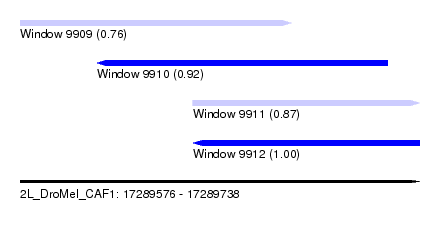
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 17,289,576 – 17,289,738 |
| Length | 162 |
| Max. P | 0.995843 |

| Location | 17,289,576 – 17,289,686 |
|---|---|
| Length | 110 |
| Sequences | 5 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 90.25 |
| Mean single sequence MFE | -39.80 |
| Consensus MFE | -31.60 |
| Energy contribution | -33.40 |
| Covariance contribution | 1.80 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.64 |
| Structure conservation index | 0.79 |
| SVM decision value | 0.50 |
| SVM RNA-class probability | 0.759451 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 17289576 110 + 22407834 AGGAUAUGUCCU---------CCUUCACUUGUUCCUCCACGCGAUGAUGACGUUGCGAUGUUGUGUCCUGUUGCGAGGUGAAGAUGAUGAUGAUCAUCAACAUCAUGAUGAUGGAGAUA ((((....))))---------.............(((((((..((((((((.((((((((........)))))))).))...((((((....))))))..))))))..)).)))))... ( -31.90) >DroSec_CAF1 139641 119 + 1 AGGAGAUGUCCUCAUUGCUCUCCUCCACUUGUUCCUCCACGCGAUGAUGACGUUGCGAUGUUGUGACCCGUUGCGAGGUGAAGACGAUGAUGAUCAUCAACAUCAUGGUGAUGGAGAUA ((((((.((.......))))))))..........((((((((.((((((((.((((((((........)))))))).))......((((.....))))..)))))).))).)))))... ( -42.10) >DroSim_CAF1 132794 119 + 1 AGGAGAUGUCCUCAUUGCUCUCCUCCACUUGUUCCUCCACGCGAUGAUGACGUUGCGAUGUUGUGCCCCGUUGCGAGGUGAAGACGAUGAUGAUCAUCAACAUCAUGGUGAUGGAGAUA ((((((.((.......))))))))..........((((((((.((((((((.((((((((........)))))))).))......((((.....))))..)))))).))).)))))... ( -42.10) >DroEre_CAF1 127868 119 + 1 AGGAGAUGUCCUCCAUGCUCUCCUCCACUGGCUGCUCCACGCGAUGAUGGCGUUCCGAUGUUGUGCCCUGUUGCGAGGUGAAGAUGAUGAUGAUCAUCAACCUCACGGUGAUGGAGAUA .((((.....))))....(((((..(((((((.((....(((((((.(((....))).)))))))....)).))(((((...((((((....)))))).))))).)))))..))))).. ( -45.10) >DroYak_CAF1 130695 119 + 1 AGGAGAUGUCCUCAUUGCUCUCCUCCACUUGUUUCUCCACGCGAUGAUGACGUUCCGAUGUUGUGCACUGUUGCGAGGUGGAGACGAUGAUGAUCAUCAACAUCACGGUGAUGGAGAUA ..(((.....))).....(((((..((((.((((((((.((((((..(((((.........))).))..)))))).)).))))))((((.((.....)).))))..))))..))))).. ( -37.80) >consensus AGGAGAUGUCCUCAUUGCUCUCCUCCACUUGUUCCUCCACGCGAUGAUGACGUUGCGAUGUUGUGCCCUGUUGCGAGGUGAAGACGAUGAUGAUCAUCAACAUCAUGGUGAUGGAGAUA ((((((.((.......))))))))..........((((((((.((((((((.((((((((........)))))))).))......((((.....))))..)))))).))).)))))... (-31.60 = -33.40 + 1.80)



| Location | 17,289,607 – 17,289,725 |
|---|---|
| Length | 118 |
| Sequences | 5 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 95.34 |
| Mean single sequence MFE | -31.56 |
| Consensus MFE | -27.02 |
| Energy contribution | -27.10 |
| Covariance contribution | 0.08 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.67 |
| Structure conservation index | 0.86 |
| SVM decision value | 1.12 |
