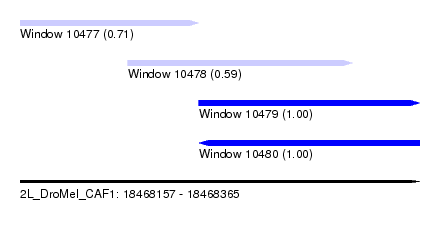

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 18,468,157 – 18,468,365 |
| Length | 208 |
| Max. P | 0.999021 |

| Location | 18,468,157 – 18,468,250 |
|---|---|
| Length | 93 |
| Sequences | 6 |
| Columns | 106 |
| Reading direction | forward |
| Mean pairwise identity | 81.73 |
| Mean single sequence MFE | -25.80 |
| Consensus MFE | -17.27 |
| Energy contribution | -17.83 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.27 |
| Mean z-score | -1.88 |
| Structure conservation index | 0.67 |
| SVM decision value | 0.38 |
| SVM RNA-class probability | 0.712144 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 18468157 93 + 22407834 --UGUUCUGAGCCAAACUAUUGUUAAAUAUCAUUACGAAUGCCA-----AA---GUAUGUAAAUGCAA---CCGGGCAUAUGUAAAUAUUGGCAUUGGUGACAUGU --(((((..((((((..((((....))))...(((((.(((((.-----..---((((....))))..---...))))).)))))...))))).)..).))))... ( -23.00) >DroSec_CAF1 27168 96 + 1 --UGUUCCGAGCCAGACUAUUGUUAAAUUUCAUUACGAAUGCCU-----AAUGCCUAUGUAAAUGCAA---CCGGGCAUAUGUAAAUAUUGGCAUUGGUGACAUGU --(((((((((((((((....)))........(((((.((((((-----..(((..........))).---..)))))).)))))....)))).)))).))))... ( -25.60) >DroSim_CAF1 22987 93 + 1 --UGUUCCGAGCCAGACUAUUGUUAAAUUUCAUUACGAAUGCCU-----AA---GUAUGUAAAUGCAA---CCGGGCGUAUGUAAAUAUUGGCAUUGGUGACAUGU --(((((((((((((((....)))........(((((.((((((-----..---((((....))))..---..)))))).)))))....)))).)))).))))... ( -25.80) >DroEre_CAF1 22469 94 + 1 --UGCUCUGAGCCGGACUCUCGUUCAUUUUCAUUACAAAUGCCU----UAA---GUAUGUAAAUGCAA---CCGGGCAUAUGUAAAUAUUGGCAUUGGUGACAUGU --........(((((((....)))).......(((((.((((((----...---((((....))))..---..)))))).))))).....)))..((....))... ( -25.10) >DroYak_CAF1 22391 94 + 1 --UGUUCCGAGCCGGACUUUUGUUCAAUUUCAUUACGAAUGCCU----UAA---GUAUGUAAAUGCAA---CCGGGCAUAUGUAAAUAUUGGCAUUGGUGACAUGU --(((((((((((((((....)))).......(((((.((((((----...---((((....))))..---..)))))).))))).....))).)))).))))... ( -28.60) >DroAna_CAF1 18448 100 + 1 GCUGUUGAGAACCAAACUC-CGUCCAGCUCGAU--CGAGAGCCGACAUUCG---CCAAGCAAAUGCAUCCGCCGGGCGCAUGUAAAUAUUGGUAUUGGUGACAUGU ((((((....