
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 18,835,647 – 18,835,767 |
| Length | 120 |
| Max. P | 0.985535 |

| Location | 18,835,647 – 18,835,767 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 83.73 |
| Mean single sequence MFE | -41.16 |
| Consensus MFE | -29.92 |
| Energy contribution | -30.80 |
| Covariance contribution | 0.88 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.78 |
| Structure conservation index | 0.73 |
| SVM decision value | 2.01 |
| SVM RNA-class probability | 0.985535 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 18835647 120 + 22407834 AGGCGCCAUCUAGCGGCAACCCGCGCAGAUAGACUUGUGAAUGAAAGAAUAGAUGGAUAUAUCGUAACAUCGAUUGUAACGUGUAAGUGCGCCAUCUAUAGGCGAGUCAUAUUUGGCGCG ..(((((((((.((((....))))..)))).(((((((...(....).((((((((.((((.(((.(((.....))).)))))))......))))))))..)))))))......))))). ( -42.20) >DroSim_CAF1 2998 120 + 1 AGGCGCCAUCUAGCGGCAACCCGCGCAGCGACACUUGUGAAUGAAAGAAUAGAUGGAUAUAUCGUAACAUCGAUUGUAACGUGUAAUUGCGCCAUCUUUAGGCGAGUUCUAUUUGGCGCG ..((((((..((((((....))))((((..(((((((..(.(((.......((((....))))......))).)..))).))))..))))(((.......)))......))..)))))). ( -36.82) >DroEre_CAF1 3070 118 + 1 UUGCGCCACCUAGCGGCAACCCGCGUAGCGAGG-UUGUUAAUGAAA-AAUCGAUGGAAAUAUCGAAACAUCGAUUUCAGCAUUGAAGUGCGCCAUCUAUGGGCGAUACUUAUCCGGCGCG ..(((((.....((((((((((((...))).))-)))))..((((.-.(((((((............)))))))))))))....(((((((((.......)))).)))))....))))). ( -45.30) >DroYak_CAF1 3100 119 + 1 AGGCGCCACCUAGCGGCAACCCGCGUACCAAGACCUGUGAAUAAAA-AAUCGAUGGAAAUAUCGAAACAUCGAUUUCAGCACUUAAGUGCGCCAUCUAUAGGCGAUACUUAUUCGGCGCG ..(((((.....((((....))))..............((((((.(-((((((((............)))))))))..((((....))))(((.......))).....))))))))))). ( -40.30) >consensus AGGCGCCACCUAGCGGCAACCCGCGCAGCGAGACUUGUGAAUGAAA_AAUAGAUGGAAAUAUCGAAACAUCGAUUGCAACAUGUAAGUGCGCCAUCUAUAGGCGAGACUUAUUCGGCGCG ..(((((.....((((....)))).............((((((....((((((((............))))))))....))))))....((((.......))))..........))))). (-29.92 = -30.80 + 0.88)