| SVM RNA-class probability | 0.918500 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 17289607 118 - 22407834 GCUGAAAAACAGUUGUUGACUGUUUGCCGGCAUCGCCUAUAUCUCCAUCAUCAUGAUGUUGAUGAUCAUCAUCAUCUUCACCUCGCAACAGGACACAACAUCGCAACGUCAUCAUCGC ((((..((((((((...))))))))..)))).....................((((((((((((((....))))))....(((......)))............))))))))...... ( -27.10) >DroSec_CAF1 139681 118 - 1 GCUGGAAAACAGUUGUUGACUGUUUGCCGGCAUCGCCUAUAUCUCCAUCACCAUGAUGUUGAUGAUCAUCAUCGUCUUCACCUCGCAACGGGUCACAACAUCGCAACGUCAUCAUCGC (((((.((((((((...)))))))).))))).......................(((((((.(((((.....((.........)).....))))))))))))................ ( -30.70) >DroSim_CAF1 132834 118 - 1 GCUGGAAAACAGUUGUUGACUGUUUGCCGGCAUCGCCUAUAUCUCCAUCACCAUGAUGUUGAUGAUCAUCAUCGUCUUCACCUCGCAACGGGGCACAACAUCGCAACGUCAUCAUCGC (((((.((((((((...)))))))).))))).....................((((((((((((((....))))))....(((((...)))))...........))))))))...... ( -32.50) >DroEre_CAF1 127908 118 - 1 GCUGGAAAACAGUUGUUGACUGUUUGCCGGCAUCGCCUAUAUCUCCAUCACCGUGAGGUUGAUGAUCAUCAUCAUCUUCACCUCGCAACAGGGCACAACAUCGGAACGCCAUCAUCGC (((((.((((((((...)))))))).)))))...((((((......))....(((((((.((((((....))))))...)))))))....))))........((....))........ ( -36.60) >DroYak_CAF1 130735 118 - 1 GCUGGAAAACAGUUGUUGACUGUUUGCCGGCAUCGCCUAUAUCUCCAUCACCGUGAUGUUGAUGAUCAUCAUCGUCUCCACCUCGCAACAGUGCACAACAUCGGAACGUCAUCAUCGC (((((.((((((((...)))))))).))))).....................((((((.(((((.((.................(((....))).(......))).))))).)))))) ( -30.90) >consensus GCUGGAAAACAGUUGUUGACUGUUUGCCGGCAUCGCCUAUAUCUCCAUCACCAUGAUGUUGAUGAUCAUCAUCGUCUUCACCUCGCAACAGGGCACAACAUCGCAACGUCAUCAUCGC (((((.((((((((...)))))))).))))).....................((((((((((((((....))))))....(((......)))............))))))))...... (-27.02 = -27.10 + 0.08)


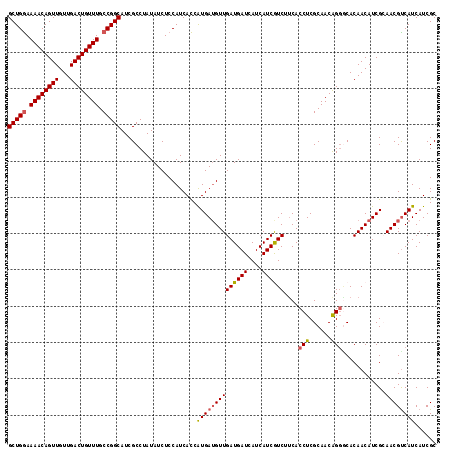
| Location | 17,289,646 – 17,289,738 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 92 |
| Reading direction | forward |
| Mean pairwise identity | 96.96 |
| Mean single sequence MFE | -25.64 |
| Consensus MFE | -24.34 |
| Energy contribution | -23.98 |
| Covariance contribution | -0.36 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.34 |
| Structure conservation index | 0.95 |
| SVM decision value | 0.89 |
| SVM RNA-class probability | 0.874618 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 17289646 92 + 22407834 GAAGAUGAUGAUGAUCAUCAACAUCAUGAUGAUGGAGAUAUAGGCGAUGCCGGCAAACAGUCAACAACUGUUUUUCAGCUGCAGUUUUCCUG ...((((.((((....)))).))))........(((((.....(((..((.((.(((((((.....))))))).)).)))))...))))).. ( -23.50) >DroSec_CAF1 139720 92 + 1 GAAGACGAUGAUGAUCAUCAACAUCAUGGUGAUGGAGAUAUAGGCGAUGCCGGCAAACAGUCAACAACUGUUUUCCAGCUGCAGUUUUCCUG ....((.((((((........)))))).))...(((((.....(((..((.((.(((((((.....))))))).)).)))))...))))).. ( -26.40) >DroSim_CAF1 132873 92 + 1 GAAGACGAUGAUGAUCAUCAACAUCAUGGUGAUGGAGAUAUAGGCGAUGCCGGCAAACAGUCAACAACUGUUUUCCAGCUGCAGUUUUCCUG ....