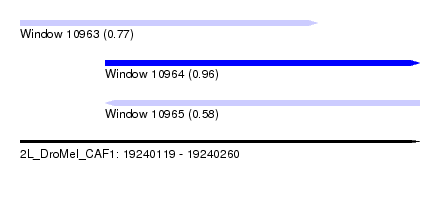
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 19,240,119 – 19,240,260 |
| Length | 141 |
| Max. P | 0.963322 |

| Location | 19,240,119 – 19,240,224 |
|---|---|
| Length | 105 |
| Sequences | 5 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 72.76 |
| Mean single sequence MFE | -22.94 |
| Consensus MFE | -4.11 |
| Energy contribution | -4.39 |
| Covariance contribution | 0.28 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.72 |
| Structure conservation index | 0.18 |
| SVM decision value | 0.52 |
| SVM RNA-class probability | 0.766607 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 19240119 105 + 22407834 CUAAAUGUU-------UGAAUGGACA--U-UGAAGCAUUGAUUAUG--CUCCAUAACAUAAAGGUGUAUAUAAAAUUCUCUCCAUUUCACUGGAUAAGUAAGAACUCUUUCAUUUCU .........-------.(((((((..--.-((.(((((.....)))--)).))..((((....)))).............)))))))...((((..(((....)))..))))..... ( -15.90) >DroSec_CAF1 59985 106 + 1 CCAAGUGUU-------UGAAUGGACA--CGUAUUGCAGUGAUUAUG--CUCCAUGCCAUACAGGUGUGUAUAAAAUUCUCUUCAUUUCACUGGAUAAGUAAGAACUCUUACUCUUCU ...((((..-------((((.(((((--(........))).(((((--(...(((((.....)))))))))))...))).))))...))))(((..((((((....))))))..))) ( -26.20) >DroSim_CAF1 61362 106 + 1 CCAAGUGUU-------UGAAUGGACA--CGUAUUGCAUUGAUUAUG--CUCCAUACCAGACAGGUGUAUAUAAAAUUCUCUUCAUUUCACUGGAUAAGUAAGAACUCUUACUCUUCU ...((((..-------((((.(((..--......((((.....)))--)...(((((.....))))).........))).))))...))))(((..((((((....))))))..))) ( -24.50) >DroEre_CAF1 59604 115 + 1 CAAAGUGUUUU-CCCUUGAAUGGUUGUCCAUACUAUAUGGACUAUAGUCUCCAGACCAUAAGUGUGCACA-AAAAUGCUCUCAAUUUCACUGCAUAUGUAGGAACUCCUACACUAUU ....((((...-(.(((..(((((((((((((...)))))))...........))))))))).).)))).-...((((.............)))).((((((....))))))..... ( -26.22) >DroYak_CAF1 61625 111 + 1 CAAAGUGUUUUAUUCUUGAAUGGUUGUAUGGAC---GUUGACUAUG--CUCCAUACCAUACAUAUGCAUA-AAAAUCCCCUCAAUUUCUGUGCAAAAAUAAGAACUCUUACAGUAUU ..(((.(((((.....((.(((((...((((((---((.....)))--.)))))))))).))..((((((-.((((.......)))).))))))......))))).)))........ ( -21.90) >consensus CAAAGUGUU_______UGAAUGGACA__CGUACUGCAUUGAUUAUG__CUCCAUACCAUACAGGUGUAUAUAAAAUUCUCUCCAUUUCACUGGAUAAGUAAGAACUCUUACACUUCU ...................((((...........................))))...........................................(((((....)))))...... ( -4.11 = -4.39 + 0.28)



