
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 17,618,989 – 17,619,129 |
| Length | 140 |
| Max. P | 0.987335 |

| Location | 17,618,989 – 17,619,089 |
|---|---|
| Length | 100 |
| Sequences | 4 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 86.39 |
| Mean single sequence MFE | -28.83 |
| Consensus MFE | -24.27 |
| Energy contribution | -24.03 |
| Covariance contribution | -0.25 |
| Combinations/Pair | 1.16 |
| Mean z-score | -2.65 |
| Structure conservation index | 0.84 |
| SVM decision value | 2.08 |
| SVM RNA-class probability | 0.987335 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
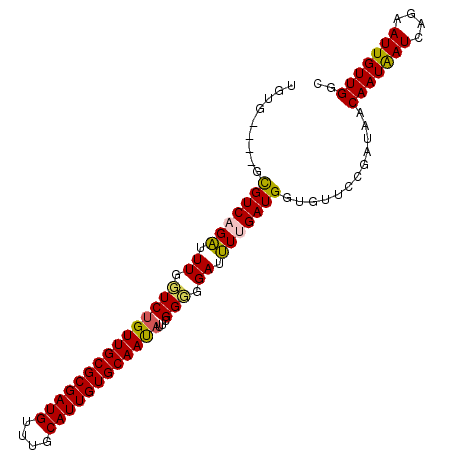
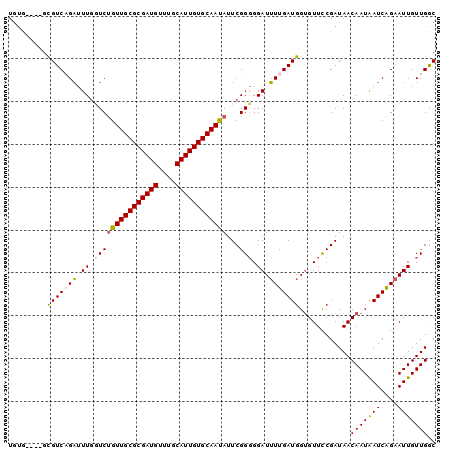
>2L_DroMel_CAF1 17618989 100 - 22407834 UUUUUUGUGCGUCAGAUUUUGUCUGUUGCGCGAUGUUUGCAUUGUGCAAUAUUCGGGGGAUUUUGAUGGUGUUCCGAUAACAAUAAUCAGAAUUGUUGGC ..........(((((.(((((..((((((((((((....))))))))))))..)))))(((((((((..((((.....))))...))))))))).))))) ( -30.30) >DroSec_CAF1 14374 96 - 1 UGUG----GUGUCUGAUUUGUUCUGUUGCGCGAUGUUUGCAUUGUGCAAUAUUCGGGGGAUUUUGAUGGUGUUUCGAUAACAAUGAUCGGAAUUGUUGGC .(..----((.(((((((((((.((((((((((((....)))))))))))).(((..(.(((.....))).)..))).))))..)))))))))..).... ( -29.60) >DroSim_CAF1 4573 96 - 1 UGUG----GUGUCUGAUUUGUUCUGUUGCGCGAUGUUUGCAUUGUGCAAUAUUCGGGGGAUUUUGAUGGUGUUUCGAUAACAAUAAUCAGAAUUGUUGGC .(..----((.(((((((((((.((((((((((((....)))))))))))).(((..(.(((.....))).)..))).))))..)))))))))..).... ( -29.90) >DroEre_CAF1 2819 92 - 1 UGUU----GCGUCAGUUUUGGUCGGUUGCGCGAUG----CAUUGUGCAACAUUCGGUGGAUAUUGAUGGCGUUCCGAUAACAAUAAUCAGGAUUGUUGAC .(..----((((((..........(((((((((..----..)))))))))..(((((....)))))))))))..).....(((((((....))))))).. ( -25.50) >consensus UGUG____GCGUCAGAUUUGGUCUGUUGCGCGAUGUUUGCAUUGUGCAAUAUUCGGGGGAUUUUGAUGGUGUUCCGAUAACAAUAAUCAGAAUUGUUGGC .........(((((((.((.(((((((((((((((....))))))))))))...))).)).)))))))............(((((((....))))))).. (-24.27 = -24.03 + -0.25)

