
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 19,272,229 – 19,272,348 |
| Length | 119 |
| Max. P | 0.954879 |

| Location | 19,272,229 – 19,272,348 |
|---|---|
| Length | 119 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 96.37 |
| Mean single sequence MFE | -31.02 |
| Consensus MFE | -28.34 |
| Energy contribution | -28.37 |
| Covariance contribution | 0.03 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.56 |
| Structure conservation index | 0.91 |
| SVM decision value | 1.45 |
| SVM RNA-class probability | 0.954879 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 19272229 119 + 22407834 AUGCAAAU-UUCUGUCGACAUCGACGCACUAAAUGCGACUUGCAAAUUAAUGAAAUGAACCUUUAACUUGAGUUUAACUUGGCGCCAUUUUAGCCAGAACGCGCGAGAUAAGGCGCCACC .(((((..-....((((....))))(((.....)))...)))))...........((((((........).)))))...((((((((((((.((......))..)))))..))))))).. ( -30.60) >DroSec_CAF1 13463 119 + 1 AUGCAAAU-UUCUGUCGACAUCGACGCACUAAAUGAGACUUGCAAAUUAAUGAAAUGAACCUUUAACUUGAGUUUAACUUGGCGCCAUUUUAGCCAGAACGUGCGAGAUAAGGCGCCACC .(((((.(-((((((((....)))))........)))).)))))...........((((((........).)))))...((((((((((((.(((.....).)))))))..))))))).. ( -28.80) >DroSim_CAF1 20346 119 + 1 AUGCAAAU-UUCUGUCGACAUCGACGCACUAAAUGCGACUUGCAAAUUAAUGAAAUGAACCUUUAACUUGAGUUUAACUUGGCGCCAUUUUAGCCAGAACGCGCGAGAUAAGGCGCCACC .(((((..-....((((....))))(((.....)))...)))))...........((((((........).)))))...((((((((((((.((......))..)))))..))))))).. ( -30.60) >DroEre_CAF1 20207 119 + 1 AUGCAAAU-UUCUGUCGACAUCGACGCACUAAAUGCGACUUGCAAAUUAAUGAAAUGAACCUUUAACUUGAGUUUAACUUGGCGCCAUUUUAGCCAGAACGGGCGAGAUAAGGCGCCACC .(((((..-....((((....))))(((.....)))...)))))...........((((((........).)))))...((((((((((((.(((......))))))))..))))))).. ( -33.40) >DroYak_CAF1 19491 119 + 1 AUGCAAAU-UUCUGUCGACAUCGACGCACUAAAUGCGACUUGCAAAUUAAUGAAAUGAACCUUUAACUUGAGUUUAACUUGGCGCCAUUUUAGCCGGAACGCGCGAGAUAAGGCGCCACC .(((((..-....((((....))))(((.....)))...)))))...........((((((........).)))))...((((((((((((.((......))..)))))..))))))).. ( -30.60) >DroAna_CAF1 35680 116 + 1 AUGCAAAUUUUCUGUCGACAUCGACGCACUAAAUGCGACUUGCAAAUUAAUGAAAUGAACCUUUAACUUGAGUUUAACUUGGCGCCUUUUUAGC----GCCGGCGAGAUAAGGCGCCGCC .(((((.......((((....))))(((.....)))...)))))...........((((((........).)))))....((((((((....((----....)).....))))))))... ( -32.10) >consensus AUGCAAAU_UUCUGUCGACAUCGACGCACUAAAUGCGACUUGCAAAUUAAUGAAAUGAACCUUUAACUUGAGUUUAACUUGGCGCCAUUUUAGCCAGAACGCGCGAGAUAAGGCGCCACC .(((((.......((((....))))(((.....)))...)))))...........((((((........).)))))...(((((((..(((.((........)).)))...))))))).. (-28.34 = -28.37 + 0.03)


