
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 17,626,614 – 17,626,715 |
| Length | 101 |
| Max. P | 0.789836 |

| Location | 17,626,614 – 17,626,715 |
|---|---|
| Length | 101 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 87.23 |
| Mean single sequence MFE | -28.80 |
| Consensus MFE | -20.36 |
| Energy contribution | -21.92 |
| Covariance contribution | 1.56 |
| Combinations/Pair | 1.16 |
| Mean z-score | -1.75 |
| Structure conservation index | 0.71 |
| SVM decision value | 0.08 |
| SVM RNA-class probability | 0.574297 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
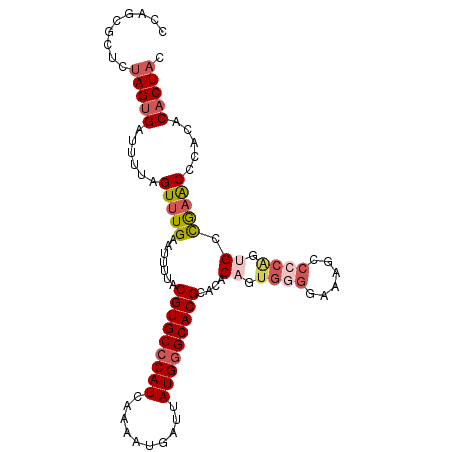
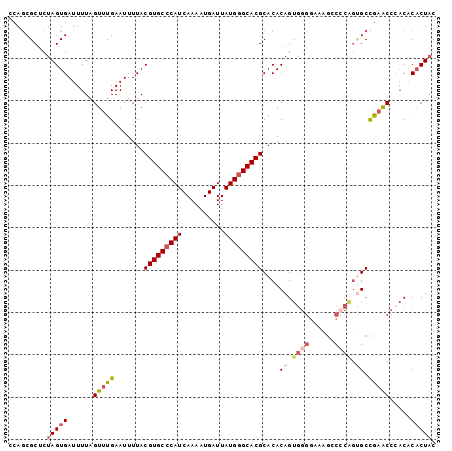
>2L_DroMel_CAF1 17626614 101 + 22407834 CCAGCGCUCUAGUGAUUUUAGUUUGAAUUUUACGUGCCCAUCAAAAUGAUUAUGGGCACGCACACAGUGGGGAAAGCCCCAGUGCCGAACCGACACACUAC .........(((((...(..(((((........((((((((..........))))))))((((....(((((....))))))))))))))..)..))))). ( -33.70) >DroSec_CAF1 21956 101 + 1 CCAGCGCUCUAGUGAUUUUAGUUUGAAUUUUACGUGCCCAUCAAAAUGAUUAUGGGCACGCACACAGUGGGGAAAGGCCCGGUGCCGAACCCACACACUAC .........(((((..................(((((((((..........)))))))))......(((((....(((.....)))...))))).))))). ( -33.30) >DroSim_CAF1 12178 101 + 1 CCAGCGCUCUAGUGAUUUUAGUUUGAAUUUUACGUGCCCAUCAAAAUGAUUAUGGGCACGCACACAGUGGGGAAAGGCCCGGUGCCGAACCCACGCACUAC ...((((....((((..((......))..))))((((((((..........)))))))))).....(((((....(((.....)))...)))))))..... ( -34.10) >DroYak_CAF1 22011 101 + 1 CCAGCGCCCUAGUGAUUUUAGUUUGAAUUUUACGUGCCCAUCAAAGUGAUUAUGGGCACGCACACAGUGGGGACAGCCACAGCGCCGCGCCCACACACUAC .........(((((..................(((((((((..((....)))))))))))......((((((...((....))....).))))).))))). ( -30.30) >DroAna_CAF1 4340 89 + 1 CCAACGCUCUAGUGAUUUUAGUUUGAAUUUUACGUGCCCAUCAAAAUGAUUAUGAGCACGCACACAGC---------CCC---GCUAAACACUCCCUCUCC ..........((((..((((((..........(((((.(((..........))).)))))........---------...---))))))))))........ ( -12.60) >consensus CCAGCGCUCUAGUGAUUUUAGUUUGAAUUUUACGUGCCCAUCAAAAUGAUUAUGGGCACGCACACAGUGGGGAAAGCCCCAGUGCCGAACCCACACACUAC .........(((((......(((((.......(((((((((..........)))))))))....((.((((......)))).)).))))).....))))). (-20.36 = -21.92 + 1.56)

