CAF1 13170 119 + 1 AUCAGGAGAUGUGACAAAUGAACUGUCACAAUAAUGCCAGCGACGAAAGGGAAUGACGACACAUGUGUUGGCACCGUUUCGGCUCUGGAGGAGUCUCCGGAUCUCUGGUUACGAG-UCCC ....((((((((((((.......)))))).......((((.(.(((((.((..((.(((((....))))).)))).))))).).))))....))))))(((.(((.......)))-))). ( -42.50) >DroEre_CAF1 10523 100 + 1 AUCAGAGGAUGUGACAAAUGAACUGUCACAAUAAUGCCAGCGACGAAAGGGAAUGACGACACACAUGCUGGC-------------------AGUCUCCGGAUUUCUGGAGACGCG-ACCC ......((.(((((((.......)))))))....(((((((..(....)...(((........)))))))))-------------------)((((((((....))))))))...-.)). ( -38.20) >DroYak_CAF1 11613 119 + 1 AUCAGUGGAUGUGACAAAUGAACUGUCACAAUAAUGCCAGCGACGAAAGGGAAUGACGACACAUAUGUUGGCACCGUUUCGCAACUGGAGGAGUCUCCGGAUCUCUGGAUACGAG-ACCG ......((.(((((((.......)))))))....((((((((.(....)...(((......))).))))))))))((((((...((((((....))))))(((....))).))))-)).. ( -37.90) >consensus AUCAGGAGAUGUGACAAAUGAACUGUCACAAUAAUGCCAGCGACGAAAGGGAAUGACGACACAUGUGUUGGCACCGUUUCGGCUCUGGAGGAGUCUCCGGAUCUCUGGAUACGAG_UCCC ....((((((((((((.......)))))).....((((((((.(....)...(((......))).))))))))...................))))))(((.(((.(....)))).))). (-28.26 = -29.10 + 0.84)



| Location | 19,318,749 – 19,318,869 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 86.18 |
| Mean single sequence MFE | -35.38 |
| Consensus MFE | -23.48 |
| Energy contribution | -25.88 |
| Covariance contribution | 2.40 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.83 |
| Structure conservation index | 0.66 |
| SVM decision value | -0.00 |
| SVM RNA-class probability | 0.532153 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 19318749 120 - 22407834 GGGAAGUCGUAUCCGGAGAUCCGAAAACUCCUCCAGAACCGAAACGGUGCCAACACAUGUGUCGUCAUUCCCUUUCGUCGCUGGCAUUAUUGUGACAGUUCAUUUGUCACAUCUCCUGAU .(((.......)))(((((.............((((...(((((.((((.(.(((....))).).))..)).)))))...))))......((((((((.....))))))))))))).... ( -32.20) >DroSec_CAF1 11333 119 - 1 GGGA-CUCGUAACCAGAGAUCCGGAGACUCCUCCAGAGCCGAAACAGUGCCAACACAUGUGUCGUCAUUCCCUUUCGUCGCUGGCAUUAUUGUGACAGUUCAUUUGUCACAUCUCCUGAU .(((-(((.......))).)))(((((.....((((.(.(((((.((((.(.(((....))).).))))...))))).).))))......((((((((.....))))))))))))).... ( -36.50) >DroSim_CAF1 13170 119 - 1 GGGA-CUCGUAACCAGAGAUCCGGAGACUCCUCCAGAGCCGAAACGGUGCCAACACAUGUGUCGUCAUUCCCUUUCGUCGCUGGCAUUAUUGUGACAGUUCAUUUGUCACAUCUCCUGAU .(((-(((.......))).)))(((((.....((((.(.(((((.((((.(.(((....))).).))..)).))))).).))))......((((((((.....))))))))))))).... ( -37.50) >DroEre_CAF1 10523 100 - 1 GGGU-CGCGUCUCCAGAAAUCCGGAGACU-------------------GCCAGCAUGUGUGUCGUCAUUCCCUUUCGUCGCUGGCAUUAUUGUGACAGUUCAUUUGUCACAUCCUCUGAU ((((-...((((((.(....).))))))(-------------------(((((((((.(((....))).......))).))))))).....(((((((.....)))))))))))...... ( -33.60) >DroYak_CAF1 11613 119 - 1 CGGU-CUCGUAUCCAGAGAUCCGGAGACUCCUCCAGUUGCGAAACGGUGCCAACAUAUGUGUCGUCAUUCCCUUUCGUCGCUGGCAUUAUUGUGACAGUUCAUUUGUCACAUCCACUGAU .(((-(((.......)))))).(((.......(((((.((((((.((((.(.(((....))).).))..)).)))))).)))))......((((((((.....)))))))))))...... ( -37.10) >consensus GGGA_CUCGUAUCCAGAGAUCCGGAGACUCCUCCAGAGCCGAAACGGUGCCAACACAUGUGUCGUCAUUCCCUUUCGUCGCUGGCAUUAUUGUGACAGUUCAUUUGUCACAUCUCCUGAU .(((.(((.......))).)))(((((..................(((((((......(((.((...........)).))))))))))..((((((((.....))))))))))))).... (-23.48 = -25.88 + 2.40)


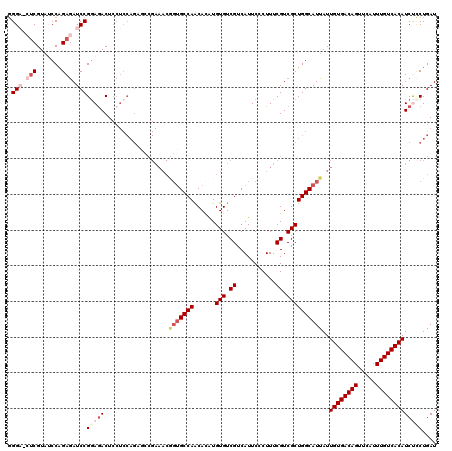
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:23:43 2006