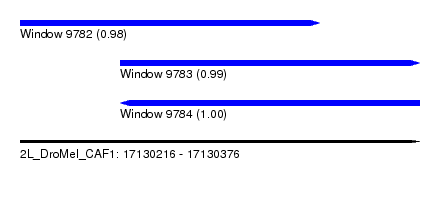

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 17,130,216 – 17,130,376 |
| Length | 160 |
| Max. P | 0.999704 |

| Location | 17,130,216 – 17,130,336 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 95.58 |
| Mean single sequence MFE | -20.56 |
| Consensus MFE | -16.72 |
| Energy contribution | -18.00 |
| Covariance contribution | 1.28 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.47 |
| Structure conservation index | 0.81 |
| SVM decision value | 1.81 |
| SVM RNA-class probability | 0.978267 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 17130216 120 + 22407834 CGCCAAACAAGAAGUUUACGCAUUUCAUACUCAAACAUUUUUACAGAAUUCUUUGUGUGGACGGAAAUCCAGCUCCCUCAAAACAAAUUCCUGGGAAAAUCCCAAGAAAAACAAUCAACA .((..........(((((((((.(((...................))).....)))))))))((....)).))..............(((.((((.....)))).)))............ ( -19.81) >DroSec_CAF1 98700 120 + 1 CGCCAAACAAAAAGUUUACGCAUUUCAUACUCAAACAUUUUUACAGAAUUGUUUCUGUGGACGGAAAUCCAGCUCCCUCAAAACAAAUUCCUGGGAAAAUCCCAAGAAAAACAAUCAACA .............((((..((..................(((((((((....))))))))).((....)).))..............(((.((((.....)))).)))))))........ ( -22.00) >DroSim_CAF1 110645 120 + 1 CGCCAAACAAAAAGUUUACGCAUUUCAUACUCAAACAUUUUUACAGAAUUGUUUCUGUGGACGGAAAUCCAGCUCCCUCAAAACAAAUUCCUGGGAAAAUCCCAAGAAAAACAAUCAACA .............((((..((..................(((((((((....))))))))).((....)).))..............(((.((((.....)))).)))))))........ ( -22.00) >DroEre_CAF1 96143 120 + 1 CGCCAAACAAAAAGUAUACGCAUUUCAUACUCAAACAUUUUUACAGAAUUGUUUCUGUGGACGGAAAUCCAGCUCCCUCAAAACAAAUUCCUGGGAAAAUCCCAAGAAAAACAAUCAACA ...................((..................(((((((((....))))))))).((....)).))..............(((.((((.....)))).)))............ ( -20.20) >DroYak_CAF1 96826 116 + 1 CGCCAAACAAAAAGUACACGCAUGUUUUACUCAAACAUUUUUACAGAAUUGUUUUUGUG----GAAAUCCAGCUCCCUCAAAACAAAUUCCUGGGAAAAUCCCAAGAAAAACAAUCAACA ...................((.(((((.....)))))(((((((((((....)))))))----))))....))..............(((.((((.....)))).)))............ ( -18.80) >consensus CGCCAAACAAAAAGUUUACGCAUUUCAUACUCAAACAUUUUUACAGAAUUGUUUCUGUGGACGGAAAUCCAGCUCCCUCAAAACAAAUUCCUGGGAAAAUCCCAAGAAAAACAAUCAACA .............((((..((..................(((((((((....))))))))).((....)).))..............(((.((((.....)))).)))))))........ (-16.72 = -18.00 + 1.28)

