
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 19,243,931 – 19,244,063 |
| Length | 132 |
| Max. P | 0.994089 |

| Location | 19,243,931 – 19,244,043 |
|---|---|
| Length | 112 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 72.99 |
| Mean single sequence MFE | -35.60 |
| Consensus MFE | -19.43 |
| Energy contribution | -20.38 |
| Covariance contribution | 0.95 |
| Combinations/Pair | 1.30 |
| Mean z-score | -2.49 |
| Structure conservation index | 0.55 |
| SVM decision value | 2.40 |
| SVM RNA-class probability | 0.993529 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 19243931 112 + 22407834 GGCAUGCAGCGAGGA-----UAUAGCCGGA-UAGCGGAAAACGCUGGCAAAGCUGAGCCGGAACUCAGCGGAUCCGGCCGCAUAAGAUUAGCCAGUAAAAUAUUGUACAUUUUGCACG-- (((((((.((..(((-----(...(((...-.((((.....)))))))...((((((......))))))..)))).)).)))).......))).(((((((.......)))))))...-- ( -37.41) >DroVir_CAF1 77016 99 + 1 AGC---CGUUCG---------------GAA-AUACGGAAAGUGUUUGUAAAACUGAGCCGGAACACUAAAGUUCC-GCCGCAAGAGAUUAG-CAGUAAAAUAUUUUAUGUUUCGCAUACU .((---.(((((---------------(..-.((((((.....))))))...))))))((((((......)))))-).(....)......)-).(..((((((...))))))..)..... ( -21.60) >DroPse_CAF1 71818 103 + 1 GGC-UACGGUCA---------------GGG-AAGCGGAAAACGCUGGCAAAGCUGAGCCGGAACUCACCAGUUCCGGCCGCAUAAGAUUAGUCCGUAAAAUAUUGUAUAUUUUGCAUGCC (((-....))).---------------...-.((((.....))))((((..((.(.(((((((((....)))))))))))).............(((((((((...))))))))).)))) ( -37.80) >DroMoj_CAF1 96605 99 + 1 AGC---CGUUCG---------------GAA-GAACGGAAAGUGUUUGCAAAGCUGAGCCGGAACUAGCAAGUUCC-GCCGCAAAAGAUUAG-CAGAAAAAUAUUUUAUGUUUCGCAUGCU ..(---(((((.---------------...-)))))).....((.(((...((((((((((((((....))))))-)..))......))))-).((((.(((...))).))))))).)). ( -31.20) >DroAna_CAF1 114234 120 + 1 GGCGAGCACUGGGUAGCCAGUGUAGCCGUGUCAGCGGAAAACGCCGGCAAAGCUGAGCCGGAACUCAACGGUUCCGGUGGCAUAAGAUUAGCCAGUAAAAUAUUGUAUAUUUUGCAUGCU (((..(((((((....))))))).(((.((((.(((.....))).)))).......(((((((((....)))))))))))).........))).(((((((((...)))))))))..... ( -50.30) >DroPer_CAF1 74993 103 + 1 GGC-UACGGUCA---------------GGG-AAGCGGAAAACGCUGGCAAAGCUGAGCCGGAACUCACCAGUUCCGGGCGCAUAAGAUUAGUCCGUAAAAUAUUGUAUAUUUUGCAUGCC (((-(.((((..---------------.(.-.((((.....))))..)...))))))))((((((....))))))((((...........))))(((((((((...)))))))))..... ( -35.30) >consensus GGC_U_CGGUCA_______________GGA_AAGCGGAAAACGCUGGCAAAGCUGAGCCGGAACUCACCAGUUCCGGCCGCAUAAGAUUAG_CAGUAAAAUAUUGUAUAUUUUGCAUGCU ...............................(((((.....)))))(((..((...((((((((......)))))))).)).............(((((((((...))))))))).))). (-19.43 = -20.38 + 0.95)