((.((((((........)))))).))...(((((.....(((..((.((.(((((((.....))))))).)).)))))...))))).. ( -26.40) >DroEre_CAF1 127947 92 + 1 GAAGAUGAUGAUGAUCAUCAACCUCACGGUGAUGGAGAUAUAGGCGAUGCCGGCAAACAGUCAACAACUGUUUUCCAGCUGCAGUUUUCCUG ...((((((....)))))).(((....)))...(((((.....(((..((.((.(((((((.....))))))).)).)))))...))))).. ( -26.20) >DroYak_CAF1 130774 92 + 1 GGAGACGAUGAUGAUCAUCAACAUCACGGUGAUGGAGAUAUAGGCGAUGCCGGCAAACAGUCAACAACUGUUUUCCAGCUGCAGUUUUCCUG ((((((((((.....))))..((((.....)))).........(((..((.((.(((((((.....))))))).)).))))).))))))... ( -25.70) >consensus GAAGACGAUGAUGAUCAUCAACAUCAUGGUGAUGGAGAUAUAGGCGAUGCCGGCAAACAGUCAACAACUGUUUUCCAGCUGCAGUUUUCCUG ((((((((((.....))))..((((.....)))).........(((..((.((.(((((((.....))))))).)).))))).))))))... (-24.34 = -23.98 + -0.36)

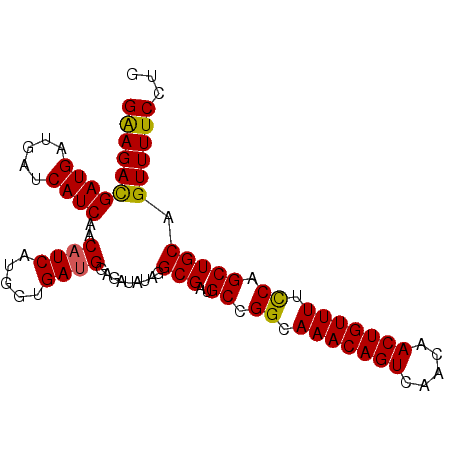
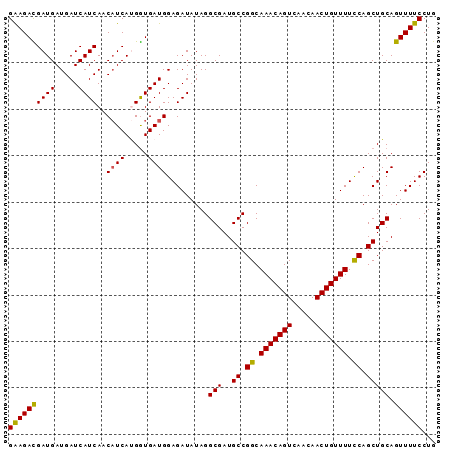
| Location | 17,289,646 – 17,289,738 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 92 |
| Reading direction | reverse |
| Mean pairwise identity | 96.96 |
| Mean single sequence MFE | -25.26 |
| Consensus MFE | -23.58 |
| Energy contribution | -23.74 |
| Covariance contribution | 0.16 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.94 |
| Structure conservation index | 0.93 |
| SVM decision value | 2.62 |
| SVM RNA-class probability | 0.995843 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 17289646 92 - 22407834 CAGGAAAACUGCAGCUGAAAAACAGUUGUUGACUGUUUGCCGGCAUCGCCUAUAUCUCCAUCAUCAUGAUGUUGAUGAUCAUCAUCAUCUUC ..(((.....((.((((..((((((((...))))))))..))))...)).......)))........((((.((((....)))).))))... ( -22.50) >DroSec_CAF1 139720 92 - 1 CAGGAAAACUGCAGCUGGAAAACAGUUGUUGACUGUUUGCCGGCAUCGCCUAUAUCUCCAUCACCAUGAUGUUGAUGAUCAUCAUCGUCUUC ..(((.....((.(((((.((((((((...)))))))).)))))...)).......)))........((((.((((....)))).))))... ( -25.10) >DroSim_CAF1 132873 92 - 1 CAGGAAAACUGCAGCUGGAAAACAGUUGUUGACUGUUUGCCGGCAUCGCCUAUAUCUCCAUCACCAUGAUGUUGAUGAUCAUCAUCGUCUUC ..(((.....((.(((((.((((((((...)))))))).)))))...)).......)))........((((.((((....)))).))))... ( -25.10) >DroEre_CAF1 127947 92 - 1 CAGGAAAACUGCAGCUGGAAAACAGUUGUUGACUGUUUGCCGGCAUCGCCUAUAUCUCCAUCACCGUGAGGUUGAUGAUCAUCAUCAUCUUC (((.....)))..(((((.((((((((...)))))))).)))))..............(((((((....)).)))))............... ( -28.30) >DroYak_CAF1 130774 92 - 1 CAGGAAAACUGCAGCUGGAAAACAGUUGUUGACUGUUUGCCGGCAUCGCCUAUAUCUCCAUCACCGUGAUGUUGAUGAUCAUCAUCGUCUCC ..(((.....((.(((((.((((((((...)))))))).)))))...)).......)))......(.((((.((((....)))).)))).). ( -25.30) >consensus CAGGAAAACUGCAGCUGGAAAACAGUUGUUGACUGUUUGCCGGCAUCGCCUAUAUCUCCAUCACCAUGAUGUUGAUGAUCAUCAUCGUCUUC (((.....)))..(((((.((((((((...)))))))).)))))..............((((.....))))..((((((....))))))... (-23.58 = -23.74 + 0.16)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:05:06 2006