(((((....-.(((..((((...--...)))).)))....(---((((.....(((((.(((...))).)))))....))))).))))))))).)) ( -26.70) >consensus __UGUUCCGAGCCAGACUAUUGUUAAAUUUCAUUACGAAUGCCU_____AA___GUAUGUAAAUGCAA___CCGGGCAUAUGUAAAUAUUGGCAUUGGUGACAUGU ..(((((((((((((((....)))........(((((.((((((..........((((....)))).......)))))).)))))....)))).)))).))))... (-17.27 = -17.83 + 0.56)


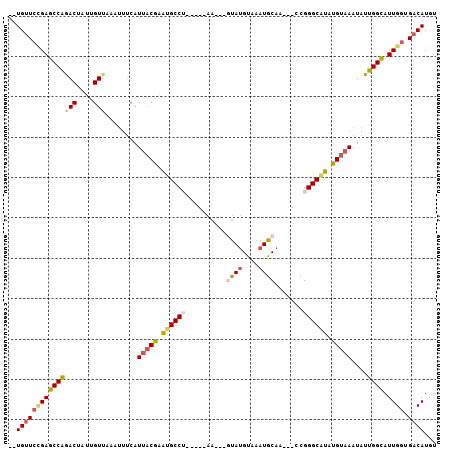
| Location | 18,468,213 – 18,468,330 |
|---|---|
| Length | 117 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 92.71 |
| Mean single sequence MFE | -33.23 |
| Consensus MFE | -26.27 |
| Energy contribution | -27.17 |
| Covariance contribution | 0.89 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.47 |
| Structure conservation index | 0.79 |
| SVM decision value | 0.11 |
| SVM RNA-class probability | 0.589452 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 18468213 117 + 22407834 AA---CCGGGCAUAUGUAAAUAUUGGCAUUGGUGACAUGUGCAAAUGUUUGUAUGGCUGAUUCCCUGAGACCCUAACUUGUGACUUUUAAUACCAGUUUCACAAGUUUUGAUCUCCGGUA .(---((((((.((((((((((((.((((((....)).)))).)))))))))))))))........((((.(..(((((((((..((.......))..)))))))))..).)))))))). ( -37.00) >DroGri_CAF1 27577 114 + 1 A------CUGCAUAUUUACACAUUGGUAUUAGUGACAUGUGCAAAUGUUUGUAUGGCUGAUUCCCUGAGACCCUAACUUGUGACUUUUAAUACCAGUUUCACAAGUUUUGAUCUCCGGUA .------...(((((....(((((.((((.........)))).)))))..)))))........((.((((.(..(((((((((..((.......))..)))))))))..).)))).)).. ( -24.90) >DroSec_CAF1 27227 117 + 1 AA---CCGGGCAUAUGUAAAUAUUGGCAUUGGUGACAUGUGCAAAUGUUUGUAUGGCUGAUUCCCUGAGACCCUAACUUGUGACUUUUAAUACCAGUUUCACAAGUUUUGAUCUCCGGUA .(---((((((.((((((((((((.((((((....)).)))).)))))))))))))))........((((.(..(((((((((..((.......))..)))))))))..).)))))))). ( -37.00) >DroSim_CAF1 23043 117 + 1 AA---CCGGGCGUAUGUAAAUAUUGGCAUUGGUGACAUGUGCAAAUGUUUGUAUGGCUGAUUCCCUGAGACCCUAACUUGUGACUUUUAAUACCAGUUUCACAAGUUUUGAUCUCCGGUA .(---((((((.((((((((((((.((((((....)).)))).)))))))))))))))