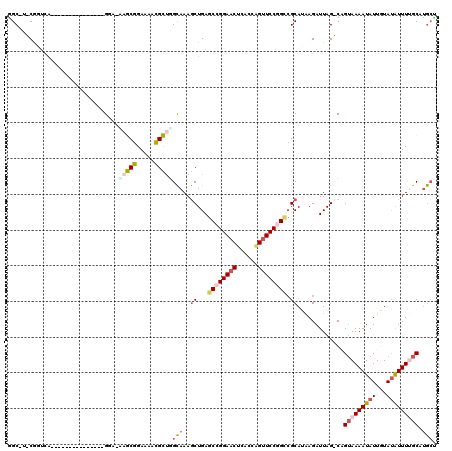
| Location | 19,243,931 – 19,244,043 |
|---|---|
| Length | 112 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 72.99 |
| Mean single sequence MFE | -30.73 |
| Consensus MFE | -16.33 |
| Energy contribution | -16.87 |
| Covariance contribution | 0.53 |
| Combinations/Pair | 1.26 |
| Mean z-score | -1.88 |
| Structure conservation index | 0.53 |
| SVM decision value | 1.06 |
| SVM RNA-class probability | 0.908833 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 19243931 112 - 22407834 --CGUGCAAAAUGUACAAUAUUUUACUGGCUAAUCUUAUGCGGCCGGAUCCGCUGAGUUCCGGCUCAGCUUUGCCAGCGUUUUCCGCUA-UCCGGCUAUA-----UCCUCGCUGCAUGCC --.((((.....))))...........(((.......(((((((.((((..(((((((....)))))))...(((((((.....)))).-...)))...)-----)))..)))))))))) ( -37.31) >DroVir_CAF1 77016 99 - 1 AGUAUGCGAAACAUAAAAUAUUUUACUG-CUAAUCUCUUGCGGC-GGAACUUUAGUGUUCCGGCUCAGUUUUACAAACACUUUCCGUAU-UUC---------------CGAACG---GCU .((.((..((((.............(((-(.........))))(-(((((......)))))).....))))..)).)).....((((..-...---------------...)))---).. ( -22.70) >DroPse_CAF1 71818 103 - 1 GGCAUGCAAAAUAUACAAUAUUUUACGGACUAAUCUUAUGCGGCCGGAACUGGUGAGUUCCGGCUCAGCUUUGCCAGCGUUUUCCGCUU-CCC---------------UGACCGUA-GCC (((....(((((((...)))))))((((...........((((((((((((....))))))))))..))......((((.....)))).-...---------------...)))).-))) ( -31.50) >DroMoj_CAF1 96605 99 - 1 AGCAUGCGAAACAUAAAAUAUUUUUCUG-CUAAUCUUUUGCGGC-GGAACUUGCUAGUUCCGGCUCAGCUUUGCAAACACUUUCCGUUC-UUC---------------CGAACG---GCU .(((.((((((.(((...))).))))..-.........((.(.(-((((((....))))))).).))))..))).........((((((-...---------------.)))))---).. ( -27.00) >DroAna_CAF1 114234 120 - 1 AGCAUGCAAAAUAUACAAUAUUUUACUGGCUAAUCUUAUGCCACCGGAACCGUUGAGUUCCGGCUCAGCUUUGCCGGCGUUUUCCGCUGACACGGCUACACUGGCUACCCAGUGCUCGCC .(((.(((((((((...)))))))..((((.........))))(((((((......)))))))....))..)))(((((.....)))))....(((..((((((....))))))...))) ( -38.50) >DroPer_CAF1 74993 103 - 1 GGCAUGCAAAAUAUACAAUAUUUUACGGACUAAUCUUAUGCGCCCGGAACUGGUGAGUUCCGGCUCAGCUUUGCCAGCGUUUUCCGCUU-CCC---------------UGACCGUA-GCC (((....(((((((...)))))))((((.............(((.((((((....)))))))))((((.......((((.....)))).-..)---------------))))))).-))) ( -27.40) >consensus AGCAUGCAAAAUAUACAAUAUUUUACUG_CUAAUCUUAUGCGGCCGGAACUGGUGAGUUCCGGCUCAGCUUUGCCAGCGUUUUCCGCUU_CCC_______________CGACCG_A_GCC .(((.(((((((((...)))))))..............((.(((((((((......)))))))))))))..))).((((.....))))................................ (-16.33 = -16.87 + 0.53)