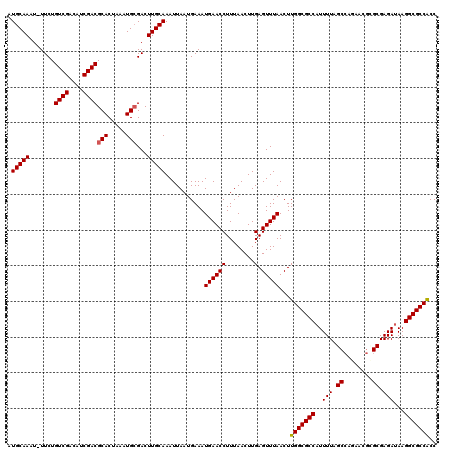
| Location | 19,272,229 – 19,272,348 |
|---|---|
| Length | 119 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 96.37 |
| Mean single sequence MFE | -31.62 |
| Consensus MFE | -29.29 |
| Energy contribution | -29.62 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -0.90 |
| Structure conservation index | 0.93 |
| SVM decision value | -0.04 |
| SVM RNA-class probability | 0.513527 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 19272229 119 - 22407834 GGUGGCGCCUUAUCUCGCGCGUUCUGGCUAAAAUGGCGCCAAGUUAAACUCAAGUUAAAGGUUCAUUUCAUUAAUUUGCAAGUCGCAUUUAGUGCGUCGAUGUCGACAGAA-AUUUGCAU ..(((((((.......((........))......)))))))..........(((((((.((......)).)))))))((((((.((((...))))((((....))))....-)))))).. ( -31.42) >DroSec_CAF1 13463 119 - 1 GGUGGCGCCUUAUCUCGCACGUUCUGGCUAAAAUGGCGCCAAGUUAAACUCAAGUUAAAGGUUCAUUUCAUUAAUUUGCAAGUCUCAUUUAGUGCGUCGAUGUCGACAGAA-AUUUGCAU ..(((((((.......((........))......)))))))..........(((((((.((......)).)))))))((((((.((((...))..((((....)))).)).-)))))).. ( -26.92) >DroSim_CAF1 20346 119 - 1 GGUGGCGCCUUAUCUCGCGCGUUCUGGCUAAAAUGGCGCCAAGUUAAACUCAAGUUAAAGGUUCAUUUCAUUAAUUUGCAAGUCGCAUUUAGUGCGUCGAUGUCGACAGAA-AUUUGCAU ..(((((((.......((........))......)))))))..........(((((((.((......)).)))))))((((((.((((...))))((((....))))....-)))))).. ( -31.42) >DroEre_CAF1 20207 119 - 1 GGUGGCGCCUUAUCUCGCCCGUUCUGGCUAAAAUGGCGCCAAGUUAAACUCAAGUUAAAGGUUCAUUUCAUUAAUUUGCAAGUCGCAUUUAGUGCGUCGAUGUCGACAGAA-AUUUGCAU ..(((((((.......(((......)))......)))))))..........(((((((.((......)).)))))))((((((.((((...))))((((....))))....-)))))).. ( -34.12) >DroYak_CAF1 19491 119 - 1 GGUGGCGCCUUAUCUCGCGCGUUCCGGCUAAAAUGGCGCCAAGUUAAACUCAAGUUAAAGGUUCAUUUCAUUAAUUUGCAAGUCGCAUUUAGUGCGUCGAUGUCGACAGAA-AUUUGCAU ..(((((((.......((.((...))))......)))))))..........(((((((.((......)).)))))))((((((.((((...))))((((....))))....-)))))).. ( -31.62) >DroAna_CAF1 35680 116 - 1 GGCGGCGCCUUAUCUCGCCGGC----GCUAAAAAGGCGCCAAGUUAAACUCAAGUUAAAGGUUCAUUUCAUUAAUUUGCAAGUCGCAUUUAGUGCGUCGAUGUCGACAGAAAAUUUGCAU (((((((........))))(((----(((.....))))))..)))......(((((((.((......)).)))))))((((((.((((...))))((((....)))).....)))))).. ( -34.20) >consensus GGUGGCGCCUUAUCUCGCGCGUUCUGGCUAAAAUGGCGCCAAGUUAAACUCAAGUUAAAGGUUCAUUUCAUUAAUUUGCAAGUCGCAUUUAGUGCGUCGAUGUCGACAGAA_AUUUGCAU ..(((((((.......((........))......)))))))..........(((((((.((......)).)))))))((((((.((((...))))((((....)))).....)))))).. (-29.29 = -29.62 + 0.33)

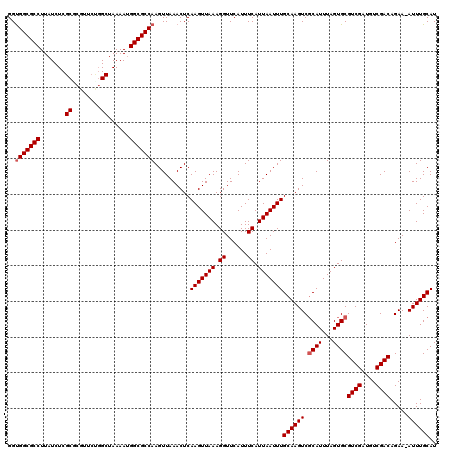
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:22:36 2006