
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 18,728,214 – 18,728,374 |
| Length | 160 |
| Max. P | 0.996235 |

| Location | 18,728,214 – 18,728,334 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 97.77 |
| Mean single sequence MFE | -26.03 |
| Consensus MFE | -25.39 |
| Energy contribution | -25.50 |
| Covariance contribution | 0.11 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.16 |
| Structure conservation index | 0.98 |
| SVM decision value | 1.16 |
| SVM RNA-class probability | 0.924228 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 18728214 120 + 22407834 UUUACACAUAACAUAUAUUUUGUAUUUAGGAUGUAUCCUUAAGCGAUCCUAUCUUCACUUCACUAUAAAACGGUAGCGCUUCAUUUUGAAUGUUUUACCGUUAUCACCGAUCGCAUGGCG ......(((...((((((((((....))))))))))......((((((.(((............))).((((((((.(((((.....))).)).))))))))......)))))))))... ( -23.80) >DroSec_CAF1 20776 119 + 1 UUAACACAUAACAUAUAUUUUGUAUUUAGGAUGUAUCCUUAAGCGAUCCUAUCUUCACUUCACUGUA-AACGGUAGCGCUUCAUUUUGAAUGUUUUACCGUUAUCACCGAUCGCAUGGCG ......(((...((((((((((....))))))))))......((((((........((......)).-((((((((.(((((.....))).)).))))))))......)))))))))... ( -27.00) >DroSim_CAF1 19253 119 + 1 UUUACACAUAACAUAUAUUUUGUAUUUAGGAUGUAUCCUUAAGCGAUCCUAUCUUCACUUCACUGUA-AACGGUAACGCUUCAUUUUGAAUGUUUUACCGUUAUCACCGAUCGCAUGGCG ......(((...((((((((((....))))))))))......((((((........((......)).-((((((((.(((((.....))).)).))))))))......)))))))))... ( -27.30) >consensus UUUACACAUAACAUAUAUUUUGUAUUUAGGAUGUAUCCUUAAGCGAUCCUAUCUUCACUUCACUGUA_AACGGUAGCGCUUCAUUUUGAAUGUUUUACCGUUAUCACCGAUCGCAUGGCG ......(((...((((((((((....))))))))))......((((((........((......))..((((((((.(((((.....))).)).))))))))......)))))))))... (-25.39 = -25.50 + 0.11)

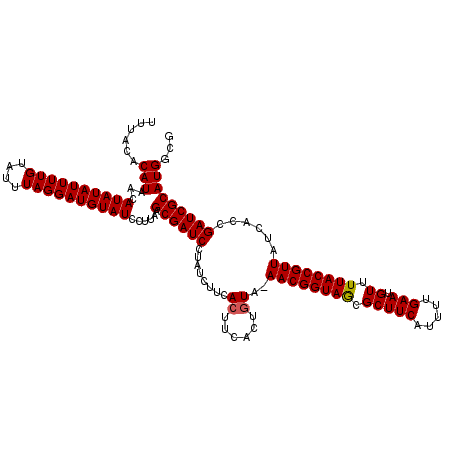
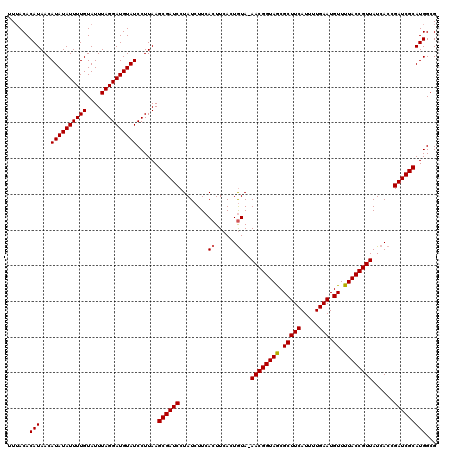
| Location | 18,728,254 – 18,728,374 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 96.22 |
| Mean single sequence MFE | -28.68 |
| Consensus MFE | -28.16 |
| Energy contribution | -28.52 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.48 |
