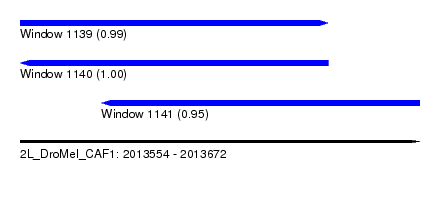

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 2,013,554 – 2,013,672 |
| Length | 118 |
| Max. P | 0.999968 |

| Location | 2,013,554 – 2,013,645 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 92 |
| Reading direction | forward |
| Mean pairwise identity | 97.37 |
| Mean single sequence MFE | -17.24 |
| Consensus MFE | -16.05 |
| Energy contribution | -16.05 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.84 |
| Structure conservation index | 0.93 |
| SVM decision value | 2.20 |
| SVM RNA-class probability | 0.990191 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 2013554 91 + 22407834 GAAAGCAAACAA-AAUAUUUCCAUUUCUUCGUUUGCGGGCCGAAAACCUUUUUCGGUUUCCUCUGCUUUUCUUUAUGUUUUGUUUCAUUCCG (((.(.((((((-(((((................(((((((((((.....))))))).....))))........)))))))))))).))).. ( -17.65) >DroSec_CAF1 31099 91 + 1 GAAAGCAAACAA-AAUAUUUCCAUUUCUUCGUUUGCGGGCCGAAAACCUUUUUCGGUUUCCUCUGCUUUUCUUUAUGUUUUGUUUCAUUCCG (((.(.((((((-(((((................(((((((((((.....))))))).....))))........)))))))))))).))).. ( -17.65) >DroSim_CAF1 29437 91 + 1 GAAAGCAAACAA-AAUAUUUCCAUUUCUUCGUUUGCGGGCCGAAAACCUUUUUCGGUUUCCUCUGCUUUUCUUUAUGUUUUGUUUCAUUCCG (((.(.((((((-(((((................(((((((((((.....))))))).....))))........)))))))))))).))).. ( -17.65) >DroEre_CAF1 28271 92 + 1 GAAAGCAAACAAAAAUAUUCCCAUUUCAUCGUUUGCGGGCCGAAAACCUUUUUCGGUUUCCUUUGCUUUUCCUUAUGUUUUGUUUCAUUCCG ((((((((((....................))))))(((((((((.....)))))))..)).....................))))...... ( -16.05) >DroYak_CAF1 32638 91 + 1 GAAAGCAAACAA-AAUAUUCCCAUUUCUUCGUUUGCGGGCCGAAAACCUUUUUCGGUUUCCUUUGCUUUUCUUUAUGUUUUGUUUCAUUCCG (((.(.((((((-(((((..........(((....)))(((((((.....))))))).................)))))))))))).))).. ( -17.20) >consensus GAAAGCAAACAA_AAUAUUUCCAUUUCUUCGUUUGCGGGCCGAAAACCUUUUUCGGUUUCCUCUGCUUUUCUUUAUGUUUUGUUUCAUUCCG ((((((((((....................))))))(((((((((.....)))))))..)).....................))))...... (-16.05 = -16.05 + 0.00)

