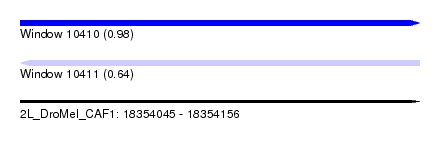

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 18,354,045 – 18,354,156 |
| Length | 111 |
| Max. P | 0.979113 |

| Location | 18,354,045 – 18,354,156 |
|---|---|
| Length | 111 |
| Sequences | 4 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 78.22 |
| Mean single sequence MFE | -33.95 |
| Consensus MFE | -21.08 |
| Energy contribution | -21.02 |
| Covariance contribution | -0.06 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.42 |
| Structure conservation index | 0.62 |
| SVM decision value | 1.83 |
| SVM RNA-class probability | 0.979113 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
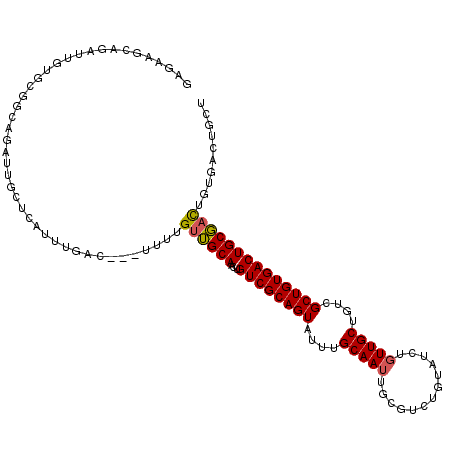
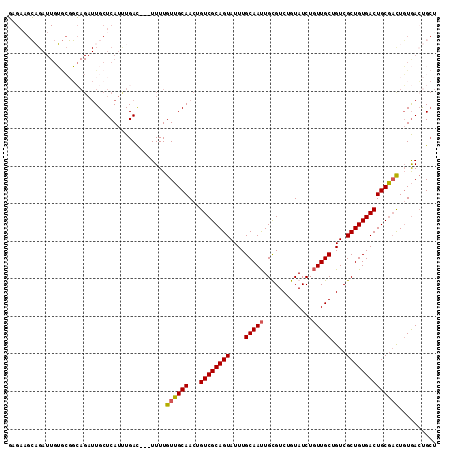
>2L_DroMel_CAF1 18354045 111 + 22407834 GAGAAGCAGAUUGUGCGGCAGAUUGCUCAUUUGAC---UUUUGUUGCAACUGUCGCAGUAUUUGCAAUUGCGUCUGUAUCUGUUGCUGUCGCUGUGACUGCGACUGUGACUGCU .(((.((((((..((((((((.((((.((......---...))..))))))))))))......((....)))))))).)))((..(.(((((.......))))).)..)).... ( -40.90) >DroSec_CAF1 197868 111 + 1 NNNNNNNNNNNNNNNNNNNNNNNUGCUCAUUUGAU---UUUUGUUGCAACUGUCGCAGUAUUUGCAAUUGUGUCUGUAUCUGUUGCUGUCGCUGUGACUGCGACUGUGACUGAU ................................(((---....((((((((((...))))...))))))...)))...(((.((..(.(((((.......))))).)..)).))) ( -23.70) >DroSim_CAF1 205876 111 + 1 GAGAAGCAGAUUGUGCGGCAGAUUGCUCAUUUGAU---UUUUGUUGCAACUGUCGCAGUAUUUGCAAUUGUGUCUGUAUCUGUUGCUGUCGCUGUGACUGCGACUGUGACUGCU .(((.((((((..((((((((.((((.((......---...))..))))))))))))((....))......)))))).)))((..(.(((((.......))))).)..)).... ( -38.80) >DroYak_CAF1 208328 114 + 1 CUGAUGGUGAUUGAAGCUGAGAUUGCUCAUCUGACUGUUUUUGUAGCAACAGUCGCAGUAUUUGCAACUGCGUCUGUAUCUCUUGCUGUCGCUGUGACUGCUCUUUUAAACGCU ...((((((((...(((.(((((((((((............)).))))((((.((((((.......)))))).)))))))))..)))))))))))................... ( -32.40) >consensus GAGAAGCAGAUUGUGCGGCAGAUUGCUCAUUUGAC___UUUUGUUGCAACUGUCGCAGUAUUUGCAAUUGCGUCUGUAUCUGUUGCUGUCGCUGUGACUGCGACUGUGACUGCU ..........................................((((((...((((((((....(((((.............)))))....)))))))))))))).......... (-21.08 = -21.02 + -0.06)

