
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 17,881,136 – 17,881,226 |
| Length | 90 |
| Max. P | 0.999991 |

| Location | 17,881,136 – 17,881,226 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 92.28 |
| Mean single sequence MFE | -28.70 |
| Consensus MFE | -25.81 |
| Energy contribution | -26.13 |
| Covariance contribution | 0.32 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.90 |
| SVM decision value | 2.85 |
| SVM RNA-class probability | 0.997414 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 17881136 90 + 22407834 GUGCCAGCAUGGCGUAUACUUGAUGCUCCUUUCCGUGCACC---GUAUGCU-----CGCAUACAUGAAGUAUACGCCCCGAAUGUUGACGUGUUAUUA ....(((((((((((((((((.((((.(......).))...---(((((..-----..))))))).)))))))))))....))))))........... ( -29.70) >DroSec_CAF1 100876 98 + 1 GUGCCAGCAUGGCGUAUACUUGAUGCUCCUUUCCGUGCACCCCUGCACACUCGCAUCGCAUACAUGAAGUAUACGCCCCGAAUGUUGACGUGUUAUUA ....(((((((((((((((((.(((.........(((((....)))))....((...))...))).)))))))))))....))))))........... ( -31.90) >DroSim_CAF1 83575 98 + 1 GUGCCAGCAUGGCGUACACUUGAUGCUCCUUUCCGUGCACCCCUGCACACUCGCAUCGCAUACAUGAAGUAUACGCCCCGAAUGUUGACGUGUUAUUA ....((((((((((((.((((.(((.........(((((....)))))....((...))...))).)))).))))))....))))))........... ( -27.80) >DroEre_CAF1 90740 95 + 1 GUGCCAGCAUGGCGUAUACUUGAUGCUCCUUUCCGUGCAUCCCUGCACACGCG---CGCAUACAUGAAGUAUACGCCCCGAAUGUUGACGUGUUAUUA ....(((((((((((((((((.(((.........(((((....)))))..(..---..)...))).)))))))))))....))))))........... ( -31.10) >DroYak_CAF1 95291 95 + 1 GUGCCAUCAUGGCGUAUACUUGAUGCUCCUUUCCAUGCACCCCUGCACACGCU---CGCAUACAUGAAGUAUACGCCCCGAAUGUUGACGUGUUAUUA ....((.((((((((((((((.(((..........((((....))))...(..---..)...))).)))))))))))....))).))........... ( -23.00) >consensus GUGCCAGCAUGGCGUAUACUUGAUGCUCCUUUCCGUGCACCCCUGCACACUCG___CGCAUACAUGAAGUAUACGCCCCGAAUGUUGACGUGUUAUUA ....(((((((((((((((((.((((........((((......)))).........)))).....)))))))))))....))))))........... (-25.81 = -26.13 + 0.32)
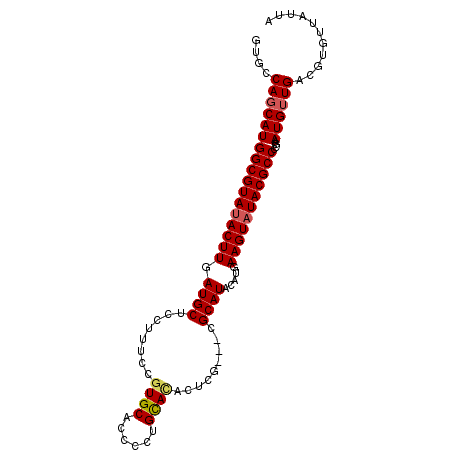


