
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 17,201,185 – 17,201,278 |
| Length | 93 |
| Max. P | 0.898899 |

| Location | 17,201,185 – 17,201,278 |
|---|---|
| Length | 93 |
| Sequences | 6 |
| Columns | 112 |
| Reading direction | reverse |
| Mean pairwise identity | 74.40 |
| Mean single sequence MFE | -18.44 |
| Consensus MFE | -13.10 |
| Energy contribution | -13.10 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.32 |
| Structure conservation index | 0.71 |
| SVM decision value | 1.01 |
| SVM RNA-class probability | 0.898899 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
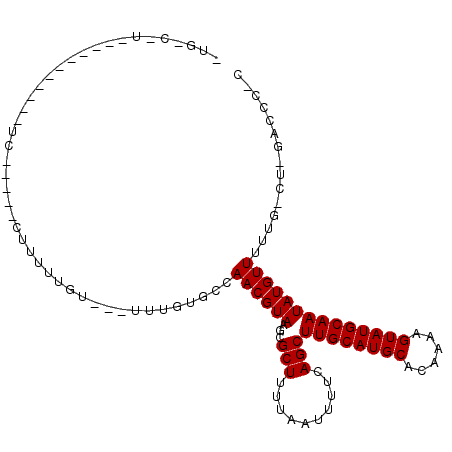
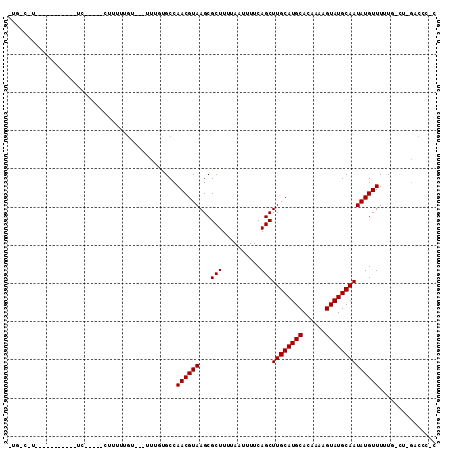
>2L_DroMel_CAF1 17201185 93 - 22407834 UUGGCUU----------UUC-----CCUCCUGUU-UUUUGUGCCAACGUAAGCGCUUUUAAUUUUCAGCUUGCAUGCACAAAAGUAUGCAAUAUGUUUUUG-CU-GACCC-C (..((..----------...-----.....((((-((((((((....((((((..............))))))..)))))))))...)))..........)-).-.)...-. ( -19.39) >DroVir_CAF1 141047 80 - 1 ---------------------------UUUCG----UUUUCGCCAACGUAAGCGCUUUUAAUUUUCAGCUUGCAUGCACAAAAGUAUGCAAUAUGUUUUUUGCUCGACCCC- ---------------------------..(((----........((((((...(((..........)))((((((((......)))))))))))))).......)))....- ( -14.76) >DroGri_CAF1 162955 106 - 1 -UUCCAGUCUUGCAGU-UUCGGGUGUUUUUUC----UUUUAGCCAACGUAAGCGCUUUUAAUUUUCAGCUUGCAUGCACAAAAGUAUGCAAUAUGUUUUUUACUCGAACCCC -...............-((((((((.......----....(((...(....).))).........((..((((((((......))))))))..)).....)))))))).... ( -19.40) >DroEre_CAF1 44801 91 - 1 UUGGCUU-----------UC-----CCUUCUGUU--UUUGUGCCAACGUAAGCGCUUUUAAUUUUCAGCUUGCAUGCACAAAAGUAUGCAAUAUGUUUUUG-CU-GACCC-C ..((...-----------..-----.......((--(((((((....((((((..............))))))..)))))))))...((((.......)))-).-..)).-. ( -18.84) >DroWil_CAF1 55371 77 - 1 ------------------UU-----CUUUUUGU----UUUCGCCAACGUAAGCGCUUUUAAUUUUCAGCUUGCAUGCACAAAAGUAUGCAAUAUGUUUUA-------CCC-U ------------------..-----........----.......((((((...(((..........)))((((((((......))))))))))))))...-------...-. ( -13.10) >DroAna_CAF1 119907 100 - 1 UGGUCCU----GGCGAGGUC-----CUUUUUGUUAUUUUGUGCCAACGUAAGCGCUUUUAAUUUUCAGCUUGCAUGCACAAAAGUAUGCAAUAUGUUUUUG-UA-GCCAC-C ((((...----.((((((..-----....((((..((((((((....((((((..............))))))..))))))))....)))).....)))))-).-)))).-. ( -25.14) >consensus _UG_C_U___________UC_____CUUUUUGU___UUUGUGCCAACGUAAGCGCUUUUAAUUUUCAGCUUGCAUGCACAAAAGUAUGCAAUAUGUUUUUG_CU_GACCC_C ............................................((((((...(((..........)))((((((((......))))))))))))))............... (-13.10 = -13.10 + -0.00)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:04:14 2006