
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 18,046,110 – 18,046,202 |
| Length | 92 |
| Max. P | 0.918187 |

| Location | 18,046,110 – 18,046,202 |
|---|---|
| Length | 92 |
| Sequences | 6 |
| Columns | 104 |
| Reading direction | forward |
| Mean pairwise identity | 76.57 |
| Mean single sequence MFE | -25.97 |
| Consensus MFE | -15.64 |
| Energy contribution | -15.50 |
| Covariance contribution | -0.14 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.92 |
| Structure conservation index | 0.60 |
| SVM decision value | 1.12 |
| SVM RNA-class probability | 0.918187 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
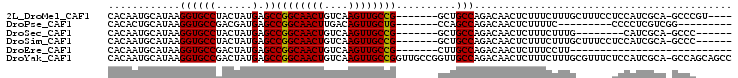
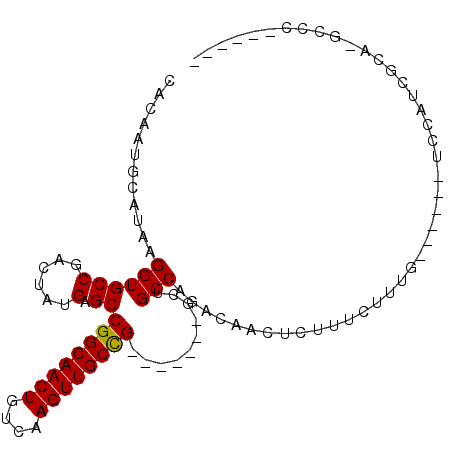
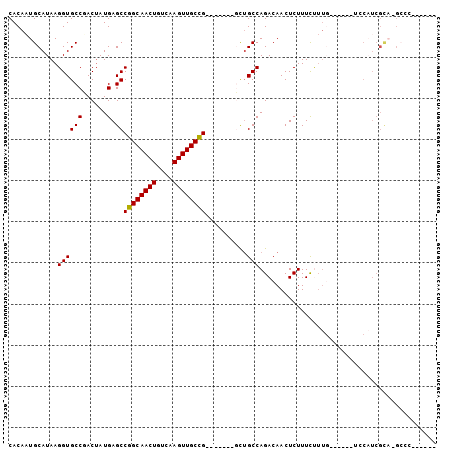
>2L_DroMel_CAF1 18046110 92 + 22407834 CACAAUGCAUAAGGUGCCUACUAUGAGCCGGCAACUGUCAAGUUGCCG-------GCUGCCAGACAACUCUUUCUUUGCUUUCCUCCAUCGCA-GCCCGU---- ............(((((........(((((((((((....))))))))-------)))((.(((........)))..))...........)))-.))...---- ( -27.00) >DroPse_CAF1 353958 79 + 1 CACACUGCAUAAGGUGCCGACGAUGAGCCGGCAACUUGACAGUUGCUG-------CCAGCCAGACAACUCUUUUC---------CCCCUCGUCGG--------- ..((((......))))(((((((((.((.(((((((....))))))))-------)))...(((....)))....---------....)))))))--------- ( -23.10) >DroSec_CAF1 262311 82 + 1 CACAAUGCAUAAGGUGCCUACUAUGAGCCGGCAACUGUCAAGUUGCCG-------GCUGCCAGACAACUCUUUCUUUG--------CAUCGCA-GCCC------ ......((....(((((........(((((((((((....))))))))-------)))...(((....)))......)--------))))...-))..------ ( -27.80) >DroSim_CAF1 262602 90 + 1 CACAAUGCAUAAGGUGCCUACUAUGAGCCGGCAACUGUCAAGUUGCCG-------GCUGCCAGACAACUCUUUCUUUGCUUUCCUCCAUCGCA-GCCC------ ............(((((........(((((((((((....))))))))-------)))((.(((........)))..))...........)))-.)).------ ( -27.00) >DroEre_CAF1 255526 70 + 1 CACAAUGCAUAAGGUGCCGACUAUGAGCCGGCAACUGUCAAGUUGCCG-------CUUGCCAGACAACUCUUUCCUU--------------------------- ......((((...)))).......((((.(((((((....))))))))-------)))...................--------------------------- ( -17.10) >DroYak_CAF1 271751 103 + 1 CACAAUGCAUAAGGUGCCGACUAUGAGCCGGCAACUGUCAAGUUGCCGGUUGCCGGUUGCCAGACAACUCUUUCUUUGCGUUUCUCCAUCGCA-GCCAGCAGCC .....(((....(((..........(((((((((((....))))))))))))))((((((.(((.(((.(.......).)))))).....)))-))).)))... ( -33.80) >consensus CACAAUGCAUAAGGUGCCGACUAUGAGCCGGCAACUGUCAAGUUGCCG_______GCUGCCAGACAACUCUUUCUUUG______UCCAUCGCA_GCCC______ ............((((((......).))((((((((....))))))))..........)))........................................... (-15.64 = -15.50 + -0.14)
