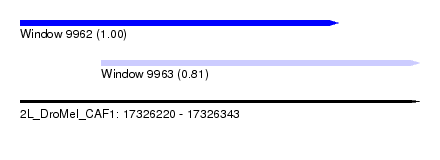

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 17,326,220 – 17,326,343 |
| Length | 123 |
| Max. P | 0.999749 |

| Location | 17,326,220 – 17,326,318 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 106 |
| Reading direction | forward |
| Mean pairwise identity | 92.49 |
| Mean single sequence MFE | -22.71 |
| Consensus MFE | -19.36 |
| Energy contribution | -19.76 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.00 |
| Mean z-score | -3.41 |
| Structure conservation index | 0.85 |
| SVM decision value | 4.00 |
| SVM RNA-class probability | 0.999749 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 17326220 98 + 22407834 --------CCUUCUGCAUAUAUUGACCAACUCAAUGACCUGCAGCUACUACAAAAUGUUGCAGUUCAAAAGGCUACAACAGCUGCAACAAUAAUAACAAUAUCAAG --------....(((((..((((((.....))))))...)))))...........((((((((((..............))))))))))................. ( -20.44) >DroSec_CAF1 23803 98 + 1 --------CAUUCUGCAUAUAUUGACCAACUCAAUGACCUGCAGCUACUACAAAAUGUUGCAGUUGAAAAGGCUACAACAGCUGCAACAAUAAUAACAAUAUCAAG --------....(((((..((((((.....))))))...)))))...........(((((((((((............)))))))))))................. ( -23.70) >DroSim_CAF1 26713 98 + 1 --------CAUUCUGCAUAUAUUGACCAACUCAAUGACCUGCAGCUACUACAAAAUGUUGCAGUUGAAAAGGCUACAACAGCUGCAACAAUAAUAACAAUAUCAAG --------....(((((..((((((.....))))))...)))))...........(((((((((((............)))))))))))................. ( -23.70) >DroEre_CAF1 25934 106 + 1 CACUCUGGCACUCUGCAUAUAUUGACCAACCCAAUGACCUGCAGCUACUGCAAAAUGUUGCAGUUCAAGAGGCUACGACAGCUGCAACAAUAAUAACAAUAUCAAG .......(((..(((((..(((((.......)))))...)))))....)))....((((((((((...(......)...))))))))))................. ( -22.90) >DroYak_CAF1 18774 98 + 1 --------CAUUCUGCAUAUAUUGACCAACUCUAUGCCCUGCAGCUACUGCAAAAUGUUGCAGUUCAAAAGGCUGCAACAGCUGCAACAAUAAUAACAAUAUCAAG --------......(((((.............)))))..(((((((..........(((((((((.....)))))))))))))))).................... ( -22.82) >consensus ________CAUUCUGCAUAUAUUGACCAACUCAAUGACCUGCAGCUACUACAAAAUGUUGCAGUUCAAAAGGCUACAACAGCUGCAACAAUAAUAACAAUAUCAAG ............(((((..((((((.....))))))...)))))...........((((((((((..............))))))))))................. (-19.36 = -19.76 + 0.40)

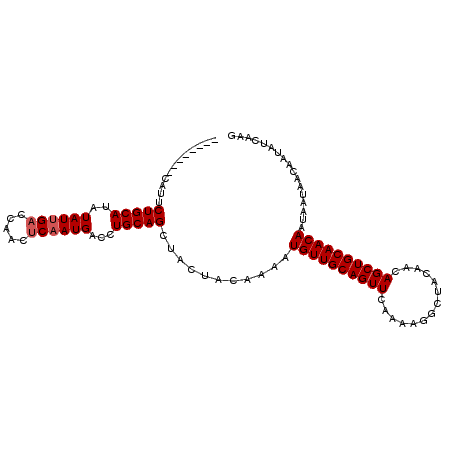

| Location | 17,326,245 – 17,326,343 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 95.45 |
| Mean single sequence MFE | -21.92 |
| Consensus MFE | -17.44 |
| Energy contribution | -17.48 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.93 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.65 |
| SVM RNA-class probability | 0.810535 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 17326245 98 + 22407834 AUGACCUGCAGCUACUACAAAAUGUUGCAGUUCAAAAGGCUACAACAGCUGCAACAAUAAUAACAAUAUCAAGUGUGCGUACUUUAUGACUUGACCUG .(((.(((((((...........))))))).)))...((((.....))))..................((((((...((((...)))))))))).... ( -19.20) >DroSec_CAF1 23828 96 + 1 AUGACCUGCAGCUACUACAAAAUGUUGCAGUUGAAAAGGCUACAACAGCUGCAACAAUAAUAACAAUAUCAAGU--GCGUACUUUAUGACUUGACCUG ........(((...........(((((((((((............)))))))))))............((((((--.((((...)))))))))).))) ( -22.10) >DroSim_CAF1 26738 96 + 1 AUGACCUGCAGCUACUACAAAAUGUUGCAGUUGAAAAGGCUACAACAGCUGCAACAAUAAUAACAAUAUCAAGU--GCGUACUUUAUGACUUGACCUG ........(((...........(((((((((((............)))))))))))............((((((--.((((...)))))))))).))) ( -22.10) >DroEre_CAF1 25967 96 + 1 AUGACCUGCAGCUACUGCAAAAUGUUGCAGUUCAAGAGGCUACGACAGCUGCAACAAUAAUAACAAUAUCAAGU--GCGUACUUUAUGACUUGACCUG ......((((((((((((((....)))))))......(....)...)))))))...............((((((--.((((...)))))))))).... ( -24.00) >DroYak_CAF1 18799 96 + 1 AUGCCCUGCAGCUACUGCAAAAUGUUGCAGUUCAAAAGGCUGCAACAGCUGCAACAAUAAUAACAAUAUCAAGU--GCGUACUUUAUGACUUCACCCG ......(((((((..........(((((((((.....)))))))))))))))).................((((--.((((...))))))))...... ( -22.20) >consensus AUGACCUGCAGCUACUACAAAAUGUUGCAGUUCAAAAGGCUACAACAGCUGCAACAAUAAUAACAAUAUCAAGU__GCGUACUUUAUGACUUGACCUG ......(((((((..........((((.((((.....)))).)))))))))))...............((((((...((((...)))))))))).... (-17.44 = -17.48 + 0.04)


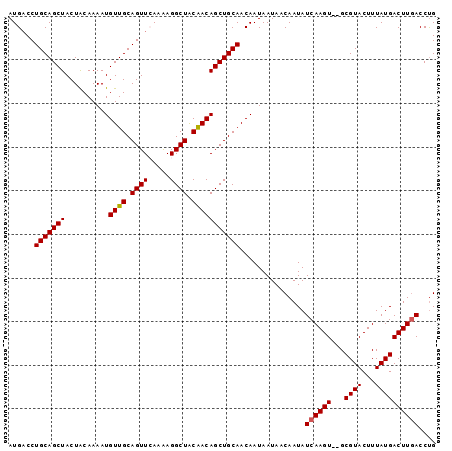
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:05:58 2006