| Location | 18,354,045 – 18,354,156 |
|---|---|
| Length | 111 |
| Sequences | 4 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 78.22 |
| Mean single sequence MFE | -24.81 |
| Consensus MFE | -12.88 |
| Energy contribution | -14.63 |
| Covariance contribution | 1.75 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.18 |
| Structure conservation index | 0.52 |
| SVM decision value | 0.22 |
| SVM RNA-class probability | 0.640385 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
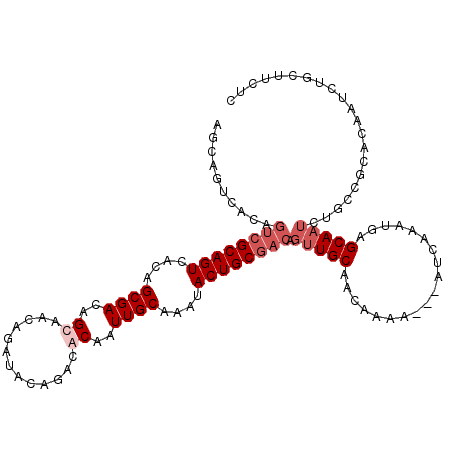
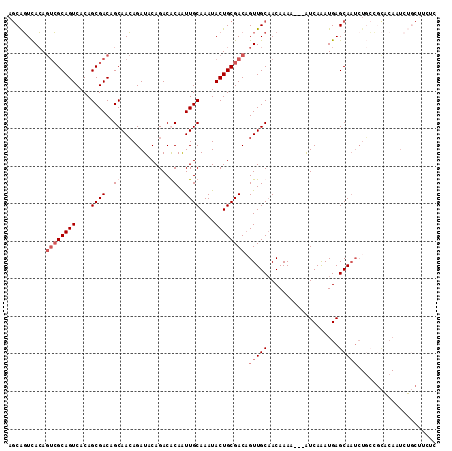
>2L_DroMel_CAF1 18354045 111 - 22407834 AGCAGUCACAGUCGCAGUCACAGCGACAGCAACAGAUACAGACGCAAUUGCAAAUACUGCGACAGUUGCAACAAAA---GUCAAAUGAGCAAUCUGCCGCACAAUCUGCUUCUC (((((.....(((((.......))))).((..(((((......(((((((((.....))))...))))).......---.((....))...)))))..)).....))))).... ( -27.20) >DroSec_CAF1 197868 111 - 1 AUCAGUCACAGUCGCAGUCACAGCGACAGCAACAGAUACAGACACAAUUGCAAAUACUGCGACAGUUGCAACAAAA---AUCAAAUGAGCANNNNNNNNNNNNNNNNNNNNNNN .(((......(((((.......))))).(((((.(.......)....(((((.....)))))..))))).......---......))).......................... ( -17.30) >DroSim_CAF1 205876 111 - 1 AGCAGUCACAGUCGCAGUCACAGCGACAGCAACAGAUACAGACACAAUUGCAAAUACUGCGACAGUUGCAACAAAA---AUCAAAUGAGCAAUCUGCCGCACAAUCUGCUUCUC (((((.....(((((.......))))).((((...............))))......((((.(((((((.......---.........)))).))).))))....))))).... ( -25.45) >DroYak_CAF1 208328 114 - 1 AGCGUUUAAAAGAGCAGUCACAGCGACAGCAAGAGAUACAGACGCAGUUGCAAAUACUGCGACUGUUGCUACAAAAACAGUCAGAUGAGCAAUCUCAGCUUCAAUCACCAUCAG (((..........((.(((.....))).))..(((((......(((((.......)))))(((((((........))))))).........))))).))).............. ( -29.30) >consensus AGCAGUCACAGUCGCAGUCACAGCGACAGCAACAGAUACAGACACAAUUGCAAAUACUGCGACAGUUGCAACAAAA___AUCAAAUGAGCAAUCUGCCGCACAAUCUGCUUCUC ..........((((((((....((((..((.............))..))))....)))))))).(((((...................)))))..................... (-12.88 = -14.63 + 1.75)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:13:17 2006