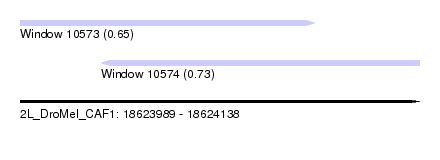

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 18,623,989 – 18,624,138 |
| Length | 149 |
| Max. P | 0.734719 |

| Location | 18,623,989 – 18,624,099 |
|---|---|
| Length | 110 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 87.10 |
| Mean single sequence MFE | -34.42 |
| Consensus MFE | -25.58 |
| Energy contribution | -25.32 |
| Covariance contribution | -0.25 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.61 |
| Structure conservation index | 0.74 |
| SVM decision value | 0.24 |
| SVM RNA-class probability | 0.647314 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
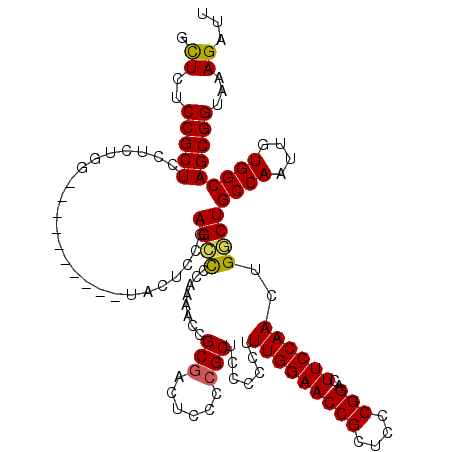
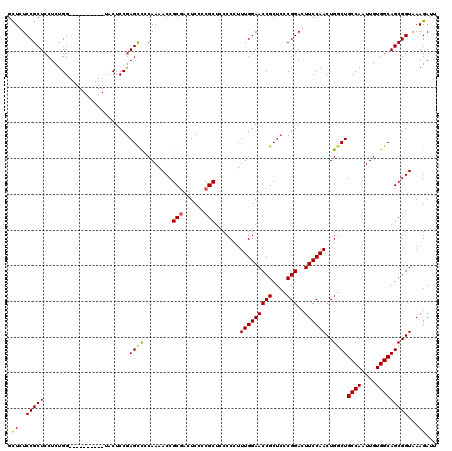
>2L_DroMel_CAF1 18623989 110 + 22407834 GCUGUCCGCUCCUCUGG----------UCCUCCGAGUUGCAAAACCGCGACUCCCCGCACCCCUUUUGGAACCGCUCCCGGACUUCCAACUGACUGCCAAUUGUGGCAGCGGUAAAGAUU ...(((((.......((----------(.(...(((((((......)))))))...).)))......(((.....)))))))).....((((.((((((....))))))))))....... ( -34.60) >DroSec_CAF1 101535 110 + 1 GCUGUCCGCUCCUCUGG----------UCCUCCGAGCUGCAAAACCGCGAAUCCCCGCUCCCCUUUUGGAACCGCUCGCGGACUUCCAACUGCCUGCCAAUUGUGGCAGCGGUAAAGAUU ...((((((......((----------....))((((.........(((......)))(((......)))...)))))))))).....(((((.(((((....))))))))))....... ( -37.90) >DroEre_CAF1 100527 120 + 1 GUUCUCCGCUCCUCUGGUAUUCUCUAGUACUCCGAGCCCCAAAACCGCGACUCCCCGCUGCCCCUUUGGAACCGCUCCCGGACUUCCAACUGGCUGCCAAUUGUGGCAGCGGUAAAGAUU .......((((....(((((......)))))..)))).........(((......)))((((...(((((((((....)))..))))))...(((((((....)))))))))))...... ( -34.70) >DroYak_CAF1 101981 109 + 1 GUUCUCCGCUCCUCUGG----------AACUCCAAGCCCCAAAACCGCUACUCUCCGCUCCCCC-UUGGAACCGCUCCCGGACUUCCAACUGGCUGCCAAUUGUGGCAGCGGUAAAGAUU ..((((((((....(((----------....)))............(((((.....((..((..-(((((((((....)))..))))))..))..)).....))))))))))...))).. ( -30.50) >consensus GCUCUCCGCUCCUCUGG__________UACUCCGAGCCCCAAAACCGCGACUCCCCGCUCCCCCUUUGGAACCGCUCCCGGACUUCCAACUGGCUGCCAAUUGUGGCAGCGGUAAAGAUU .((..(((((........................((((........(((......))).......(((((((((....)))..))))))..))))((((....)))))))))...))... (-25.58 = -25.32 + -0.25)

