
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 18,520,754 – 18,520,870 |
| Length | 116 |
| Max. P | 0.860808 |

| Location | 18,520,754 – 18,520,870 |
|---|---|
| Length | 116 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 80.58 |
| Mean single sequence MFE | -22.76 |
| Consensus MFE | -16.25 |
| Energy contribution | -16.62 |
| Covariance contribution | 0.38 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.66 |
| Structure conservation index | 0.71 |
| SVM decision value | 0.28 |
| SVM RNA-class probability | 0.666925 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 18520754 116 + 22407834 -AAUAAAAAAGAAAUGCUAACUAUUGAUGUCAUAAUGUUGUUAAAUAACGUAUAUGCGGCAUGUGUCGUGCAUUUCUAUUUGCUUU---UUAACGCUAUUUUAUUUUUUUUUUUUGUUUU -((((((((((((((((.................(((((((...)))))))....(((((....)))))))))))))....((...---.....))..............)))))))).. ( -20.90) >DroPse_CAF1 49817 96 + 1 CAGCAAAGAAGAAAUGAUAACUAUUGGUGGCAUAAUGCUGUUAAAUAACGUAUACGCCGCAUGCGCCGUGCAUUUCUAUUUGCGGUUU--------------------CGUUUUUG---- ..(((((..(((((((..........(((((...((((.(((....)))))))..)))))..........))))))).))))).....--------------------........---- ( -23.95) >DroSec_CAF1 41145 112 + 1 -AAUAAAAAAGAAAUGCUAACUAUUGAUGUCAUAAUGUUGUUAAAUAACGUAUAUGCGGCAUGUGCCGUGCAUUUCUAUUUGCUUUUUUUUAACGCC-------UUUUUUUUUUUGUUUU -(((((((((((((.((.(((...((....))....)))((((((..........(((((....)))))(((........))).....)))))))).-------.))))))))))))).. ( -23.40) >DroSim_CAF1 42061 107 + 1 -AAUAAAAAAGAAAUGCUAACUAUUGAUGUCAUAAUGUUGUUAAAUAACGUAUAUGCGGCAUGUGCCGUGCAUUUCUAUUUGCUUU--UUUAACGCU----------UUUUUUUUGUUUU -((((((((((((((((.................(((((((...)))))))....(((((....)))))))))))))....((...--......)).----------...)))))))).. ( -22.80) >consensus _AAUAAAAAAGAAAUGCUAACUAUUGAUGUCAUAAUGUUGUUAAAUAACGUAUAUGCGGCAUGUGCCGUGCAUUUCUAUUUGCUUU___UUAACGCU__________UUUUUUUUGUUUU .....(((.((((((((.................(((((((...)))))))....(((((....))))))))))))).)))....................................... (-16.25 = -16.62 + 0.38)

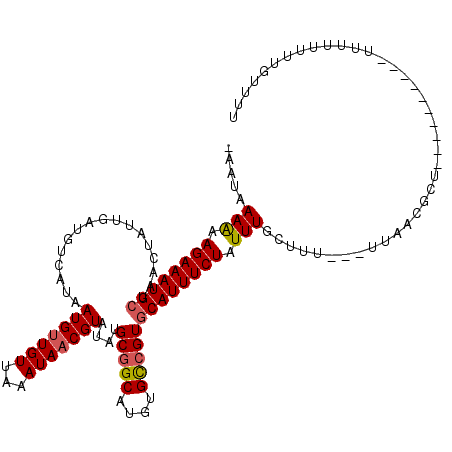
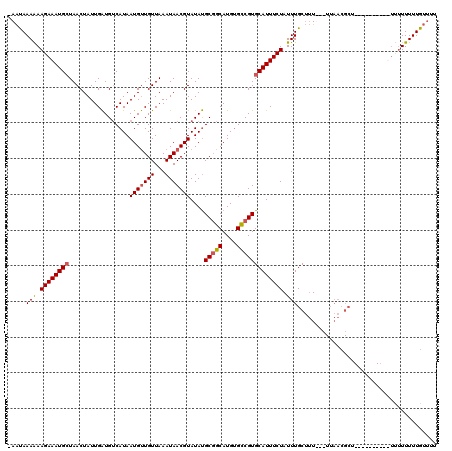
| Location | 18,520,754 – 18,520,870 |
|---|---|
| Length | 116 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 80.58 |
| Mean single sequence MFE | -18.52 |
| Consensus MFE | -11.82 |
| Energy contribution | -12.20 |
| Covariance contribution | 0.38 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.08 |
| Structure conservation index | 0.64 |
| SVM decision value | 0.83 |
| SVM RNA-class probability | 0.860808 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 18520754 116 - 22407834 AAAACAAAAAAAAAAAAUAAAAUAGCGUUAA---AAAGCAAAUAGAAAUGCACGACACAUGCCGCAUAUACGUUAUUUAACAACAUUAUGACAUCAAUAGUUAGCAUUUCUUUUUUAUU- ...............(((((((..((.....---...))....((((((((.((........)).......(((((...........)))))...........)))))))).)))))))- ( -14.40) >DroPse_CAF1 49817 96 - 1 ----CAAAAACG--------------------AAACCGCAAAUAGAAAUGCACGGCGCAUGCGGCGUAUACGUUAUUUAACAGCAUUAUGCCACCAAUAGUUAUCAUUUCUUCUUUGCUG ----........--------------------.....(((((.(((((((.(..((....(.((((((...(((.......)))..)))))).).....))..))))))))..))))).. ( -21.10) >DroSec_CAF1 41145 112 - 1 AAAACAAAAAAAAAAA-------GGCGUUAAAAAAAAGCAAAUAGAAAUGCACGGCACAUGCCGCAUAUACGUUAUUUAACAACAUUAUGACAUCAAUAGUUAGCAUUUCUUUUUUAUU- ................-------.((...........))(((.((((((((.((((....)))).......(((((...........)))))...........)))))))).)))....- ( -19.10) >DroSim_CAF1 42061 107 - 1 AAAACAAAAAAAA----------AGCGUUAAA--AAAGCAAAUAGAAAUGCACGGCACAUGCCGCAUAUACGUUAUUUAACAACAUUAUGACAUCAAUAGUUAGCAUUUCUUUUUUAUU- .............----------.((......--...))(((.((((((((.((((....)))).......(((((...........)))))...........)))))))).)))....- ( -19.50) >consensus AAAACAAAAAAAA__________AGCGUUAA___AAAGCAAAUAGAAAUGCACGGCACAUGCCGCAUAUACGUUAUUUAACAACAUUAUGACAUCAAUAGUUAGCAUUUCUUUUUUAUU_ ...........................................((((((((.((((....))))((((...(((.......)))..)))).............))))))))......... (-11.82 = -12.20 + 0.38)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:15:14 2006