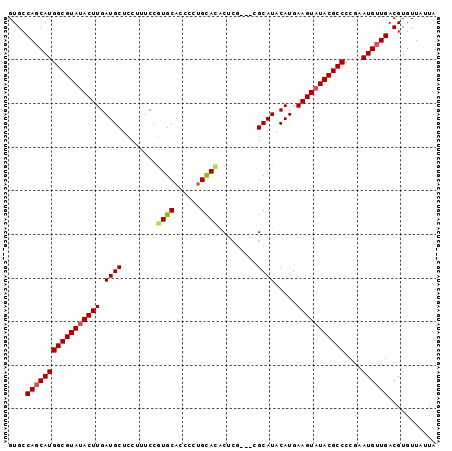
| Location | 17,881,136 – 17,881,226 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 92.28 |
| Mean single sequence MFE | -34.68 |
| Consensus MFE | -33.96 |
| Energy contribution | -33.20 |
| Covariance contribution | -0.76 |
| Combinations/Pair | 1.19 |
| Mean z-score | -2.90 |
| Structure conservation index | 0.98 |
| SVM decision value | 5.61 |
| SVM RNA-class probability | 0.999991 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 17881136 90 - 22407834 UAAUAACACGUCAACAUUCGGGGCGUAUACUUCAUGUAUGCG-----AGCAUAC---GGUGCACGGAAAGGAGCAUCAAGUAUACGCCAUGCUGGCAC .........((((.(((....(((((((((((...(((((..-----..)))))---(((((.(......).))))))))))))))))))).)))).. ( -33.00) >DroSec_CAF1 100876 98 - 1 UAAUAACACGUCAACAUUCGGGGCGUAUACUUCAUGUAUGCGAUGCGAGUGUGCAGGGGUGCACGGAAAGGAGCAUCAAGUAUACGCCAUGCUGGCAC .........((((.(((....(((((((((((..(((((((.......)))))))..(((((.(......).))))))))))))))))))).)))).. ( -33.60) >DroSim_CAF1 83575 98 - 1 UAAUAACACGUCAACAUUCGGGGCGUAUACUUCAUGUAUGCGAUGCGAGUGUGCAGGGGUGCACGGAAAGGAGCAUCAAGUGUACGCCAUGCUGGCAC .........(((((((((((.(.(((((((.....))))))).).)))))))(((((.((((((.((........))..)))))).)).))))))).. ( -37.70) >DroEre_CAF1 90740 95 - 1 UAAUAACACGUCAACAUUCGGGGCGUAUACUUCAUGUAUGCG---CGCGUGUGCAGGGAUGCACGGAAAGGAGCAUCAAGUAUACGCCAUGCUGGCAC .........((((.(((.....((((((((.....)))))))---)((((((((...(((((.(......).)))))..)))))))).))).)))).. ( -36.10) >DroYak_CAF1 95291 95 - 1 UAAUAACACGUCAACAUUCGGGGCGUAUACUUCAUGUAUGCG---AGCGUGUGCAGGGGUGCAUGGAAAGGAGCAUCAAGUAUACGCCAUGAUGGCAC .........((((....(((.(((((((((((..(((((((.---...)))))))..(((((..........)))))))))))))))).))))))).. ( -33.00) >consensus UAAUAACACGUCAACAUUCGGGGCGUAUACUUCAUGUAUGCG___CGAGUGUGCAGGGGUGCACGGAAAGGAGCAUCAAGUAUACGCCAUGCUGGCAC .........((((.(((....(((((((((((..(((((((.......)))))))..(((((.(......).))))))))))))))))))).)))).. (-33.96 = -33.20 + -0.76)


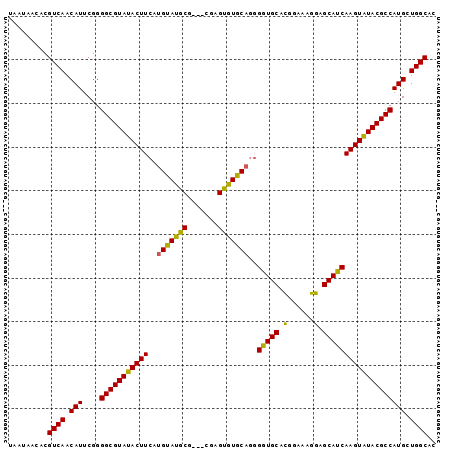
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:09:37 2006