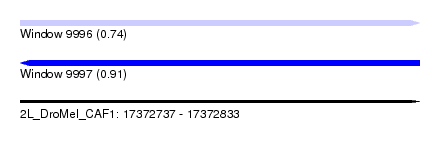

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 17,372,737 – 17,372,833 |
| Length | 96 |
| Max. P | 0.914418 |

| Location | 17,372,737 – 17,372,833 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 97 |
| Reading direction | forward |
| Mean pairwise identity | 92.80 |
| Mean single sequence MFE | -14.64 |
| Consensus MFE | -11.80 |
| Energy contribution | -12.28 |
| Covariance contribution | 0.48 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.58 |
| Structure conservation index | 0.81 |
| SVM decision value | 0.45 |
| SVM RNA-class probability | 0.741522 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 17372737 96 + 22407834 GCGACACCUGCUCAAUUCUGCUUUCACUGGCAAUUUUCCAAGAGUUUCCACUCUAUUUCCCCGCUCAUUCCCCCCUUUUUUUUCUGG-CCCCAGCUG (((...............((((......))))........(((((....))))).......)))...................(((.-...)))... ( -13.00) >DroSec_CAF1 8709 95 + 1 GCGACACCUGCUCAAUUCUGCUUUCACUGGCAGUUUUCCAAGAGUUUCCACUCUAUUUCCCCGCUCAUUCCCCCCUUU-UUUUCUGG-CCCCAGCUG (((..........((..(((((......)))))..))...(((((....))))).......)))..............-....(((.-...)))... ( -15.50) >DroSim_CAF1 8914 95 + 1 GCGACACCUGCUCAAUUCUGCUUUCACUGGCAGUUUUCCAAGAGUUUCCACUCUAUUUCCCCGCCCAUUCCCCCCUUU-UUUUCUGG-CCCCAGCUG (((..........((..(((((......)))))..))...(((((....))))).......)))..............-....(((.-...)))... ( -15.10) >DroEre_CAF1 8589 93 + 1 GCGACACCUGCUCAAUUCUGCUUUCACUGGCAGUUUUCCAAGAGUUUCCACUCUAUUUCCCCGCUCAUUCCCCCUUUU----UAUGGCCCACAGCUG (((..........((..(((((......)))))..))...(((((....))))).......)))..............----...(((.....))). ( -14.80) >DroYak_CAF1 8947 93 + 1 CCGACACCUGCUCGAUUCUGCUUUCACUGGCAGUUUUCCAAGAGUUUCCACUCUAUUUCCCCGCUCAUUCCCCCUUUU----GUUGACCCCCAGCUG .(((((.......((..(((((......)))))...))..(((((....))))).......................)----))))........... ( -14.80) >consensus GCGACACCUGCUCAAUUCUGCUUUCACUGGCAGUUUUCCAAGAGUUUCCACUCUAUUUCCCCGCUCAUUCCCCCCUUU_UUUUCUGG_CCCCAGCUG (((..............(((((......))))).......(((((....))))).......)))...................(((.....)))... (-11.80 = -12.28 + 0.48)

