| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 18,165,787 – 18,165,881 |
| Length | 94 |
| Max. P | 0.678647 |

| Location | 18,165,787 – 18,165,881 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 94 |
| Reading direction | forward |
| Mean pairwise identity | 84.58 |
| Mean single sequence MFE | -19.91 |
| Consensus MFE | -14.68 |
| Energy contribution | -14.72 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.16 |
| Mean z-score | -1.75 |
| Structure conservation index | 0.74 |
| SVM decision value | 0.30 |
| SVM RNA-class probability | 0.678647 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 18165787 94 + 22407834 UAAUUACAAGUUCAGCUGAGUUUUCAAAUGAGUUUUCCCUUUCAGCUGAAGCUUUAAUUUGCGUACUUGAAAUUAAUGCCGAUUGAAAGUUACU (((((.((((((((((((((....................))))))))).((........))..))))).)))))................... ( -19.15) >DroSec_CAF1 8334 94 + 1 UAAUUACAAGUUCAGCUGAGUUUUCAAAUGAGUUUUCCCUUUCAGCUGAAGCUUUAAUUUGCGUACUUGAAAUUAAUGCCGAUUGAAAGUUACU (((((.((((((((((((((....................))))))))).((........))..))))).)))))................... ( -19.15) >DroSim_CAF1 8352 94 + 1 UAAUUACAAGUUCAGCUGAGUUUUCAAAUGAGUUUUCCCUUUCAGCUGAACCUUCAAUUUGCGUAUCUGAAAUUAAUGCCGAUUGAAAGUUACU (((((....(((((((((((....................))))))))))).(((((((.((((...........)))).)))))))))))).. ( -23.35) >DroEre_CAF1 8529 88 + 1 UAAUUACAAGUUCAGCUGAGUUUUUAAAUGAGUUUUCCCUUUCAGCUGGAGCUUUAAGUUUCGUUCUUGGUUGUACUG------GAAAUAUAAC ......((((..((((((((.....(((....))).....)))))))(((((.....))))))..))))((((((.(.------...))))))) ( -17.40) >DroYak_CAF1 8647 88 + 1 UAAUUACAAGUCCAGCUGAGUUUUUAAAUGAGUUUUCCCUUUCAGCUGGAGCUUUAAUUUGCAUACUUGGUUGUACUC------AAAAUAUAAU ......((((((((((((((.....(((....))).....))))))))))((........))...))))((((((...------....)))))) ( -20.50) >consensus UAAUUACAAGUUCAGCUGAGUUUUCAAAUGAGUUUUCCCUUUCAGCUGAAGCUUUAAUUUGCGUACUUGAAAUUAAUGCCGAUUGAAAGUUACU ......((((((((((((((.....(((....))).....))))))))))((........))...))))......................... (-14.68 = -14.72 + 0.04)

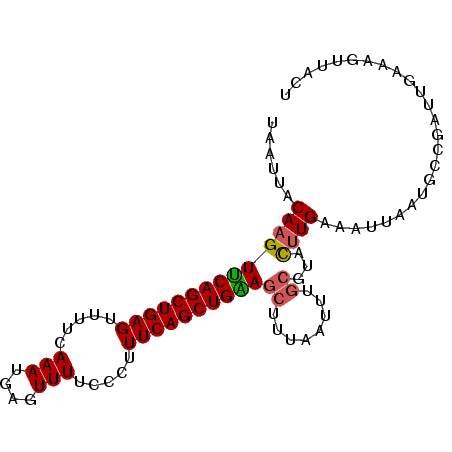
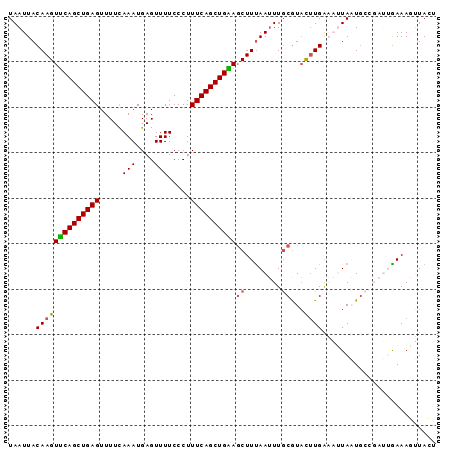
| Location | 18,165,787 – 18,165,881 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 94 |
| Reading direction | reverse |
| Mean pairwise identity | 84.58 |
| Mean single sequence MFE | -17.53 |
| Consensus MFE | -13.48 |
| Energy contribution | -13.48 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.66 |
| Structure conservation index | 0.77 |
| SVM decision value | 0.24 |
| SVM RNA-class probability | 0.648122 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 18165787 94 - 22407834 AGUAACUUUCAAUCGGCAUUAAUUUCAAGUACGCAAAUUAAAGCUUCAGCUGAAAGGGAAAACUCAUUUGAAAACUCAGCUGAACUUGUAAUUA ...................(((((.(((((..((........)).((((((((..(((....)))...(....).))))))))))))).))))) ( -18.20) >DroSec_CAF1 8334 94 - 1 AGUAACUUUCAAUCGGCAUUAAUUUCAAGUACGCAAAUUAAAGCUUCAGCUGAAAGGGAAAACUCAUUUGAAAACUCAGCUGAACUUGUAAUUA ...................(((((.(((((..((........)).((((((((..(((....)))...(....).))))))))))))).))))) ( -18.20) >DroSim_CAF1 8352 94 - 1 AGUAACUUUCAAUCGGCAUUAAUUUCAGAUACGCAAAUUGAAGGUUCAGCUGAAAGGGAAAACUCAUUUGAAAACUCAGCUGAACUUGUAAUUA ........(((((..((...............))..)))))((((((((((((..(((....)))...(....).))))))))))))....... ( -21.56) >DroEre_CAF1 8529 88 - 1 GUUAUAUUUC------CAGUACAACCAAGAACGAAACUUAAAGCUCCAGCUGAAAGGGAAAACUCAUUUAAAAACUCAGCUGAACUUGUAAUUA ..........------...(((((..(((.......))).......(((((((....(((......)))......)))))))...))))).... ( -10.10) >DroYak_CAF1 8647 88 - 1 AUUAUAUUUU------GAGUACAACCAAGUAUGCAAAUUAAAGCUCCAGCUGAAAGGGAAAACUCAUUUAAAAACUCAGCUGGACUUGUAAUUA ....((.(((------(.((((......)))).)))).))(((.(((((((((....(((......)))......))))))))))))....... ( -19.60) >consensus AGUAACUUUCAAUCGGCAUUAAUUUCAAGUACGCAAAUUAAAGCUUCAGCUGAAAGGGAAAACUCAUUUGAAAACUCAGCUGAACUUGUAAUUA ...................................((((((((.(((((((((..(((....)))...(....).)))))))))))).))))). (-13.48 = -13.48 + 0.00)

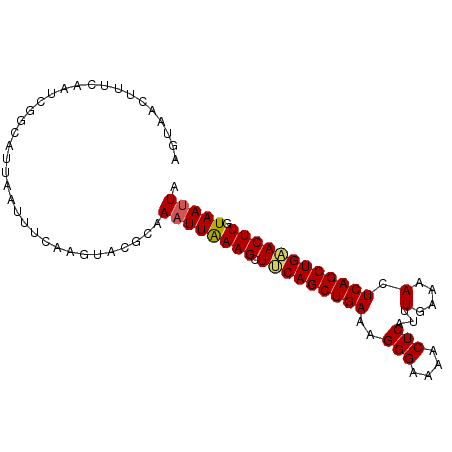
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:11:51 2006