| Location | 17,619,018 – 17,619,129 |
|---|---|
| Length | 111 |
| Sequences | 4 |
| Columns | 111 |
| Reading direction | reverse |
| Mean pairwise identity | 88.38 |
| Mean single sequence MFE | -27.52 |
| Consensus MFE | -23.15 |
| Energy contribution | -23.40 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.15 |
| Mean z-score | -1.81 |
| Structure conservation index | 0.84 |
| SVM decision value | 0.76 |
| SVM RNA-class probability | 0.844166 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
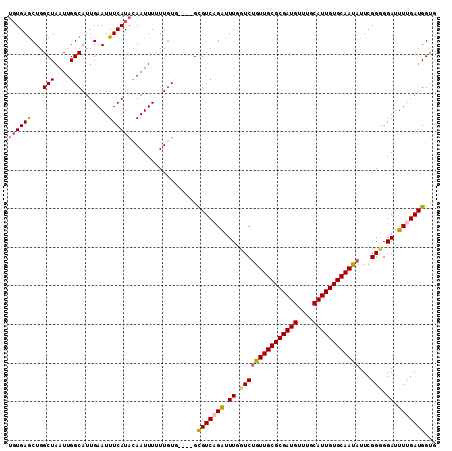
>2L_DroMel_CAF1 17619018 111 - 22407834 UGUGAGCUGGCUAAUUGGCAUUGAAUUUCAUACAAUUUUUUUUUUUGUGCGUCAGAUUUUGUCUGUUGCGCGAUGUUUGCAUUGUGCAAUAUUCGGGGGAUUUUGAUGGUG .....((..(....)..))...........(((((.........)))))(((((((.(((.((((((((((((((....))))))))))))...)).))).)))))))... ( -29.20) >DroSec_CAF1 14403 107 - 1 UGUGAGCUGGCUAAUUGGCAUUGAAUUUCAUACAAUUUUUUGUG----GUGUCUGAUUUGUUCUGUUGCGCGAUGUUUGCAUUGUGCAAUAUUCGGGGGAUUUUGAUGGUG .....((..(....)..))((..((.(((.(((((....)))))----...(((((.......((((((((((((....)))))))))))).)))))))).))..)).... ( -26.80) >DroSim_CAF1 4602 107 - 1 UGUGAGCUGGCUAAUUGGCAUUGAAUUUCAUACAAUUUUUUGUG----GUGUCUGAUUUGUUCUGUUGCGCGAUGUUUGCAUUGUGCAAUAUUCGGGGGAUUUUGAUGGUG .....((..(....)..))((..((.(((.(((((....)))))----...(((((.......((((((((((((....)))))))))))).)))))))).))..)).... ( -26.80) >DroEre_CAF1 2848 103 - 1 UUUGAACUGGCUAAUUGGCAUUGAAUUUCAUUUAAUUUUUUGUU----GCGUCAGUUUUGGUCGGUUGCGCGAUG----CAUUGUGCAACAUUCGGUGGAUAUUGAUGGCG ...((((((((....(((((..(((((......)))))..))))----).))))))))..(((((((((((((..----..)))))))))..(((((....))))))))). ( -27.30) >consensus UGUGAGCUGGCUAAUUGGCAUUGAAUUUCAUACAAUUUUUUGUG____GCGUCAGAUUUGGUCUGUUGCGCGAUGUUUGCAUUGUGCAAUAUUCGGGGGAUUUUGAUGGUG ((((((...(((....))).......)))))).................(((((((.((.(((((((((((((((....))))))))))))...))).)).)))))))... (-23.15 = -23.40 + 0.25)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:08:19 2006