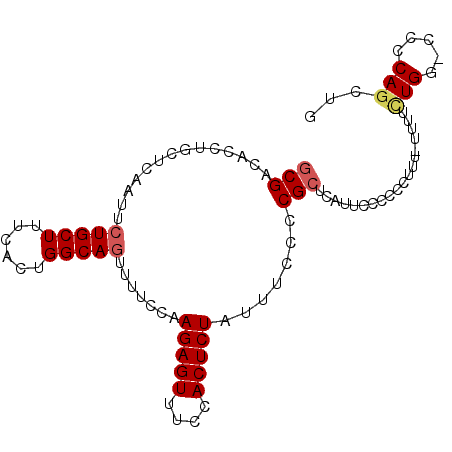
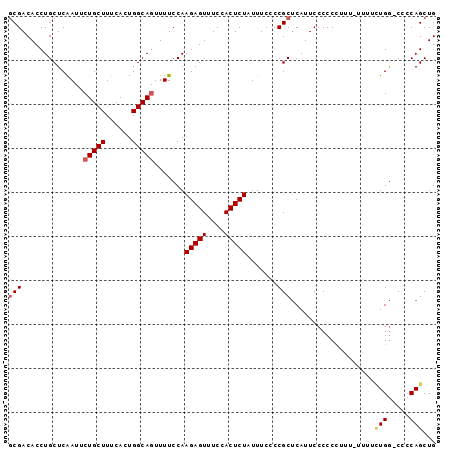
| Location | 17,372,737 – 17,372,833 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 97 |
| Reading direction | reverse |
| Mean pairwise identity | 92.80 |
| Mean single sequence MFE | -21.92 |
| Consensus MFE | -18.98 |
| Energy contribution | -18.82 |
| Covariance contribution | -0.16 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.56 |
| Structure conservation index | 0.87 |
| SVM decision value | 1.10 |
| SVM RNA-class probability | 0.914418 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 17372737 96 - 22407834 CAGCUGGGG-CCAGAAAAAAAAGGGGGGAAUGAGCGGGGAAAUAGAGUGGAAACUCUUGGAAAAUUGCCAGUGAAAGCAGAAUUGAGCAGGUGUCGC ..(((.(..-((..............))..).)))........(((((....)))))..((...((((((((.........)))).))))...)).. ( -19.64) >DroSec_CAF1 8709 95 - 1 CAGCUGGGG-CCAGAAAA-AAAGGGGGGAAUGAGCGGGGAAAUAGAGUGGAAACUCUUGGAAAACUGCCAGUGAAAGCAGAAUUGAGCAGGUGUCGC ..(((.(..-((......-.......))..).)))........(((((....)))))..((...((((((((.........)))).))))...)).. ( -21.92) >DroSim_CAF1 8914 95 - 1 CAGCUGGGG-CCAGAAAA-AAAGGGGGGAAUGGGCGGGGAAAUAGAGUGGAAACUCUUGGAAAACUGCCAGUGAAAGCAGAAUUGAGCAGGUGUCGC ..((.((.(-((......-.............(((((......(((((....))))).......))))).((....))...........))).)))) ( -24.62) >DroEre_CAF1 8589 93 - 1 CAGCUGUGGGCCAUA----AAAAGGGGGAAUGAGCGGGGAAAUAGAGUGGAAACUCUUGGAAAACUGCCAGUGAAAGCAGAAUUGAGCAGGUGUCGC ((.((((...((...----.....))....((.((((......(((((....))))).......))))))((....))........)))).)).... ( -21.12) >DroYak_CAF1 8947 93 - 1 CAGCUGGGGGUCAAC----AAAAGGGGGAAUGAGCGGGGAAAUAGAGUGGAAACUCUUGGAAAACUGCCAGUGAAAGCAGAAUCGAGCAGGUGUCGG ..(((.((..((...----...........((.((((......(((((....))))).......))))))((....)).)).)).)))......... ( -22.32) >consensus CAGCUGGGG_CCAGAAAA_AAAGGGGGGAAUGAGCGGGGAAAUAGAGUGGAAACUCUUGGAAAACUGCCAGUGAAAGCAGAAUUGAGCAGGUGUCGC ((.(((........................((.((((......(((((....))))).......))))))((....)).........))).)).... (-18.98 = -18.82 + -0.16)

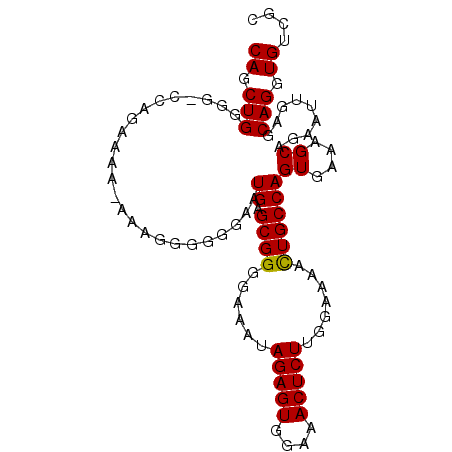
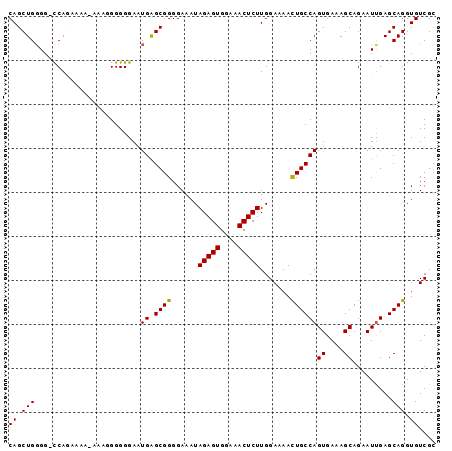
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:06:32 2006