| Structure conservation index | 0.98 |
| SVM decision value | 1.80 |
| SVM RNA-class probability | 0.977666 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 18728254 120 + 22407834 AAGCGAUCCUAUCUUCACUUCACUAUAAAACGGUAGCGCUUCAUUUUGAAUGUUUUACCGUUAUCACCGAUCGCAUGGCGUUACGUCUUACGACCGCUUUACGUAGUGGAUCUCUUCUGC (((.(((((............(((((..((((((((.(((((.....))).)).)))))))).....((.(((...((((...))))...))).))......)))))))))).))).... ( -27.30) >DroSec_CAF1 20816 119 + 1 AAGCGAUCCUAUCUUCACUUCACUGUA-AACGGUAGCGCUUCAUUUUGAAUGUUUUACCGUUAUCACCGAUCGCAUGGCGUUACGUCUUACGACCGCUUUACGUAGUGGAUCUCUUCUGC ..((((((........((......)).-((((((((.(((((.....))).)).))))))))......))))))..((((...))))....(((((((......))))).))........ ( -30.30) >DroSim_CAF1 19293 119 + 1 AAGCGAUCCUAUCUUCACUUCACUGUA-AACGGUAACGCUUCAUUUUGAAUGUUUUACCGUUAUCACCGAUCGCAUGGCGUUACGUCUUACGACCGCUUUACGUAGUGGAUCUCUUCUGC ..((((((........((......)).-((((((((.(((((.....))).)).))))))))......))))))..((((...))))....(((((((......))))).))........ ( -30.60) >DroEre_CAF1 21040 116 + 1 -AGCGAUCCUAUCUUCACUUCACUGUA-AACGGUAACGCUUCAUUUUGAAUGUUUCACCGUUAUCACCGAUCGCAUGGCGUUACGUCU--CGACCGCUUUACGUAGUGGAUCUCUUCUGC -.((((((........((......)).-((((((((((.(((.....)))))))..))))))......))))))..((((...)))).--.(((((((......))))).))........ ( -27.60) >DroYak_CAF1 21060 117 + 1 AAGCGAUCCUAUCUUCACUUCACUGUA-AACGGUAACGCUUCAUUUUGAAUGUUUAACCGUUAUCAUCGAUCGCAUGGCGUUACGUCU--CGACCGCUUUACGUAGUGGAUCUCUUCUGU ..((((((........((......)).-((((((((((.(((.....)))))))..))))))......))))))..((((...)))).--.(((((((......))))).))........ ( -27.60) >consensus AAGCGAUCCUAUCUUCACUUCACUGUA_AACGGUAACGCUUCAUUUUGAAUGUUUUACCGUUAUCACCGAUCGCAUGGCGUUACGUCUUACGACCGCUUUACGUAGUGGAUCUCUUCUGC ..((((((........((......))..((((((((.(((((.....))).)).))))))))......))))))..((((...))))....(((((((......))))).))........ (-28.16 = -28.52 + 0.36)

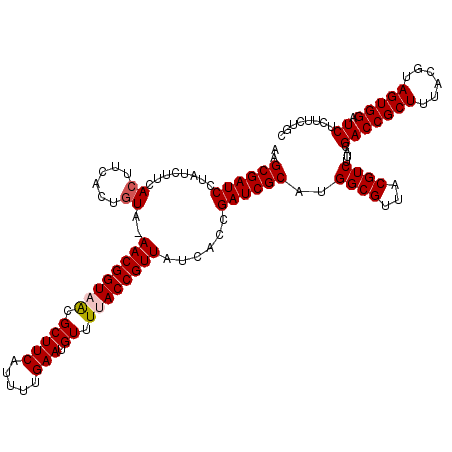
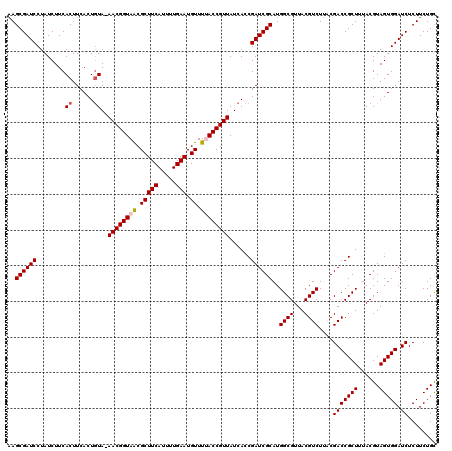
| Location | 18,728,254 – 18,728,374 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 96.22 |
| Mean single sequence MFE | -32.30 |
