
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 1,833,817 – 1,833,914 |
| Length | 97 |
| Max. P | 0.506254 |

| Location | 1,833,817 – 1,833,914 |
|---|---|
| Length | 97 |
| Sequences | 6 |
| Columns | 103 |
| Reading direction | reverse |
| Mean pairwise identity | 71.98 |
| Mean single sequence MFE | -25.82 |
| Consensus MFE | -9.38 |
| Energy contribution | -9.47 |
| Covariance contribution | 0.08 |
| Combinations/Pair | 1.20 |
| Mean z-score | -1.87 |
| Structure conservation index | 0.36 |
| SVM decision value | -0.05 |
| SVM RNA-class probability | 0.506254 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
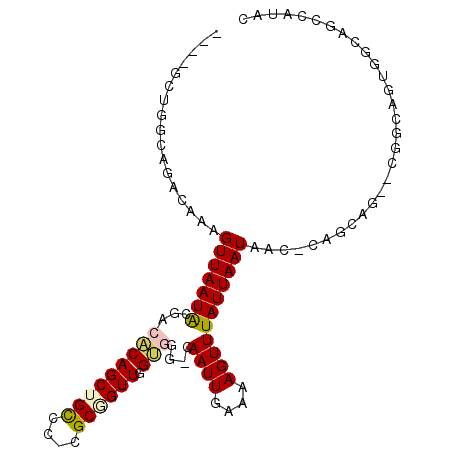
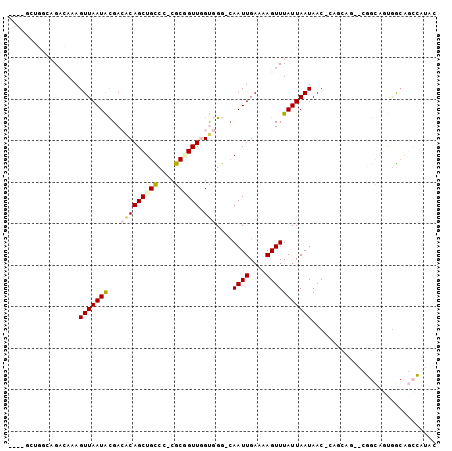
>2L_DroMel_CAF1 1833817 97 - 22407834 ----GCUGGCAGACAAAGUUAAUACGACACAGCUGCCCUCGCGGUUGGUGGG-CAAUUGAAAAGUUUAUUAAUAAC-CCGCAGAACGGCAGUGGCAGCCAUAC ----..((((.......(((.....))).....((((((.((.(((.(((((-..(((((........)))))..)-))))..))).)))).))))))))... ( -29.40) >DroGri_CAF1 14148 82 - 1 -------CCGAGCAAAAGUUAAUGUGA-ACAGCUGUCC-CGCUGUUCGCUGCAAAAUUGAAAAGUUUAUUAAUAAC-AAAAAU----GAGCCGACG------- -------.((.((....((....((((-(((((.....-.))))))))).)).(((((....))))).........-......----..))))...------- ( -20.40) >DroSim_CAF1 2085 97 - 1 ----GUUGGCAGACAAAGUUAAUACGACACAGCUGCCCUCGCGGUUGGUGGG-CAAUUGAAAAGUUUAUUAAUAAC-CCGCAGAACGGCAGUGGCAGCCAUAC ----((((((.......))))))........((((((((.((.(((.(((((-..(((((........)))))..)-))))..))).)))).))))))..... ( -31.20) >DroYak_CAF1 2138 97 - 1 ----GCUGGCAGACAAAGUUAAUACGACACAGCUGCCCUCGCGGUUGGUGGG-CAAUUGAAAAGUUUAUUAAUAAC-CCGCAGAACGGCAGUGGCAGCCAUAC ----..((((.......(((.....))).....((((((.((.(((.(((((-..(((((........)))))..)-))))..))).)))).))))))))... ( -29.40) >DroMoj_CAF1 14930 89 - 1 GUAGACGAAAAGAAAAAGUUAAUGCGA-GCAGCUGUCA-CGCUGUUUGCUGG-CAAUUGAAAAGUUUAUUAAUAACCAAAAUC----GUGCUGGCA------- .(((((((...............((((-(((((.....-.)))))))))(((-..(((((........)))))..)))...))----)).)))...------- ( -22.10) >DroAna_CAF1 1995 90 - 1 -------GACAGACAAAGUUAAUACGACACAGCUGCCC-CGCAGUUGGUGGG-CAAUUGAAAAGUUUAUUAAUAAC-CACCA---AGGCAGCAAAAGCUAUGG -------..........(((.....)))...((((((.-.....(((((((.-..(((((........)))))..)-)))))---)))))))........... ( -22.40) >consensus ____GCUGGCAGACAAAGUUAAUACGACACAGCUGCCC_CGCGGUUGGUGGG_CAAUUGAAAAGUUUAUUAAUAAC_CAGCAG__CGGCAGUGGCAGCCAUAC .................(((((((...(((((((((....)))))).)))....((((....))))))))))).............................. ( -9.38 = -9.47 + 0.08)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:41:11 2006