| Location | 17,626,614 – 17,626,715 |
|---|---|
| Length | 101 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 87.23 |
| Mean single sequence MFE | -34.38 |
| Consensus MFE | -24.74 |
| Energy contribution | -25.30 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.27 |
| Mean z-score | -2.08 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.58 |
| SVM RNA-class probability | 0.789836 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
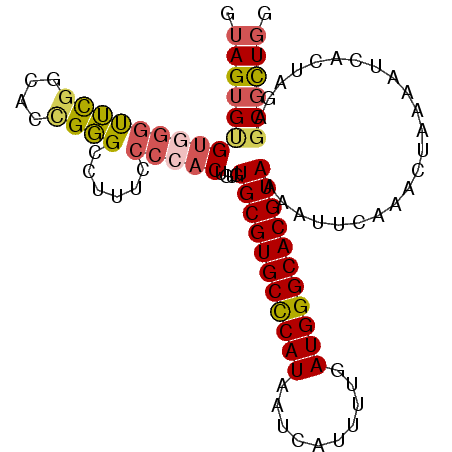
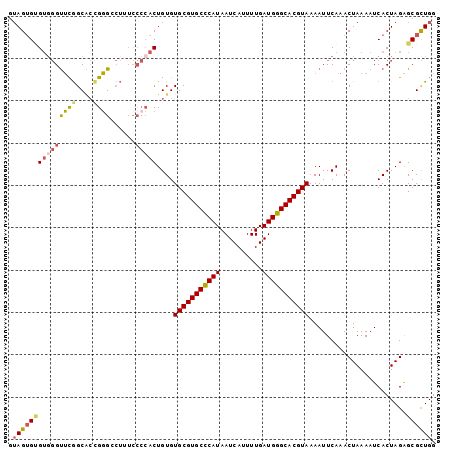
>2L_DroMel_CAF1 17626614 101 - 22407834 GUAGUGUGUCGGUUCGGCACUGGGGCUUUCCCCACUGUGUGCGUGCCCAUAAUCAUUUUGAUGGGCACGUAAAAUUCAAACUAAAAUCACUAGAGCGCUGG .((((((...(((...(((.(((((....))))).))).(((((((((((..........))))))))))).................)))...)))))). ( -35.70) >DroSec_CAF1 21956 101 - 1 GUAGUGUGUGGGUUCGGCACCGGGCCUUUCCCCACUGUGUGCGUGCCCAUAAUCAUUUUGAUGGGCACGUAAAAUUCAAACUAAAAUCACUAGAGCGCUGG .(((((((((((...(((.....)))....)))))....(((((((((((..........))))))))))).......................)))))). ( -35.80) >DroSim_CAF1 12178 101 - 1 GUAGUGCGUGGGUUCGGCACCGGGCCUUUCCCCACUGUGUGCGUGCCCAUAAUCAUUUUGAUGGGCACGUAAAAUUCAAACUAAAAUCACUAGAGCGCUGG .(((((((((((...(((.....)))....)))))....(((((((((((..........))))))))))).......................)))))). ( -38.40) >DroYak_CAF1 22011 101 - 1 GUAGUGUGUGGGCGCGGCGCUGUGGCUGUCCCCACUGUGUGCGUGCCCAUAAUCACUUUGAUGGGCACGUAAAAUUCAAACUAAAAUCACUAGGGCGCUGG .(((((((((((.(((((......))))).)))))....(((((((((((..........))))))))))).......................)))))). ( -40.60) >DroAna_CAF1 4340 89 - 1 GGAGAGGGAGUGUUUAGC---GGG---------GCUGUGUGCGUGCUCAUAAUCAUUUUGAUGGGCACGUAAAAUUCAAACUAAAAUCACUAGAGCGUUGG ...(....(((((((((.---..(---------..(...(((((((((((..........)))))))))))..)..)...)))))..))))....)..... ( -21.40) >consensus GUAGUGUGUGGGUUCGGCACCGGGCCUUUCCCCACUGUGUGCGUGCCCAUAAUCAUUUUGAUGGGCACGUAAAAUUCAAACUAAAAUCACUAGAGCGCUGG .(((((((((((((((....))))......)))))....(((((((((((..........))))))))))).......................)))))). (-24.74 = -25.30 + 0.56)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:08:21 2006