| Location | 19,240,149 – 19,240,260 |
|---|---|
| Length | 111 |
| Sequences | 5 |
| Columns | 113 |
| Reading direction | forward |
| Mean pairwise identity | 78.16 |
| Mean single sequence MFE | -20.81 |
| Consensus MFE | -4.34 |
| Energy contribution | -5.50 |
| Covariance contribution | 1.16 |
| Combinations/Pair | 1.20 |
| Mean z-score | -2.44 |
| Structure conservation index | 0.21 |
| SVM decision value | 1.55 |
| SVM RNA-class probability | 0.963322 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 19240149 111 + 22407834 AUUAUG--CUCCAUAACAUAAAGGUGUAUAUAAAAUUCUCUCCAUUUCACUGGAUAAGUAAGAACUCUUUCAUUUCUUACCAGCUUGAUGAACCACCUGCCAGCAUAAUUCCA ((((((--((...........(((((..............((((......))))...(((((((.........))))))).............)))))...)))))))).... ( -18.94) >DroSec_CAF1 60016 108 + 1 AUUAUG--CUCCAUGCCAUACAGGUGUGUAUAAAAUUCUCUUCAUUUCACUGGAUAAGUAAGAACUCUUACUCUUCUUAGCAGCUUGAUGAAC---CUGCCAGCAUAAUUCCA ((((((--((.((((((.....))))))............((((((...(((((..((((((....))))))..))....)))...)))))).---.....)))))))).... ( -25.50) >DroSim_CAF1 61393 108 + 1 AUUAUG--CUCCAUACCAGACAGGUGUAUAUAAAAUUCUCUUCAUUUCACUGGAUAAGUAAGAACUCUUACUCUUCUUAGCAGCUUGAUGAAC---CUGCCAGCAUAAUUCCA ((((((--((..(((((.....))))).............((((((...(((((..((((((....))))))..))....)))...)))))).---.....)))))))).... ( -24.00) >DroEre_CAF1 59643 97 + 1 ACUAUAGUCUCCAGACCAUAAGUGUGCACA-AAAAUGCUCUCAAUUUCACUGCAUAUGUAGGAACUCCUACACUAUUUACCAGCCUGAUGGAC---UUGCC------------ .....(((((.(((.......(((((((..-.((((.......))))...)))))))(((((....))))).............)))..))))---)....------------ ( -20.60) >DroYak_CAF1 61662 107 + 1 ACUAUG--CUCCAUACCAUACAUAUGCAUA-AAAAUCCCCUCAAUUUCUGUGCAAAAAUAAGAACUCUUACAGUAUUUACCAGCUUGAUGAAC---UUGCCAGCAUAAUUCCA ..((((--((.......((((...((((((-.((((.......)))).))))))....((((....))))..))))......((..(.....)---..)).))))))...... ( -15.00) >consensus AUUAUG__CUCCAUACCAUACAGGUGUAUAUAAAAUUCUCUCCAUUUCACUGGAUAAGUAAGAACUCUUACACUUCUUACCAGCUUGAUGAAC___CUGCCAGCAUAAUUCCA ........................................((((((...(((....((((((....))))))........)))...))))))..................... ( -4.34 = -5.50 + 1.16)


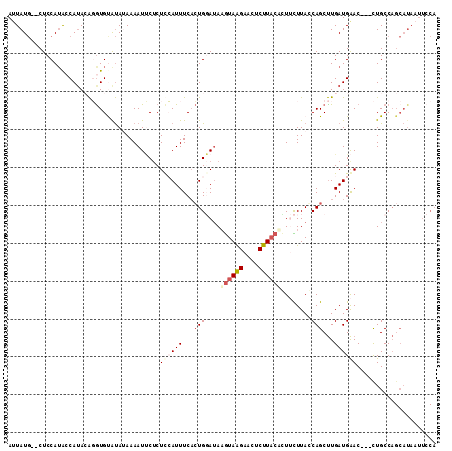