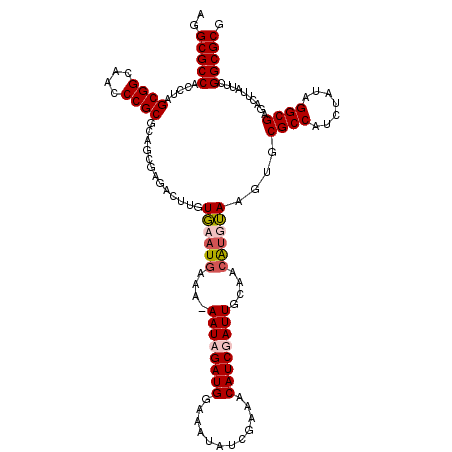
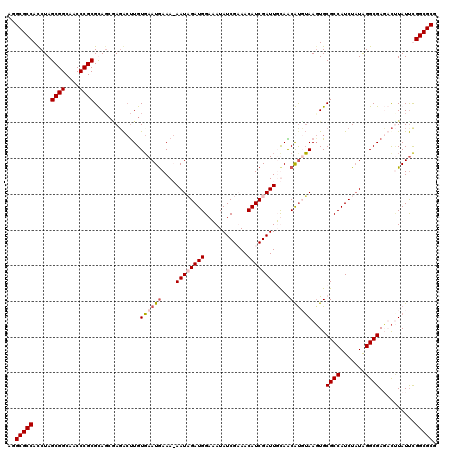
| Location | 18,835,647 – 18,835,767 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 83.73 |
| Mean single sequence MFE | -37.00 |
| Consensus MFE | -27.58 |
| Energy contribution | -27.07 |
| Covariance contribution | -0.50 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.92 |
| Structure conservation index | 0.75 |
| SVM decision value | 0.74 |
| SVM RNA-class probability | 0.838750 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 18835647 120 - 22407834 CGCGCCAAAUAUGACUCGCCUAUAGAUGGCGCACUUACACGUUACAAUCGAUGUUACGAUAUAUCCAUCUAUUCUUUCAUUCACAAGUCUAUCUGCGCGGGUUGCCGCUAGAUGGCGCCU .(((((......((((((((.......))))........(((.(((.....))).)))...........................)))).(((((.((((....)))))))))))))).. ( -33.20) >DroSim_CAF1 2998 120 - 1 CGCGCCAAAUAGAACUCGCCUAAAGAUGGCGCAAUUACACGUUACAAUCGAUGUUACGAUAUAUCCAUCUAUUCUUUCAUUCACAAGUGUCGCUGCGCGGGUUGCCGCUAGAUGGCGCCU .((((((..((((((((((....((.((((((........((.(((.....))).))(((......))).................))))))))..)))))))....)))..)))))).. ( -32.10) >DroEre_CAF1 3070 118 - 1 CGCGCCGGAUAAGUAUCGCCCAUAGAUGGCGCACUUCAAUGCUGAAAUCGAUGUUUCGAUAUUUCCAUCGAUU-UUUCAUUAACAA-CCUCGCUACGCGGGUUGCCGCUAGGUGGCGCAA .(((((.(((....)))(((..(((.((((.......((((..((((((((((............))))))))-)).))))...((-((.(((...)))))))))))))))))))))).. ( -39.90) >DroYak_CAF1 3100 119 - 1 CGCGCCGAAUAAGUAUCGCCUAUAGAUGGCGCACUUAAGUGCUGAAAUCGAUGUUUCGAUAUUUCCAUCGAUU-UUUUAUUCACAGGUCUUGGUACGCGGGUUGCCGCUAGGUGGCGCCU .((((((..((.(((((((((...(((((.((((....))))....(((((....)))))....)))))((..-......))..))))...)))))((((....))))))..)))))).. ( -42.80) >consensus CGCGCCAAAUAAGAAUCGCCUAUAGAUGGCGCACUUAAACGCUAAAAUCGAUGUUACGAUAUAUCCAUCGAUU_UUUCAUUCACAAGUCUCGCUACGCGGGUUGCCGCUAGAUGGCGCCU .((((((.................(((((.((........))....((((......))))....))))).......................(((.((((....))))))).)))))).. (-27.58 = -27.07 + -0.50)

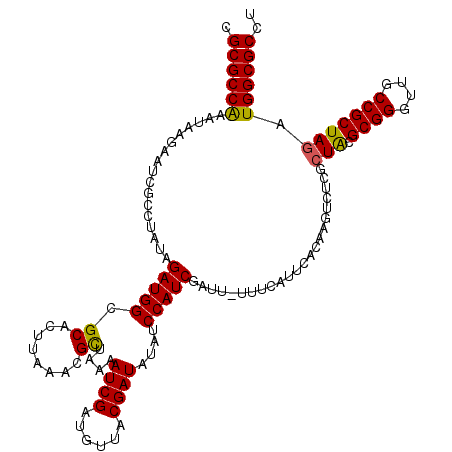

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:18:51 2006