| Location | 18,624,019 – 18,624,138 |
|---|---|
| Length | 119 |
| Sequences | 4 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 95.38 |
| Mean single sequence MFE | -47.90 |
| Consensus MFE | -43.91 |
| Energy contribution | -44.10 |
| Covariance contribution | 0.19 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.65 |
| Structure conservation index | 0.92 |
| SVM decision value | 0.43 |
| SVM RNA-class probability | 0.734719 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
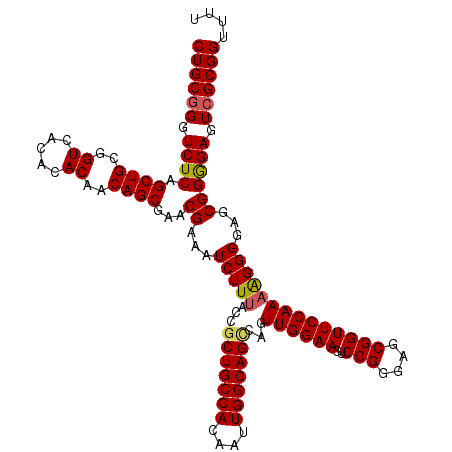
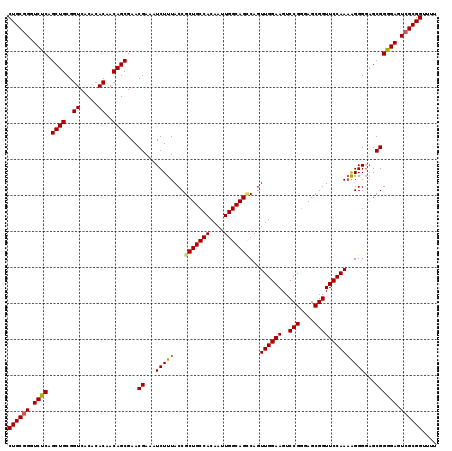
>2L_DroMel_CAF1 18624019 119 - 22407834 CUGCGGGUCUCAGCUGCGGUCACACACAACAGCGAACGAAAUCUUUACCGCUGCCACAAUUGGCAGUCAGUUGGAAGUCCGGGAGCGGUUCCAAAAGGGGUGCGGGGAGUCGCGGUUUU ((((((.((((.((((..((.....))..))))...((...(((((...(((((((....)))))))...((((((..(((....))))))))))))))...)))))).)))))).... ( -46.30) >DroSec_CAF1 101565 119 - 1 CUGCGGGUCUCAGCUGCGGUCACACACAACAGCGAACGAAAUCUUUACCGCUGCCACAAUUGGCAGGCAGUUGGAAGUCCGCGAGCGGUUCCAAAAGGGGAGCGGGGAUUCGCGGUUUU (((((((((((.(((((((.(.....((((.((((......)).......((((((....)))))))).))))...).)))).))).(((((......))))).))))))))))).... ( -48.70) >DroEre_CAF1 100567 119 - 1 CUGCGGGUCUCGGCUGCGGUCACACACAACAGCGAACGAAAUCUUUACCGCUGCCACAAUUGGCAGCCAGUUGGAAGUCCGGGAGCGGUUCCAAAGGGGCAGCGGGGAGUCGCGGUUUU ((((((.((((.((((((.((.(........).)).)..........(((((((((....)))))))...((((((..(((....))))))))).)).))))).)))).)))))).... ( -51.20) >DroYak_CAF1 102011 118 - 1 CUGCGGGUCUCAGCUGCGGUCACACACAACAGCGAACGAAAUCUUUACCGCUGCCACAAUUGGCAGCCAGUUGGAAGUCCGGGAGCGGUUCCAA-GGGGGAGCGGAGAGUAGCGGUUUU ((((.(.((((.((((..((.....))..))))...((...(((...(((((((((....)))))))...((((((..(((....)))))))))-)).))).)))))).).)))).... ( -45.40) >consensus CUGCGGGUCUCAGCUGCGGUCACACACAACAGCGAACGAAAUCUUUACCGCUGCCACAAUUGGCAGCCAGUUGGAAGUCCGGGAGCGGUUCCAAAAGGGGAGCGGGGAGUCGCGGUUUU ((((((.((((.((((..((.....))..))))...((...(((((...(((((((....)))))))...((((((..(((....))))))))))))))...)))))).)))))).... (-43.91 = -44.10 + 0.19)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:15:55 2006