| Location | 19,243,965 – 19,244,063 |
|---|---|
| Length | 98 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 76.06 |
| Mean single sequence MFE | -36.90 |
| Consensus MFE | -23.75 |
| Energy contribution | -24.33 |
| Covariance contribution | 0.59 |
| Combinations/Pair | 1.21 |
| Mean z-score | -2.94 |
| Structure conservation index | 0.64 |
| SVM decision value | 2.45 |
| SVM RNA-class probability | 0.994089 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 19243965 98 + 22407834 ACGCUGGCAAAGCUGAGCCGGAACUCAGCGGAUCCGGCCGCAUAAGAUUAGCCAGUAAAAUAUUGUACAUUUUGCACG-------------------AGGGUCCA---ACUGAGUGCCAA ....(((((..((((((......))))))(((((((((............))).(((((((.......)))))))...-------------------.)))))).---......))))). ( -34.70) >DroPse_CAF1 71841 117 + 1 ACGCUGGCAAAGCUGAGCCGGAACUCACCAGUUCCGGCCGCAUAAGAUUAGUCCGUAAAAUAUUGUAUAUUUUGCAUGCCCCAGCUCCACAGAAGCACGAGUCCA---UCUGAGUGCCAA ....(((((.(((((.(((((((((....))))))))).((((..((....)).(((((((((...)))))))))))))..)))))...((((.((....))...---))))..))))). ( -39.10) >DroSim_CAF1 65271 98 + 1 ACGCUGGCAAAGCUGAGCCGGAACUCAGCGGAUCCGGCCGCAUAAGAUUAGCCAGUAAAAUAUUGUACAUUUUGCACG-------------------AGGGUCCA---ACUGAGUGCCAA ....(((((..((((((......))))))(((((((((............))).(((((((.......)))))))...-------------------.)))))).---......))))). ( -34.70) >DroMoj_CAF1 96626 105 + 1 GUGUUUGCAAAGCUGAGCCGGAACUAGCAAGUUCC-GCCGCAAAAGAUUAG-CAGAAAAAUAUUUUAUGUUUCGCAUGCUCCAA-------------AAUGUCGAGAGAUAAAGUGUAAA ...(((((....((..(((((((((....))))))-)..))...))....)-))))...(((((((((.((((((((.......-------------.))).))))).)))))))))... ( -30.50) >DroAna_CAF1 114274 114 + 1 ACGCCGGCAAAGCUGAGCCGGAACUCAACGGUUCCGGUGGCAUAAGAUUAGCCAGUAAAAUAUUGUAUAUUUUGCAUGCUCCA---CAGCAGAAGGGCGAGUCCA---GCUGAGUGCCAU .....((((.(((((.(((((((((....))))))))).......((((.(((.(((((((((...))))))))).((((...---.))))....))).))))))---)))...)))).. ( -45.70) >DroPer_CAF1 75016 117 + 1 ACGCUGGCAAAGCUGAGCCGGAACUCACCAGUUCCGGGCGCAUAAGAUUAGUCCGUAAAAUAUUGUAUAUUUUGCAUGCCCCAGCUCCACAGAAGCACGAGUCCA---UCUGAGUGCCAA ....(((((.....((((.((((((....))))))(((.((((..((....)).(((((((((...)))))))))))))))).))))..((((.((....))...---))))..))))). ( -36.70) >consensus ACGCUGGCAAAGCUGAGCCGGAACUCACCAGUUCCGGCCGCAUAAGAUUAGCCAGUAAAAUAUUGUAUAUUUUGCAUGC_CCA______________AGAGUCCA___ACUGAGUGCCAA ....(((((..((...(((((((((....))))))))).)).............((((((((.....)))))))).......................................))))). (-23.75 = -24.33 + 0.59)


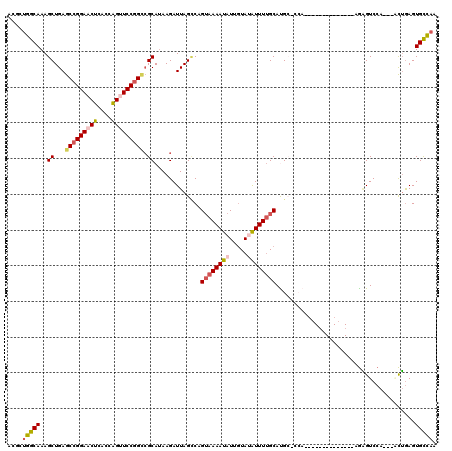