| Location | 19,240,149 – 19,240,260 |
|---|---|
| Length | 111 |
| Sequences | 5 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 78.16 |
| Mean single sequence MFE | -27.28 |
| Consensus MFE | -8.08 |
| Energy contribution | -9.32 |
| Covariance contribution | 1.24 |
| Combinations/Pair | 1.33 |
| Mean z-score | -1.89 |
| Structure conservation index | 0.30 |
| SVM decision value | 0.09 |
| SVM RNA-class probability | 0.580604 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
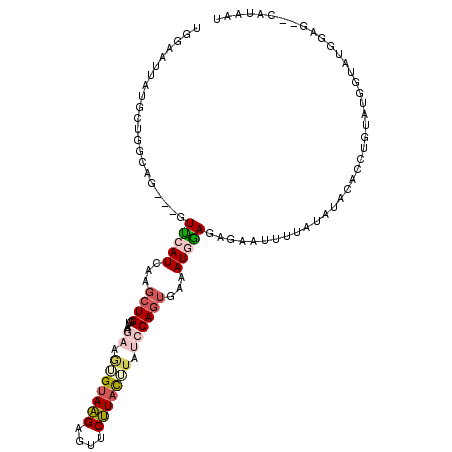
>2L_DroMel_CAF1 19240149 111 - 22407834 UGGAAUUAUGCUGGCAGGUGGUUCAUCAAGCUGGUAAGAAAUGAAAGAGUUCUUACUUAUCCAGUGAAAUGGAGAGAAUUUUAUAUACACCUUUAUGUUAUGGAG--CAUAAU ....((((((((((((((((.(((((....((....))..)))))(((((((((.....((((......))))))))))))).....))))....))))....))--)))))) ( -24.90) >DroSec_CAF1 60016 108 - 1 UGGAAUUAUGCUGGCAG---GUUCAUCAAGCUGCUAAGAAGAGUAAGAGUUCUUACUUAUCCAGUGAAAUGAAGAGAAUUUUAUACACACCUGUAUGGCAUGGAG--CAUAAU ....(((((((((((((---.((....)).))))))....(((((((....))))))).(((((((((((.......))))))).....((.....))..)))))--)))))) ( -28.10) >DroSim_CAF1 61393 108 - 1 UGGAAUUAUGCUGGCAG---GUUCAUCAAGCUGCUAAGAAGAGUAAGAGUUCUUACUUAUCCAGUGAAAUGAAGAGAAUUUUAUAUACACCUGUCUGGUAUGGAG--CAUAAU ....(((((((((((((---.((....)).))))))....(((((((....))))))).(((((((((((.......)))))))....(((.....))).)))))--)))))) ( -29.90) >DroEre_CAF1 59643 97 - 1 ------------GGCAA---GUCCAUCAGGCUGGUAAAUAGUGUAGGAGUUCCUACAUAUGCAGUGAAAUUGAGAGCAUUUU-UGUGCACACUUAUGGUCUGGAGACUAUAGU ------------....(---(((..((((((((((.....(((((((....))))))).((((..((((((....).)))))-..)))).)))...))))))).))))..... ( -26.60) >DroYak_CAF1 61662 107 - 1 UGGAAUUAUGCUGGCAA---GUUCAUCAAGCUGGUAAAUACUGUAAGAGUUCUUAUUUUUGCACAGAAAUUGAGGGGAUUUU-UAUGCAUAUGUAUGGUAUGGAG--CAUAGU ....((((((((...((---((((.((((.((((((((....(((((....))))).))))).)))...))))..))))))(-(((((....)))))).....))--)))))) ( -26.90) >consensus UGGAAUUAUGCUGGCAG___GUUCAUCAAGCUGGUAAGAAGUGUAAGAGUUCUUACUUAUCCAGUGAAAUGGAGAGAAUUUUAUAUACACCUGUAUGGUAUGGAG__CAUAAU .....................(((((...((((....((.(((((((....))))))).))))))...)))))........................................ ( -8.08 = -9.32 + 1.24)


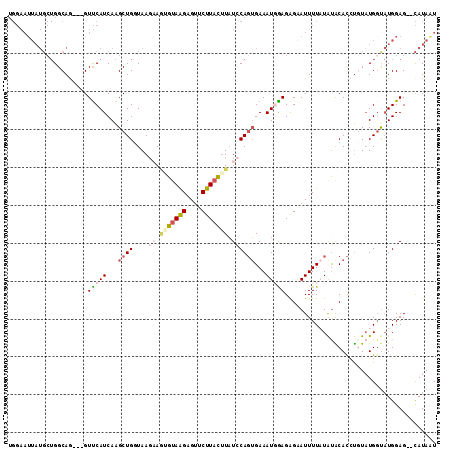
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:21:55 2006