........((((.(..(((((((((..((.......))..)))))))))..).)))))))). ( -37.00) >DroWil_CAF1 28887 117 + 1 UA---CAGUUCAAAUAAUAUUAUUGGUAUUAGUGACAUGUGCAAAUGUUUGUAUGGCUGAUUCCCUGAGACCCUAACUUGUGACUUUUAAUACCAGUUUCACAAGUUUUGAUCUCCGGUA ..---(((((((..((((((.....)))))).)))...((((((....)))))).))))....((.((((.(..(((((((((..((.......))..)))))))))..).)))).)).. ( -25.90) >DroAna_CAF1 18508 120 + 1 AUCCGCCGGGCGCAUGUAAAUAUUGGUAUUGGUGACAUGUGCAAAUGUUUGUAUGGCUGAUUCCCUGAGACCCUAACUUGUGACUUUUAAUACCAGUUUCACAAGUUUUGAUCUCCGGUA ....(((((((.((((((((((((.((((((....)).)))).)))))))))))))))........((((.(..(((((((((..((.......))..)))))))))..).)))))))). ( -37.60) >consensus AA___CCGGGCAUAUGUAAAUAUUGGCAUUGGUGACAUGUGCAAAUGUUUGUAUGGCUGAUUCCCUGAGACCCUAACUUGUGACUUUUAAUACCAGUUUCACAAGUUUUGAUCUCCGGUA ........(((.((((((((((((.((((.........)))).))))))))))))))).....((.((((.(..(((((((((..((.......))..)))))))))..).)))).)).. (-26.27 = -27.17 + 0.89)

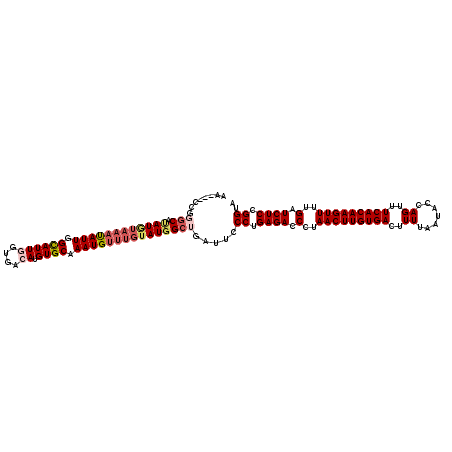
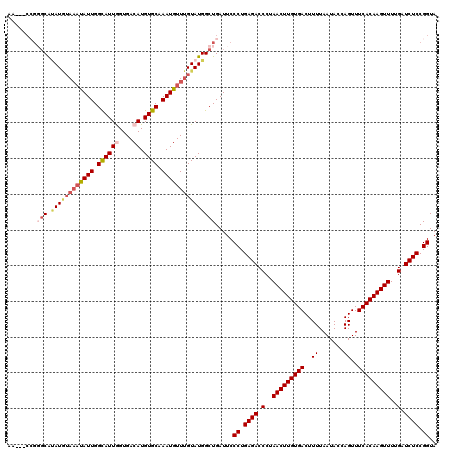
| Location | 18,468,250 – 18,468,365 |
|---|---|
| Length | 115 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 96.41 |
| Mean single sequence MFE | -28.72 |
| Consensus MFE | -28.20 |
| Energy contribution | -28.20 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.75 |
| Structure conservation index | 0.98 |
| SVM decision value | 3.33 |
| SVM RNA-class probability | 0.999016 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 18468250 115 + 22407834 GCAAAUGUUUGUAUGGCUGAUUCCCUGAGACCCUAACUUGUGACUUUUAAUACCAGUUUCACAAGUUUUGAUCUCCGGUAUUGGACGCAAACUUGCUGAUGUUAGUAAA----AAA-UAA ((((..((((((....(..((..((.((((.(..(((((((((..((.......))..)))))))))..).)))).)).))..)..)))))))))).............----...-... ( -28.20) >DroGri_CAF1 27611 115 + 1 GCAAAUGUUUGUAUGGCUGAUUCCCUGAGACCCUAACUUGUGACUUUUAAUACCAGUUUCACAAGUUUUGAUCUCCGGUAUUGGACGCAAACUUGCUGAUGUUAGUAAA----AAA-AAA ((((..((((((....(..((..((.((((.(..(((((((((..((.......))..)))))))))..).)))).)).))..)..)))))))))).............----...