
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 17,357,040 – 17,357,132 |
| Length | 92 |
| Max. P | 0.872216 |

| Location | 17,357,040 – 17,357,132 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 94 |
| Reading direction | forward |
| Mean pairwise identity | 93.13 |
| Mean single sequence MFE | -19.54 |
| Consensus MFE | -15.54 |
| Energy contribution | -16.02 |
| Covariance contribution | 0.48 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.88 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.88 |
| SVM RNA-class probability | 0.872216 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 17357040 92 + 22407834 UUAUUAAUGCAGCGUUCCAUUAAAGAAAUUAAUGGCGA-ACAAA-AAAUAAGGGUGCUGCAAGCUGGACCUGGCGAGAAAAUCGAAAAGAGGGC .......(((((((..(((((((.....))))))(...-.)...-......)..))))))).......(((..(((.....))).....))).. ( -18.60) >DroSec_CAF1 58890 93 + 1 UUAUUAAUGCAGCAUUCCAUUAAAGAAAUUAAUGGCGAAACAAA-AAAUAAGGGUGCUGCAAGCAGGACCUGGCAAGAAAAUCGAAAAGAGGGC .......(((......(((((((.....))))))).........-......((((.(((....))).)))).)))................... ( -18.10) >DroSim_CAF1 59493 93 + 1 UUAUUAAUGCAGCAUUCCAUUAAAGAAAUUAAUGGCGAAACAAA-AAAUAAGGGUGCUGCAAGCAGGACCUGGCGAGAAAAUCGAAAAGAGGGC .......((((((((((((((((.....))))))).........-.......))))))))).......(((..(((.....))).....))).. ( -20.29) >DroEre_CAF1 56667 93 + 1 UUAUUAAUGCAGCAUUCCAUUAAAGAACUUAAUGGCGA-ACAAAAAAAUAAGGGUCCUGCAAGCAGGACCUGGCGAGAAAAUCGGAAAGAGGGC ........((.((...(((((((.....)))))))...-............((((((((....)))))))).))((.....)).........)) ( -22.60) >DroYak_CAF1 53068 92 + 1 UUAUUAAUGCAGCAUUCCAUUAAAGAAAAUAAUGGCGA-ACAAA-AAGUAAGGGUGCUGCAAGCAGUAUCUAAAGAGAAAAUCGGAAAGAGGGC .......(((((((((((((((.......)))))((..-.....-..))..)))))))))).((....(((...((.....))....)))..)) ( -18.10) >consensus UUAUUAAUGCAGCAUUCCAUUAAAGAAAUUAAUGGCGA_ACAAA_AAAUAAGGGUGCUGCAAGCAGGACCUGGCGAGAAAAUCGAAAAGAGGGC .......((((((((((((((((.....))))))).................))))))))).......(((..(((.....))).....))).. (-15.54 = -16.02 + 0.48)


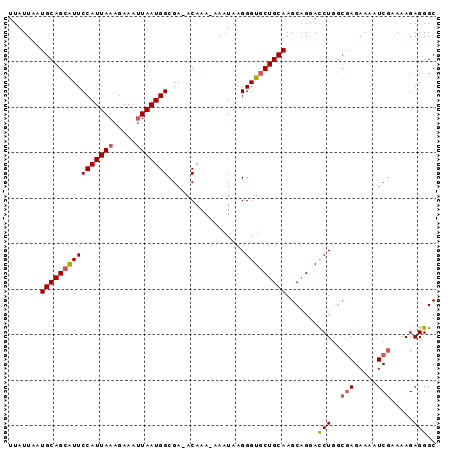
| Location | 17,357,040 – 17,357,132 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 94 |
| Reading direction | reverse |
| Mean pairwise identity | 93.13 |
| Mean single sequence MFE | -17.31 |
| Consensus MFE | -12.74 |
| Energy contribution | -13.58 |
| Covariance contribution | 0.84 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.89 |
| Structure conservation index | 0.74 |
| SVM decision value | -0.02 |
| SVM RNA-class probability | 0.521320 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 17357040 92 - 22407834 GCCCUCUUUUCGAUUUUCUCGCCAGGUCCAGCUUGCAGCACCCUUAUUU-UUUGU-UCGCCAUUAAUUUCUUUAAUGGAACGCUGCAUUAAUAA ((...(((..(((.....)))..)))....)).((((((.....((...-..)).-...(((((((.....)))))))...))))))....... ( -16.90) >DroSec_CAF1 58890 93 - 1 GCCCUCUUUUCGAUUUUCUUGCCAGGUCCUGCUUGCAGCACCCUUAUUU-UUUGUUUCGCCAUUAAUUUCUUUAAUGGAAUGCUGCAUUAAUAA ...................(((..(((.(((....))).))).......-...((....(((((((.....)))))))...)).)))....... ( -15.60) >DroSim_CAF1 59493 93 - 1 GCCCUCUUUUCGAUUUUCUCGCCAGGUCCUGCUUGCAGCACCCUUAUUU-UUUGUUUCGCCAUUAAUUUCUUUAAUGGAAUGCUGCAUUAAUAA ((...(((..(((.....)))..)))....)).(((((((.........-.........(((((((.....)))))))..)))))))....... ( -17.65) >DroEre_CAF1 56667 93 - 1 GCCCUCUUUCCGAUUUUCUCGCCAGGUCCUGCUUGCAGGACCCUUAUUUUUUUGU-UCGCCAUUAAGUUCUUUAAUGGAAUGCUGCAUUAAUAA ....................((..(((((((....)))))))...........((-...((((((((...))))))))...)).))........ ( -21.60) >DroYak_CAF1 53068 92 - 1 GCCCUCUUUCCGAUUUUCUCUUUAGAUACUGCUUGCAGCACCCUUACUU-UUUGU-UCGCCAUUAUUUUCUUUAAUGGAAUGCUGCAUUAAUAA .................................(((((((.....((..-...))-...((((((.......))))))..)))))))....... ( -14.80) >consensus GCCCUCUUUUCGAUUUUCUCGCCAGGUCCUGCUUGCAGCACCCUUAUUU_UUUGU_UCGCCAUUAAUUUCUUUAAUGGAAUGCUGCAUUAAUAA ((...(((..(((.....)))..)))....)).(((((((...................(((((((.....)))))))..)))))))....... (-12.74 = -13.58 + 0.84)


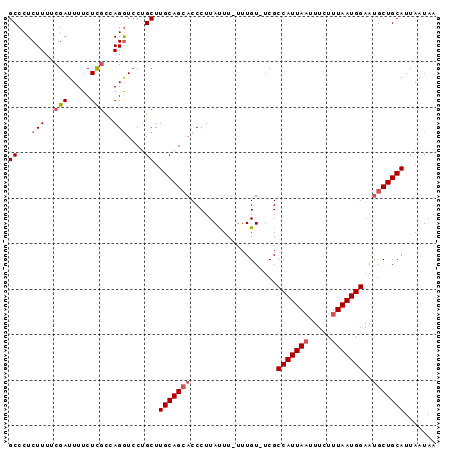
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:06:24 2006