| Consensus MFE | -32.24 |
| Energy contribution | -32.76 |
| Covariance contribution | 0.52 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.70 |
| Structure conservation index | 1.00 |
| SVM decision value | 2.67 |
| SVM RNA-class probability | 0.996235 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 18728254 120 - 22407834 GCAGAAGAGAUCCACUACGUAAAGCGGUCGUAAGACGUAACGCCAUGCGAUCGGUGAUAACGGUAAAACAUUCAAAAUGAAGCGCUACCGUUUUAUAGUGAAGUGAAGAUAGGAUCGCUU ....(((.(((((....((((..((((((....)))....)))..))))((((.......))))...((.((((..((((((((....))))))))..)))))).......))))).))) ( -32.30) >DroSec_CAF1 20816 119 - 1 GCAGAAGAGAUCCACUACGUAAAGCGGUCGUAAGACGUAACGCCAUGCGAUCGGUGAUAACGGUAAAACAUUCAAAAUGAAGCGCUACCGUU-UACAGUGAAGUGAAGAUAGGAUCGCUU ....(((.(((((....((((..((((((....)))....)))..)))).(((.((..(((((((.....(((.....)))....)))))))-..)).)))..........))))).))) ( -33.70) >DroSim_CAF1 19293 119 - 1 GCAGAAGAGAUCCACUACGUAAAGCGGUCGUAAGACGUAACGCCAUGCGAUCGGUGAUAACGGUAAAACAUUCAAAAUGAAGCGUUACCGUU-UACAGUGAAGUGAAGAUAGGAUCGCUU ....(((.(((((....((((..((((((....)))....)))..)))).(((.((..((((((((....(((.....)))...))))))))-..)).)))..........))))).))) ( -35.90) >DroEre_CAF1 21040 116 - 1 GCAGAAGAGAUCCACUACGUAAAGCGGUCG--AGACGUAACGCCAUGCGAUCGGUGAUAACGGUGAAACAUUCAAAAUGAAGCGUUACCGUU-UACAGUGAAGUGAAGAUAGGAUCGCU- .....((.(((((....((((..(((..((--...))...)))..)))).(((.((..((((((((....(((.....)))...))))))))-..)).)))..........))))).))- ( -31.60) >DroYak_CAF1 21060 117 - 1 ACAGAAGAGAUCCACUACGUAAAGCGGUCG--AGACGUAACGCCAUGCGAUCGAUGAUAACGGUUAAACAUUCAAAAUGAAGCGUUACCGUU-UACAGUGAAGUGAAGAUAGGAUCGCUU ....(((.(((((....((((..(((..((--...))...)))..)))).(((.((..((((((..(((.(((.....)))..)))))))))-..)).)))..........))))).))) ( -28.00) >consensus GCAGAAGAGAUCCACUACGUAAAGCGGUCGUAAGACGUAACGCCAUGCGAUCGGUGAUAACGGUAAAACAUUCAAAAUGAAGCGUUACCGUU_UACAGUGAAGUGAAGAUAGGAUCGCUU ....(((.(((((....((((..((((((....)))....)))..)))).(((.((..((((((((....(((.....)))...))))))))...)).)))..........))))).))) (-32.24 = -32.76 + 0.52)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:17:46 2006