-... ( -28.20) >DroSec_CAF1 27264 116 + 1 GCAAAUGUUUGUAUGGCUGAUUCCCUGAGACCCUAACUUGUGACUUUUAAUACCAGUUUCACAAGUUUUGAUCUCCGGUAUUGGACGCAAACUUGCUGAUGUUAGUAAA---AAAA-UAA ((((..((((((....(..((..((.((((.(..(((((((((..((.......))..)))))))))..).)))).)).))..)..)))))))))).............---....-... ( -28.20) >DroWil_CAF1 28924 116 + 1 GCAAAUGUUUGUAUGGCUGAUUCCCUGAGACCCUAACUUGUGACUUUUAAUACCAGUUUCACAAGUUUUGAUCUCCGGUAUUGGACGCAAACUUGCUGAUGUUGGCAAA----AAGCUAA ((....((((((....(..((..((.((((.(..(((((((((..((.......))..)))))))))..).)))).)).))..)..))))))(((((......))))).----..))... ( -31.20) >DroYak_CAF1 22485 115 + 1 GCAAAUGUUUGUAUGGCUGAUUCCCUGAGACCCUAACUUGUGACUUUUAAUACCAGUUUCACAAGUUUUGAUCUCCGGUAUUGGACGCAAACUUGCUGAUGUUAGUAAA----AAA-UAA ((((..((((((....(..((..((.((((.(..(((((((((..((.......))..)))))))))..).)))).)).))..)..)))))))))).............----...-... ( -28.20) >DroAna_CAF1 18548 119 + 1 GCAAAUGUUUGUAUGGCUGAUUCCCUGAGACCCUAACUUGUGACUUUUAAUACCAGUUUCACAAGUUUUGAUCUCCGGUAUUGGACGCAAACUUGCUGAUGUUAAUACAAAAAAGA-UAA ((((..((((((....(..((..((.((((.(..(((((((((..((.......))..)))))))))..).)))).)).))..)..))))))))))...(((....))).......-... ( -28.30) >consensus GCAAAUGUUUGUAUGGCUGAUUCCCUGAGACCCUAACUUGUGACUUUUAAUACCAGUUUCACAAGUUUUGAUCUCCGGUAUUGGACGCAAACUUGCUGAUGUUAGUAAA____AAA_UAA ((((..((((((....(..((..((.((((.(..(((((((((..((.......))..)))))))))..).)))).)).))..)..))))))))))........................ (-28.20 = -28.20 + -0.00)
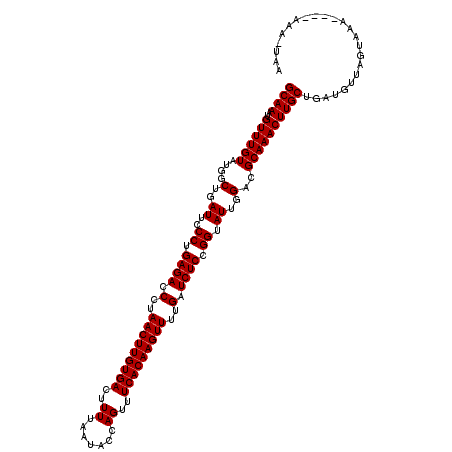



| Location | 18,468,250 – 18,468,365 |
|---|---|
| Length | 115 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 96.41 |
| Mean single sequence MFE | -27.28 |
| Consensus MFE | -26.70 |
| Energy contribution | -26.70 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.78 |
| Structure conservation index | 0.98 |
| SVM decision value | 3.33 |