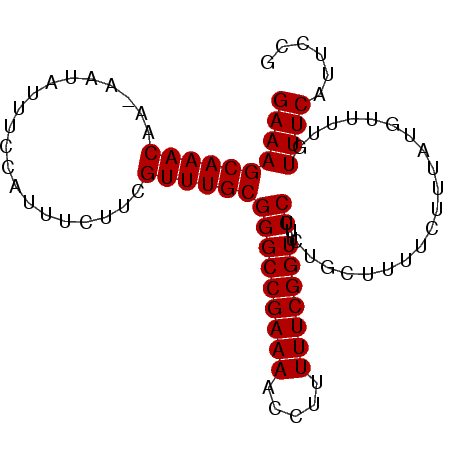
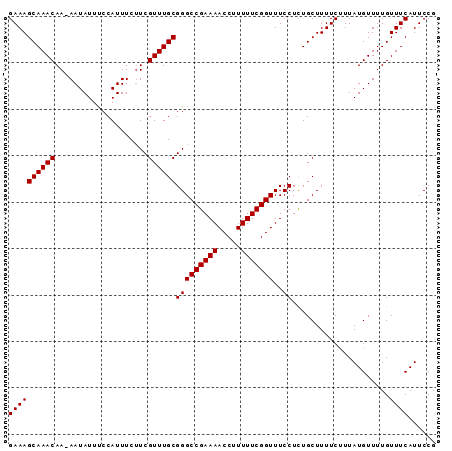
| Location | 2,013,554 – 2,013,645 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 92 |
| Reading direction | reverse |
| Mean pairwise identity | 97.37 |
| Mean single sequence MFE | -21.42 |
| Consensus MFE | -21.50 |
| Energy contribution | -21.50 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.91 |
| Structure conservation index | 1.00 |
| SVM decision value | 5.01 |
| SVM RNA-class probability | 0.999968 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
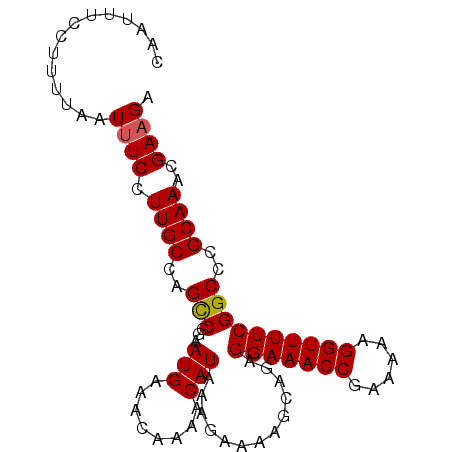
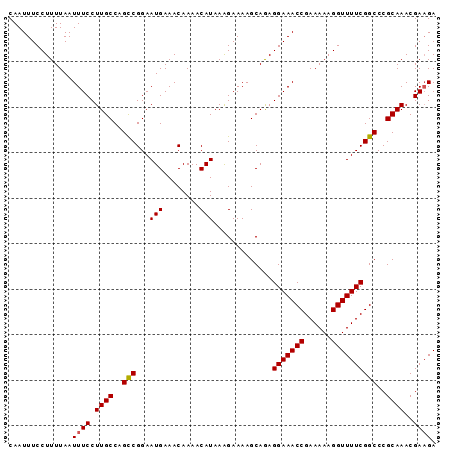
>2L_DroMel_CAF1 2013554 91 - 22407834 CGGAAUGAAACAAAACAUAAAGAAAAGCAGAGGAAACCGAAAAAGGUUUUCGGCCCGCAAACGAAGAAAUGGAAAUAUU-UUGUUUGCUUUC .((.(((........)))........((...(((((((......))))))).))))((((((((((..((....)).))-)))))))).... ( -21.80) >DroSec_CAF1 31099 91 - 1 CGGAAUGAAACAAAACAUAAAGAAAAGCAGAGGAAACCGAAAAAGGUUUUCGGCCCGCAAACGAAGAAAUGGAAAUAUU-UUGUUUGCUUUC .((.(((........)))........((...(((((((......))))))).))))((((((((((..((....)).))-)))))))).... ( -21.80) >DroSim_CAF1 29437 91 - 1 CGGAAUGAAACAAAACAUAAAGAAAAGCAGAGGAAACCGAAAAAGGUUUUCGGCCCGCAAACGAAGAAAUGGAAAUAUU-UUGUUUGCUUUC .((.(((........)))........((...(((((((......))))))).))))((((((((((..((....)).))-)))))))).... ( -21.80) >DroEre_CAF1 28271 92 - 1 CGGAAUGAAACAAAACAUAAGGAAAAGCAAAGGAAACCGAAAAAGGUUUUCGGCCCGCAAACGAUGAAAUGGGAAUAUUUUUGUUUGCUUUC .((.(((........)))........((...(((((((......))))))).))))((((((((..(((((....))))))))))))).... ( -21.20) >DroYak_CAF1 32638 91 - 1 CGGAAUGAAACAAAACAUAAAGAAAAGCAAAGGAAACCGAAAAAGGUUUUCGGCCCGCAAACGAAGAAAUGGGAAUAUU-UUGUUUGCUUUC .((.(((........)))........((...(((((((......))))))).))))((((((((((...........))-)))))))).... ( -20.50) >consensus CGGAAUGAAACAAAACAUAAAGAAAAGCAGAGGAAACCGAAAAAGGUUUUCGGCCCGCAAACGAAGAAAUGGAAAUAUU_UUGUUUGCUUUC .((.(((........)))........((...(((((((......))))))).))))((((((((.((.((....)).)).)))))))).... (-21.50 = -21.50 + 0.00)

