
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 18,748,481 – 18,748,581 |
| Length | 100 |
| Max. P | 0.999200 |

| Location | 18,748,481 – 18,748,581 |
|---|---|
| Length | 100 |
| Sequences | 6 |
| Columns | 103 |
| Reading direction | forward |
| Mean pairwise identity | 92.28 |
| Mean single sequence MFE | -38.65 |
| Consensus MFE | -34.86 |
| Energy contribution | -35.25 |
| Covariance contribution | 0.39 |
| Combinations/Pair | 1.07 |
| Mean z-score | -3.00 |
| Structure conservation index | 0.90 |
| SVM decision value | 3.43 |
| SVM RNA-class probability | 0.999200 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 18748481 100 + 22407834 UUCAAGGCGGUGCUGGGUGGUGCAGCCUCUGCGGUUGCGGCGCGAAAUGGCUAAAAGCAAAUUGACGCCGCAACAAAUGCCAGA---GCAGCAAAAGCAAAGC ......((..(((((..((((((((...)))).((((((((((((....((.....))...))).)))))))))....))))..---.)))))...))..... ( -39.60) >DroSec_CAF1 12071 100 + 1 UUCAAGGCGGUGCUGGGUGGUGCAGCCUCUGCGGUUGCGGCGCGAAAUGGCUAAAAGCAAAUUGACGCCGCAACAAAUGCCAGA---GCAGCAAAAGCAAUGC ......((..(((((..((((((((...)))).((((((((((((....((.....))...))).)))))))))....))))..---.)))))...))..... ( -39.60) >DroSim_CAF1 12291 100 + 1 UUCAGGGCGGUGCUGGGUGGUGCAGCCUCUGCGGUUGCGGCGCGAAAUGGCUAAAAGCAAAUUGACGCCGCAACAAAUGCCAGA---GCAGCAAAAGCAAUGC ..((..((..(((((..((((((((...)))).((((((((((((....((.....))...))).)))))))))....))))..---.)))))...))..)). ( -40.70) >DroEre_CAF1 12420 100 + 1 UUCAAGGCGGUGCUGGGUGGUGCAGCCUCUGCGGUUGCGGCGCGAAAUGGCUAAAAGCAAAUUGACGCCGCAACAAAUGCCAGA---GCAGCAACAGCAAAGC ......((..(((((.....(((.((.(((((.((((((((((((....((.....))...))).)))))))))....).))))---)).))).)))))..)) ( -40.50) >DroYak_CAF1 12108 103 + 1 UUCAAGGCGGUGCUGGGUGGUGCAGCCUCUGCGGUUGCGGCGCGAAAUGGCUAAAAGCAAAUUGACGCCGCAACAAAUGCCAGAUUUGCAGCAAAAGCAAAGC ......((..((((...((.(((((..(((((.((((((((((((....((.....))...))).)))))))))....).)))).))))).))..))))..)) ( -40.90) >DroAna_CAF1 9661 90 + 1 ------GUGGU----GGUGGUGCAGUCUCUGCGGUUGUGGCGCGAAAUGGCUUAAAGCAAAUUGACGCCGCAACAAAUGCCAAA---GCAGAAAAGGCAAAGC ------((..(----(((..(((((...)))))((((((((((((....((.....))...))).)))))))))....))))..---)).............. ( -30.60) >consensus UUCAAGGCGGUGCUGGGUGGUGCAGCCUCUGCGGUUGCGGCGCGAAAUGGCUAAAAGCAAAUUGACGCCGCAACAAAUGCCAGA___GCAGCAAAAGCAAAGC ......((..(((((..((((((((...)))).((((((((((((....((.....))...))).)))))))))....))))......)))))...))..... (-34.86 = -35.25 + 0.39)