| SVM RNA-class probability | 0.999021 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 18468250 115 - 22407834 UUA-UUU----UUUACUAACAUCAGCAAGUUUGCGUCCAAUACCGGAGAUCAAAACUUGUGAAACUGGUAUUAAAAGUCACAAGUUAGGGUCUCAGGGAAUCAGCCAUACAAACAUUUGC ...-...----.............(((((((((.((......((.((((((..(((((((((..((.........)))))))))))..)))))).))......))....)))))..)))) ( -26.70) >DroGri_CAF1 27611 115 - 1 UUU-UUU----UUUACUAACAUCAGCAAGUUUGCGUCCAAUACCGGAGAUCAAAACUUGUGAAACUGGUAUUAAAAGUCACAAGUUAGGGUCUCAGGGAAUCAGCCAUACAAACAUUUGC ...-...----.............(((((((((.((......((.((((((..(((((((((..((.........)))))))))))..)))))).))......))....)))))..)))) ( -26.70) >DroSec_CAF1 27264 116 - 1 UUA-UUUU---UUUACUAACAUCAGCAAGUUUGCGUCCAAUACCGGAGAUCAAAACUUGUGAAACUGGUAUUAAAAGUCACAAGUUAGGGUCUCAGGGAAUCAGCCAUACAAACAUUUGC ...-....---.............(((((((((.((......((.((((((..(((((((((..((.........)))))))))))..)))))).))......))....)))))..)))) ( -26.70) >DroWil_CAF1 28924 116 - 1 UUAGCUU----UUUGCCAACAUCAGCAAGUUUGCGUCCAAUACCGGAGAUCAAAACUUGUGAAACUGGUAUUAAAAGUCACAAGUUAGGGUCUCAGGGAAUCAGCCAUACAAACAUUUGC ...(((.----.((.((.......(((....)))...........((((((..(((((((((..((.........)))))))))))..)))))).)).))..)))............... ( -28.60) >DroYak_CAF1 22485 115 - 1 UUA-UUU----UUUACUAACAUCAGCAAGUUUGCGUCCAAUACCGGAGAUCAAAACUUGUGAAACUGGUAUUAAAAGUCACAAGUUAGGGUCUCAGGGAAUCAGCCAUACAAACAUUUGC ...-...----.............(((((((((.((......((.((((((..(((((((((..((.........)))))))))))..)))))).))......))....)))))..)))) ( -26.70) >DroAna_CAF1 18548 119 - 1 UUA-UCUUUUUUGUAUUAACAUCAGCAAGUUUGCGUCCAAUACCGGAGAUCAAAACUUGUGAAACUGGUAUUAAAAGUCACAAGUUAGGGUCUCAGGGAAUCAGCCAUACAAACAUUUGC ...-.....(((((((........(((....)))........((.((((((..(((((((((..((.........)))))))))))..)))))).)).........)))))))....... ( -28.30) >consensus UUA_UUU____UUUACUAACAUCAGCAAGUUUGCGUCCAAUACCGGAGAUCAAAACUUGUGAAACUGGUAUUAAAAGUCACAAGUUAGGGUCUCAGGGAAUCAGCCAUACAAACAUUUGC ........................(((((((((.((......((.((((((..(((((((((..((.........)))))))))))..)))))).))......))....)))))..)))) (-26.70 = -26.70 + -0.00)
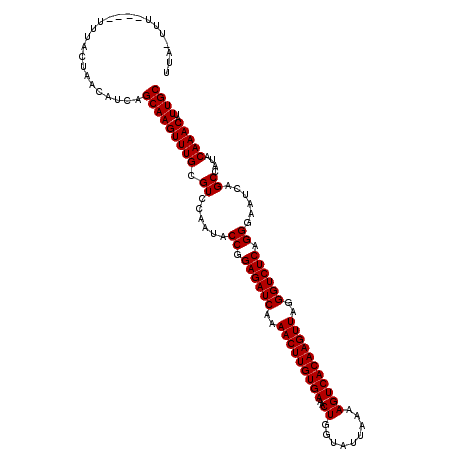


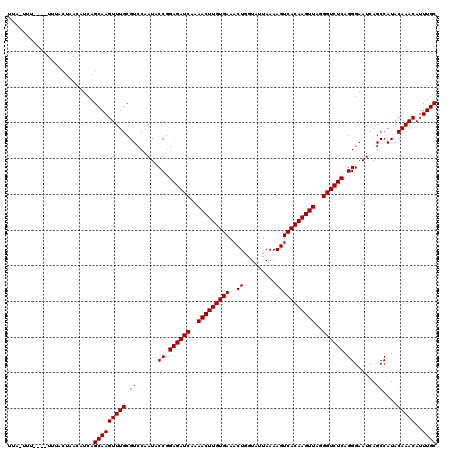
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:14:25 2006