| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 1,950,894 – 1,951,002 |
| Length | 108 |
| Max. P | 0.995248 |

| Location | 1,950,894 – 1,951,002 |
|---|---|
| Length | 108 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 74.14 |
| Mean single sequence MFE | -51.00 |
| Consensus MFE | -28.28 |
| Energy contribution | -31.27 |
| Covariance contribution | 2.98 |
| Combinations/Pair | 1.16 |
| Mean z-score | -2.68 |
| Structure conservation index | 0.55 |
| SVM decision value | 2.56 |
| SVM RNA-class probability | 0.995248 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 1950894 108 + 22407834 CUCGCAGCGGCCGCCGCUGCCACAAACUGCAGUCAGUCGAUCAGCAGAUCGCUCCUGGAGGUGGCCAUCGAGGAUAUGGCCGUGGUGGUGG------------UGGUCACCGUGGAGCUG ....(((((((((((((..((((...((.(((..((.(((((....))))))).))).))..((((((.......))))))))))..))))------------))))).(....).)))) ( -55.10) >DroEre_CAF1 2543 108 + 1 CUAGCAGCGGCCGCCGCCGCCACAAACUGCAGUCAGUCGAUCAGCAGGUCGCUCCUGGAGCUGGCCAUCGAGGAUAUGGCCGUGGUGGUGG------------UGGUCACCGUGGAGCUG ....(((((((((((((((((((...((.(((..((.(((((....))))))).))).))..((((((.......))))))))))))))))------------))))).(....).)))) ( -56.40) >DroAna_CAF1 22682 105 + 1 CUUGCAGCGGCUGCCGCUGCCACAAAUUGCAGUCAGUCGAUCAACAGAUCGCUCCUGGAAUUG-CAAUUGCUG-----------GUGCUGGUACUGGUACUGGCGG---CCAUGGAACUC .((.((..(((((((((.((((..((((((((((((.(((((....)))))...)))..))))-)))))..))-----------)))).((((....)))))))))---)).)).))... ( -41.50) >consensus CUAGCAGCGGCCGCCGCUGCCACAAACUGCAGUCAGUCGAUCAGCAGAUCGCUCCUGGAGCUGGCCAUCGAGGAUAUGGCCGUGGUGGUGG____________UGGUCACCGUGGAGCUG .((((.(.(((((((((((((((...((.(((..((.(((((....))))))).))).))..((((((.......))))))))))))))))............))))).).))))..... (-28.28 = -31.27 + 2.98)

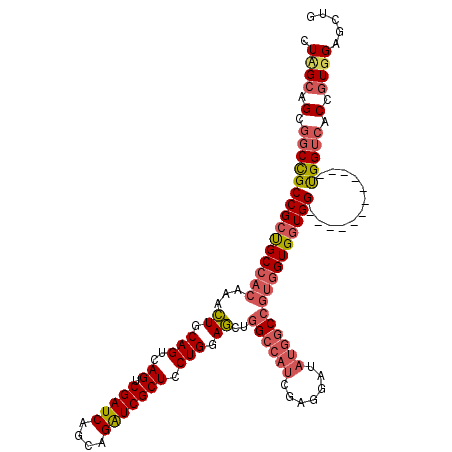
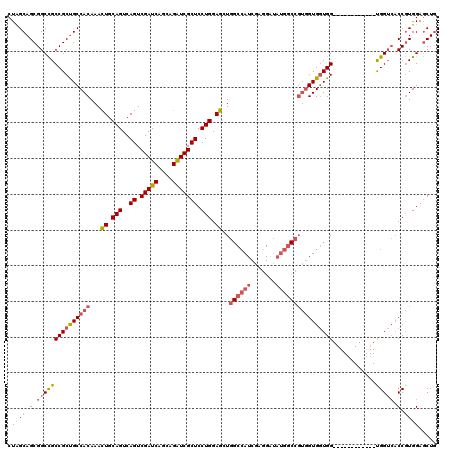
| Location | 1,950,894 – 1,951,002 |
|---|---|
| Length | 108 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 74.14 |
| Mean single sequence MFE | -41.87 |
| Consensus MFE | -29.29 |
| Energy contribution | -32.26 |
| Covariance contribution | 2.98 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.35 |
| Structure conservation index | 0.70 |
| SVM decision value | 0.49 |
| SVM RNA-class probability | 0.756509 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 1950894 108 - 22407834 CAGCUCCACGGUGACCA------------CCACCACCACGGCCAUAUCCUCGAUGGCCACCUCCAGGAGCGAUCUGCUGAUCGACUGACUGCAGUUUGUGGCAGCGGCGGCCGCUGCGAG ..(((((..((((....------------.)))).....((((((.......)))))).......)))))((((....))))(((((....)))))....(((((((...)))))))... ( -46.10) >DroEre_CAF1 2543 108 - 1 CAGCUCCACGGUGACCA------------CCACCACCACGGCCAUAUCCUCGAUGGCCAGCUCCAGGAGCGACCUGCUGAUCGACUGACUGCAGUUUGUGGCGGCGGCGGCCGCUGCUAG ..(((((..((((....------------.)))).....((((((.......)))))).......)))))...((((.(.((....))).))))...(..(((((....)))))..)... ( -44.30) >DroAna_CAF1 22682 105 - 1 GAGUUCCAUGG---CCGCCAGUACCAGUACCAGCAC-----------CAGCAAUUG-CAAUUCCAGGAGCGAUCUGUUGAUCGACUGACUGCAAUUUGUGGCAGCGGCAGCCGCUGCAAG ........(((---..((..(((....)))..)).)-----------))(((((((-((....(((...(((((....))))).)))..))))).)))).(((((((...)))))))... ( -35.20) >consensus CAGCUCCACGGUGACCA____________CCACCACCACGGCCAUAUCCUCGAUGGCCAACUCCAGGAGCGAUCUGCUGAUCGACUGACUGCAGUUUGUGGCAGCGGCGGCCGCUGCAAG ..(((((..((((.................)))).....((((((.......)))))).......)))))((((....))))(((((....)))))....(((((((...)))))))... (-29.29 = -32.26 + 2.98)

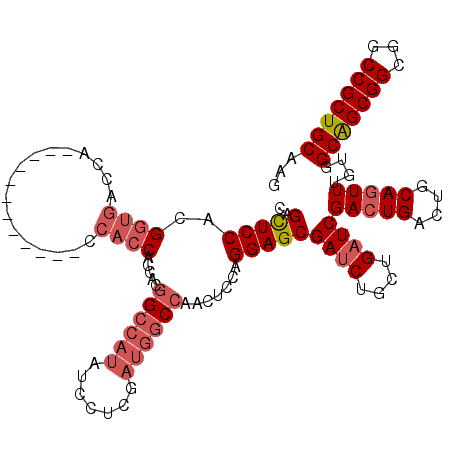
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:41:53 2006