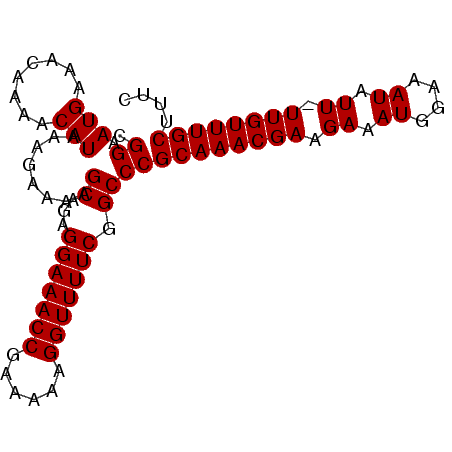
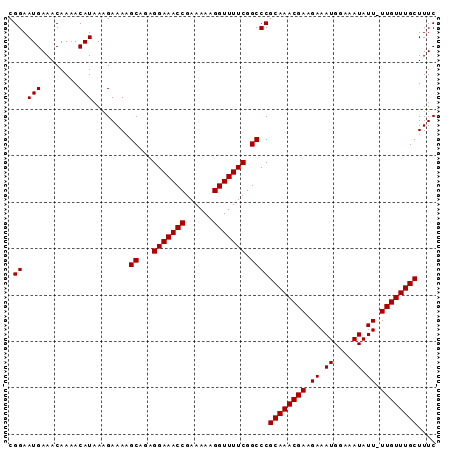
| Location | 2,013,578 – 2,013,672 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 94 |
| Reading direction | reverse |
| Mean pairwise identity | 98.09 |
| Mean single sequence MFE | -18.68 |
| Consensus MFE | -17.72 |
| Energy contribution | -17.76 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.57 |
| Structure conservation index | 0.95 |
| SVM decision value | 1.40 |
| SVM RNA-class probability | 0.950598 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 2013578 94 - 22407834 CAAUUUCCUUUUAAUUUCCUUGCCAGUCGGAAUGAAACAAAACAUAAAGAAAAGCAGAGGAAACCGAAAAAGGUUUUCGGCCCGCAAACGAAGA ......((((((..(((((((((...((...(((........)))...))...))).))))))....))))))((((((.........)))))) ( -17.80) >DroSec_CAF1 31123 94 - 1 CAAUUUCCUUUUAAUUUCCUUGCCAGCCGGAAUGAAACAAAACAUAAAGAAAAGCAGAGGAAACCGAAAAAGGUUUUCGGCCCGCAAACGAAGA ..............((((.((((..(((...(((........))).............(((((((......))))))))))..))))..)))). ( -18.60) >DroSim_CAF1 29461 94 - 1 CAAUUUCCUUUUAAUUUCCUUGCCAGCCGGAAUGAAACAAAACAUAAAGAAAAGCAGAGGAAACCGAAAAAGGUUUUCGGCCCGCAAACGAAGA ..............((((.((((..(((...(((........))).............(((((((......))))))))))..))))..)))). ( -18.60) >DroEre_CAF1 28296 94 - 1 CAAUUUCCUUUUAAUUUCCUUGCCAGCCGGAAUGAAACAAAACAUAAGGAAAAGCAAAGGAAACCGAAAAAGGUUUUCGGCCCGCAAACGAUGA ...((((((((...(((((((.((....)).(((........))))))))))...))))))))((((((.....)))))).............. ( -19.80) >DroYak_CAF1 32662 94 - 1 CAAUUUCCUUUUAAUUUCCUUGCCAGCCGGAAUGAAACAAAACAUAAAGAAAAGCAAAGGAAACCGAAAAAGGUUUUCGGCCCGCAAACGAAGA ..............((((.((((..(((...(((........))).............(((((((......))))))))))..))))..)))). ( -18.60) >consensus CAAUUUCCUUUUAAUUUCCUUGCCAGCCGGAAUGAAACAAAACAUAAAGAAAAGCAGAGGAAACCGAAAAAGGUUUUCGGCCCGCAAACGAAGA ..............((((.((((..(((...(((........))).............(((((((......))))))))))..))))..)))). (-17.72 = -17.76 + 0.04)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:43:10 2006