
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 18,635,556 – 18,635,661 |
| Length | 105 |
| Max. P | 0.774938 |

| Location | 18,635,556 – 18,635,661 |
|---|---|
| Length | 105 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 82.29 |
| Mean single sequence MFE | -32.55 |
| Consensus MFE | -18.91 |
| Energy contribution | -20.63 |
| Covariance contribution | 1.72 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.77 |
| Structure conservation index | 0.58 |
| SVM decision value | 0.54 |
| SVM RNA-class probability | 0.774938 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 18635556 105 + 22407834 GCAUGAACAAGGCGAAAGCCGUCCACUCGAAUGGGCCUUAUAUAUU--AUUUGCCCAUAUCUA--------UAUCUUUGAUCGCUUCUA-----UGCACAUUUCUGUGUAGAGGCGUUGU ....((.((((((....)))........((((((((..((.....)--)...)))))).))..--------.....))).))(((((((-----((((......)))))))))))..... ( -33.20) >DroSec_CAF1 108434 102 + 1 GCAUGAACAAGGCGAAAGCCGUCCACUCGAAUGGGCCUUAUAUA-----UUUGCCCAUAUCUA--------UAUCUUUGAUCGCUUCUA-----UGCACAUUUCUGUGUAGAGGCGUUGU ....((.((((((....)))........((((((((........-----...)))))).))..--------.....))).))(((((((-----((((......)))))))))))..... ( -32.90) >DroSim_CAF1 110007 102 + 1 GCAUGAACAAGGCGAAAGCCGUCCACUCGAAUGGGCCUUAUAUA-----UUUGCCCAUAUCUA--------UAUCUUUGAUCGCUUCUA-----UGCACAUUUCUGUGUAGAGGCGUUGU ....((.((((((....)))........((((((((........-----...)))))).))..--------.....))).))(((((((-----((((......)))))))))))..... ( -32.90) >DroEre_CAF1 107350 102 + 1 GCAUGAACAAGGCGAAAGCCGUCCACUCGAAUGGGCCUUAUAUA-----UUUGCCCAUAUCUA--------UAUCUUUGAUCGCUUCUA-----UGCACAUUUCUGUGUAGAGGCGUUGU ....((.((((((....)))........((((((((........-----...)))))).))..--------.....))).))(((((((-----((((......)))))))))))..... ( -32.90) >DroYak_CAF1 108764 107 + 1 GCAUGAACAAGGCGAAAGCCGUCCACUCGAAUGGGCCUUAUAUA-----UUUGCCCAUAUCUA--------UAUCUUUGAUCGCUUCUAUGCUAUGCACAUUUCUGUGUAGAGGCGUUGU ....((.((((((....)))........((((((((........-----...)))))).))..--------.....))).))((((((((((.............))))))))))..... ( -31.22) >DroAna_CAF1 98580 97 + 1 GCACGAACAAGGCGAAAG-------------UGGGC-UUUUAUGUUGGAUAUGC-CGGGUCUAGGCAGGGGUAUAUGUGAGCAUUUCUA-----UGCCCAUUUCAGUCUUUGG---CUGG ((.(((...((((..(((-------------(((((-....(((((.(((((((-(..((....))...))))))).).))))).....-----.))))))))..))))))))---)... ( -32.20) >consensus GCAUGAACAAGGCGAAAGCCGUCCACUCGAAUGGGCCUUAUAUA_____UUUGCCCAUAUCUA________UAUCUUUGAUCGCUUCUA_____UGCACAUUUCUGUGUAGAGGCGUUGU ....((.((((((....)))..........((((((................))))))..................))).))((((((.......((((......))))))))))..... (-18.91 = -20.63 + 1.72)

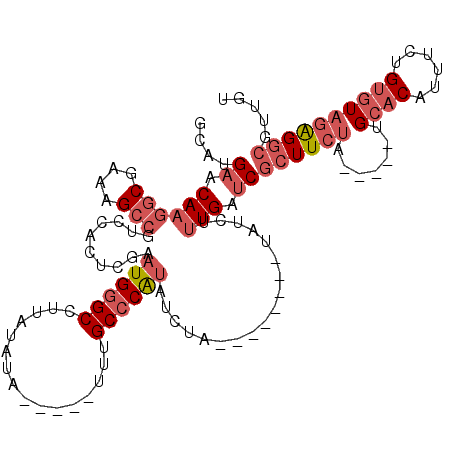
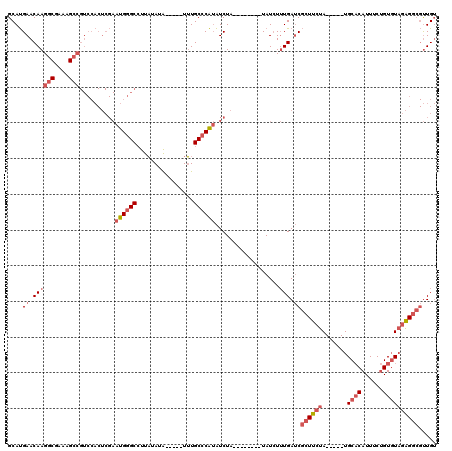
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:16:03 2006