| Location | 19,243,965 – 19,244,063 |
|---|---|
| Length | 98 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 76.06 |
| Mean single sequence MFE | -33.58 |
| Consensus MFE | -18.52 |
| Energy contribution | -19.08 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.57 |
| Structure conservation index | 0.55 |
| SVM decision value | 0.04 |
| SVM RNA-class probability | 0.551647 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 19243965 98 - 22407834 UUGGCACUCAGU---UGGACCCU-------------------CGUGCAAAAUGUACAAUAUUUUACUGGCUAAUCUUAUGCGGCCGGAUCCGCUGAGUUCCGGCUCAGCUUUGCCAGCGU ..((.(((((((---.(((.((.-------------------.((((.....))))...........((((..........)))))).)))))))))).))(((........)))..... ( -33.00) >DroPse_CAF1 71841 117 - 1 UUGGCACUCAGA---UGGACUCGUGCUUCUGUGGAGCUGGGGCAUGCAAAAUAUACAAUAUUUUACGGACUAAUCUUAUGCGGCCGGAACUGGUGAGUUCCGGCUCAGCUUUGCCAGCGU (((((((.((((---.(.(....).).)))).)((((((..(((((.(((((((...)))))))..(((....))))))))((((((((((....))))))))))))))))))))))... ( -40.80) >DroSim_CAF1 65271 98 - 1 UUGGCACUCAGU---UGGACCCU-------------------CGUGCAAAAUGUACAAUAUUUUACUGGCUAAUCUUAUGCGGCCGGAUCCGCUGAGUUCCGGCUCAGCUUUGCCAGCGU ..((.(((((((---.(((.((.-------------------.((((.....))))...........((((..........)))))).)))))))))).))(((........)))..... ( -33.00) >DroMoj_CAF1 96626 105 - 1 UUUACACUUUAUCUCUCGACAUU-------------UUGGAGCAUGCGAAACAUAAAAUAUUUUUCUG-CUAAUCUUUUGCGGC-GGAACUUGCUAGUUCCGGCUCAGCUUUGCAAACAC ............(((.(((....-------------))))))..((((((.(.............(((-(.........))))(-((((((....))))))).....).))))))..... ( -20.70) >DroAna_CAF1 114274 114 - 1 AUGGCACUCAGC---UGGACUCGCCCUUCUGCUG---UGGAGCAUGCAAAAUAUACAAUAUUUUACUGGCUAAUCUUAUGCCACCGGAACCGUUGAGUUCCGGCUCAGCUUUGCCGGCGU .((((((.((((---.((......))....))))---.).(((....(((((((...)))))))....))).......)))))((((....(((((((....)))))))....))))... ( -34.20) >DroPer_CAF1 75016 117 - 1 UUGGCACUCAGA---UGGACUCGUGCUUCUGUGGAGCUGGGGCAUGCAAAAUAUACAAUAUUUUACGGACUAAUCUUAUGCGCCCGGAACUGGUGAGUUCCGGCUCAGCUUUGCCAGCGU (((((((.((((---.(.(....).).)))).)(((((((((((((.(((((((...)))))))..(((....))))))))..((((((((....))))))))))))))))))))))... ( -39.80) >consensus UUGGCACUCAGU___UGGACUCG______________UGG_GCAUGCAAAAUAUACAAUAUUUUACUGGCUAAUCUUAUGCGGCCGGAACCGGUGAGUUCCGGCUCAGCUUUGCCAGCGU .(((((.........................................(((((((...)))))))...............(((((((((((......)))))))))..))..))))).... (-18.52 = -19.08 + 0.56)

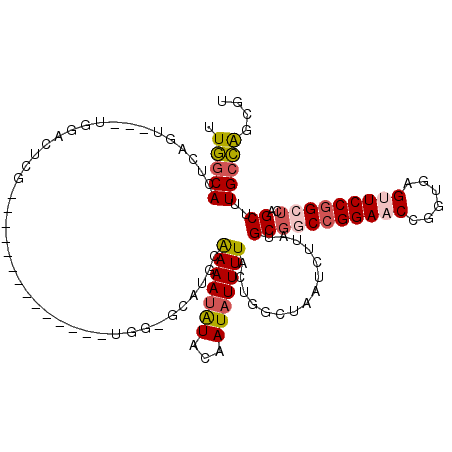

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:21:58 2006