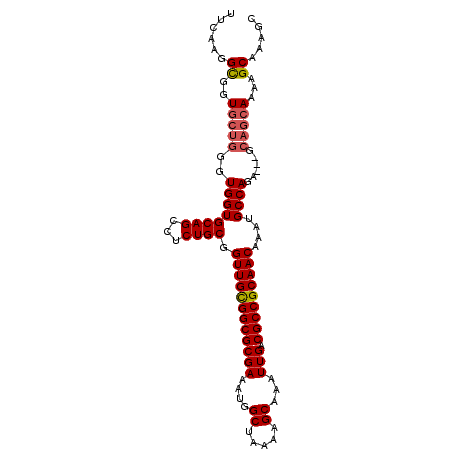
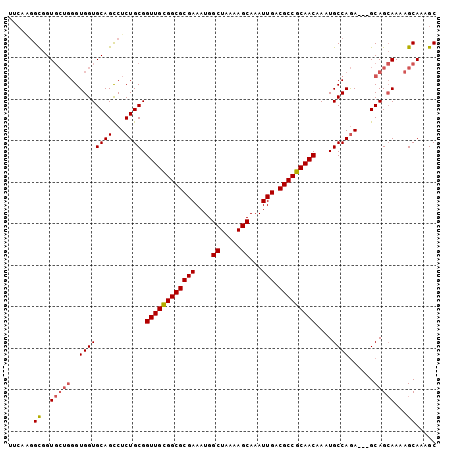
| Location | 18,748,481 – 18,748,581 |
|---|---|
| Length | 100 |
| Sequences | 6 |
| Columns | 103 |
| Reading direction | reverse |
| Mean pairwise identity | 92.28 |
| Mean single sequence MFE | -34.58 |
| Consensus MFE | -30.70 |
| Energy contribution | -31.23 |
| Covariance contribution | 0.53 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.67 |
| Structure conservation index | 0.89 |
| SVM decision value | 2.68 |
| SVM RNA-class probability | 0.996324 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 18748481 100 - 22407834 GCUUUGCUUUUGCUGC---UCUGGCAUUUGUUGCGGCGUCAAUUUGCUUUUAGCCAUUUCGCGCCGCAACCGCAGAGGCUGCACCACCCAGCACCGCCUUGAA ((...(((((((((((---....)))...((((((((((.((...((.....))...)).)))))))))).)))))))).))..................... ( -36.20) >DroSec_CAF1 12071 100 - 1 GCAUUGCUUUUGCUGC---UCUGGCAUUUGUUGCGGCGUCAAUUUGCUUUUAGCCAUUUCGCGCCGCAACCGCAGAGGCUGCACCACCCAGCACCGCCUUGAA ((...(((((((((((---....)))...((((((((((.((...((.....))...)).)))))))))).))))))))(((........)))..))...... ( -36.70) >DroSim_CAF1 12291 100 - 1 GCAUUGCUUUUGCUGC---UCUGGCAUUUGUUGCGGCGUCAAUUUGCUUUUAGCCAUUUCGCGCCGCAACCGCAGAGGCUGCACCACCCAGCACCGCCCUGAA ((...(((((((((((---....)))...((((((((((.((...((.....))...)).)))))))))).))))))))(((........)))..))...... ( -36.70) >DroEre_CAF1 12420 100 - 1 GCUUUGCUGUUGCUGC---UCUGGCAUUUGUUGCGGCGUCAAUUUGCUUUUAGCCAUUUCGCGCCGCAACCGCAGAGGCUGCACCACCCAGCACCGCCUUGAA ((..(((((.(((.((---((((.(....((((((((((.((...((.....))...)).)))))))))).))))).)).))).....)))))..))...... ( -36.40) >DroYak_CAF1 12108 103 - 1 GCUUUGCUUUUGCUGCAAAUCUGGCAUUUGUUGCGGCGUCAAUUUGCUUUUAGCCAUUUCGCGCCGCAACCGCAGAGGCUGCACCACCCAGCACCGCCUUGAA ((...(((((((((((.......)))...((((((((((.((...((.....))...)).)))))))))).)))))))).))..................... ( -35.50) >DroAna_CAF1 9661 90 - 1 GCUUUGCCUUUUCUGC---UUUGGCAUUUGUUGCGGCGUCAAUUUGCUUUAAGCCAUUUCGCGCCACAACCGCAGAGACUGCACCACC----ACCAC------ ....(((..(((((((---.((((((.....)))(((((.((...((.....))...)).))))).)))..)))))))..))).....----.....------ ( -26.00) >consensus GCUUUGCUUUUGCUGC___UCUGGCAUUUGUUGCGGCGUCAAUUUGCUUUUAGCCAUUUCGCGCCGCAACCGCAGAGGCUGCACCACCCAGCACCGCCUUGAA ((...(((((((((((.......)))...((((((((((.((...((.....))...)).)))))))))).)))))))).))..................... (-30.70 = -31.23 + 0.53)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:18:15 2006