| Location | 18,046,110 – 18,046,202 |
|---|---|
| Length | 92 |
| Sequences | 6 |
| Columns | 104 |
| Reading direction | reverse |
| Mean pairwise identity | 76.57 |
| Mean single sequence MFE | -31.42 |
| Consensus MFE | -15.95 |
| Energy contribution | -16.12 |
| Covariance contribution | 0.17 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.05 |
| Structure conservation index | 0.51 |
| SVM decision value | 0.72 |
| SVM RNA-class probability | 0.832669 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
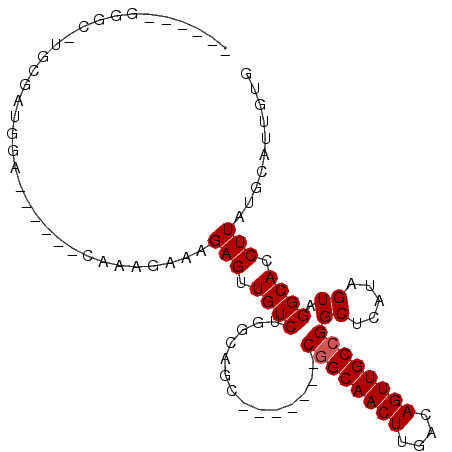
>2L_DroMel_CAF1 18046110 92 - 22407834 ----ACGGGC-UGCGAUGGAGGAAAGCAAAGAAAGAGUUGUCUGGCAGC-------CGGCAACUUGACAGUUGCCGGCUCAUAGUAGGCACCUUAUGCAUUGUG ----......-.((((((((((..(((.........)))(((((..(((-------((((((((....))))))))))).....))))).))))...)))))). ( -30.90) >DroPse_CAF1 353958 79 - 1 ---------CCGACGAGGGG---------GAAAAGAGUUGUCUGGCUGG-------CAGCAACUGUCAAGUUGCCGGCUCAUCGUCGGCACCUUAUGCAGUGUG ---------.(((((((((.---------((......)).)))((((((-------((((.........))))))))))..))))))((((........)))). ( -28.70) >DroSec_CAF1 262311 82 - 1 ------GGGC-UGCGAUG--------CAAAGAAAGAGUUGUCUGGCAGC-------CGGCAACUUGACAGUUGCCGGCUCAUAGUAGGCACCUUAUGCAUUGUG ------....-.((((((--------((....(((.((.(((((..(((-------((((((((....))))))))))).....)))))))))).)))))))). ( -34.30) >DroSim_CAF1 262602 90 - 1 ------GGGC-UGCGAUGGAGGAAAGCAAAGAAAGAGUUGUCUGGCAGC-------CGGCAACUUGACAGUUGCCGGCUCAUAGUAGGCACCUUAUGCAUUGUG ------....-.((((((((((..(((.........)))(((((..(((-------((((((((....))))))))))).....))))).))))...)))))). ( -30.90) >DroEre_CAF1 255526 70 - 1 ---------------------------AAGGAAAGAGUUGUCUGGCAAG-------CGGCAACUUGACAGUUGCCGGCUCAUAGUCGGCACCUUAUGCAUUGUG ---------------------------((((...((((((((.......-------.))))))))......(((((((.....))))))))))).......... ( -23.90) >DroYak_CAF1 271751 103 - 1 GGCUGCUGGC-UGCGAUGGAGAAACGCAAAGAAAGAGUUGUCUGGCAACCGGCAACCGGCAACUUGACAGUUGCCGGCUCAUAGUCGGCACCUUAUGCAUUGUG ((.(((((((-((((.........))).......((((((((.(....).)))))(((((((((....))))))))))))..))))))))))............ ( -39.80) >consensus ______GGGC_UGCGAUGGA______CAAAGAAAGAGUUGUCUGGCAGC_______CGGCAACUUGACAGUUGCCGGCUCAUAGUAGGCACCUUAUGCAUUGUG ..................................(((.((((..............((((((((....))))))))((.....)).)))).))).......... (-15.95 = -16.12 + 0.17)

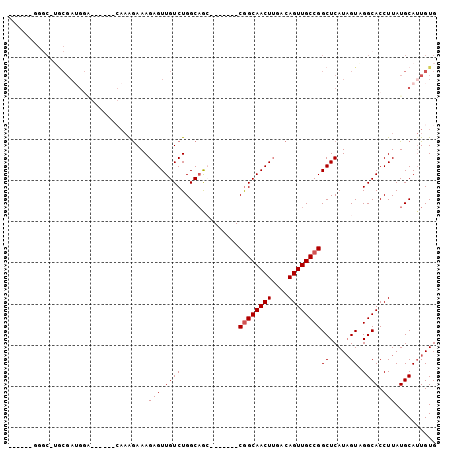
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:10:35 2006