| Location | 17,130,256 – 17,130,376 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 96.92 |
| Mean single sequence MFE | -22.10 |
| Consensus MFE | -17.96 |
| Energy contribution | -19.04 |
| Covariance contribution | 1.08 |
| Combinations/Pair | 1.04 |
| Mean z-score | -3.00 |
| Structure conservation index | 0.81 |
| SVM decision value | 2.00 |
| SVM RNA-class probability | 0.985113 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
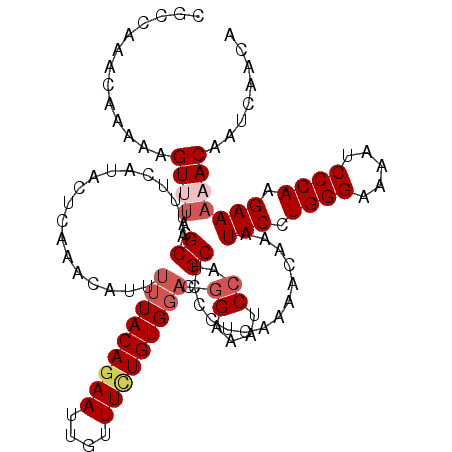
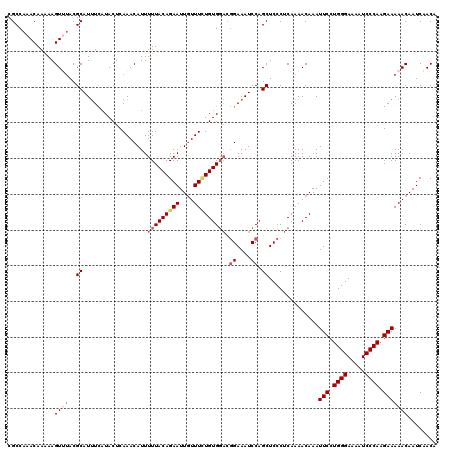
>2L_DroMel_CAF1 17130256 120 + 22407834 UUACAGAAUUCUUUGUGUGGACGGAAAUCCAGCUCCCUCAAAACAAAUUCCUGGGAAAAUCCCAAGAAAAACAAUCAACAAAUUGAAAGCCAAAGACCAAAACAAUCGAACAAGUCCAUU .((((((....))))))(((((((....)).(((.............(((.((((.....)))).)))...((((......))))..))).......................))))).. ( -22.10) >DroSec_CAF1 98740 120 + 1 UUACAGAAUUGUUUCUGUGGACGGAAAUCCAGCUCCCUCAAAACAAAUUCCUGGGAAAAUCCCAAGAAAAACAAUCAACAAAUUGAAAGCCAAAGACCAAAACAAUCGAACAAGUCCAUU ..((((((....))))))((((((....)).(((.............(((.((((.....)))).)))...((((......))))..))).......................))))... ( -24.60) >DroSim_CAF1 110685 120 + 1 UUACAGAAUUGUUUCUGUGGACGGAAAUCCAGCUCCCUCAAAACAAAUUCCUGGGAAAAUCCCAAGAAAAACAAUCAACAAAUUGAAAGCCAAAGACCAAAACAAUCGAACAAGUCCAUU ..((((((....))))))((((((....)).(((.............(((.((((.....)))).)))...((((......))))..))).......................))))... ( -24.60) >DroEre_CAF1 96183 120 + 1 UUACAGAAUUGUUUCUGUGGACGGAAAUCCAGCUCCCUCAAAACAAAUUCCUGGGAAAAUCCCAAGAAAAACAAUCAACAAAUUGAAAGCCAAAGACCAAGACAAUCGAACAAAUCCAUU ..((((((....))))))(((.((....)).(((.............(((.((((.....)))).)))...((((......))))..)))........................)))... ( -20.80) >DroYak_CAF1 96866 116 + 1 UUACAGAAUUGUUUUUGUG----GAAAUCCAGCUCCCUCAAAACAAAUUCCUGGGAAAAUCCCAAGAAAAACAAUCAACAAAUUGAAAGCCAAAGACCAAAACAAUCGAACAAAUCCAUU .....((.(((((((.(((----(....)).(((.............(((.((((.....)))).)))...((((......))))..))).....)).)))))))))............. ( -18.40) >consensus UUACAGAAUUGUUUCUGUGGACGGAAAUCCAGCUCCCUCAAAACAAAUUCCUGGGAAAAUCCCAAGAAAAACAAUCAACAAAUUGAAAGCCAAAGACCAAAACAAUCGAACAAGUCCAUU ..((((((....))))))((((((....)).(((.............(((.((((.....)))).)))...((((......))))..))).......................))))... (-17.96 = -19.04 + 1.08)

| Location | 17,130,256 – 17,130,376 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 96.92 |
| Mean single sequence MFE | -32.42 |
| Consensus MFE | -29.74 |
| Energy contribution | -30.42 |
| Covariance contribution | 0.68 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.72 |
| Structure conservation index | 0.92 |
| SVM decision value | 3.92 |
| SVM RNA-class probability | 0.999704 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
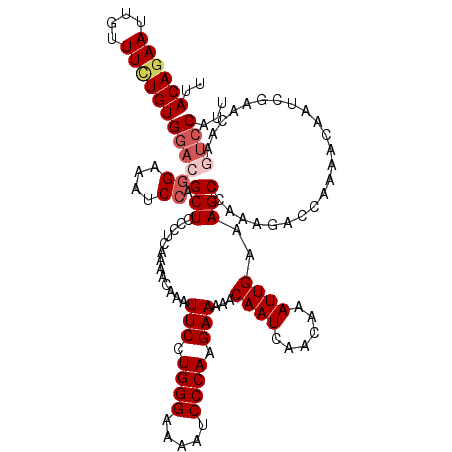
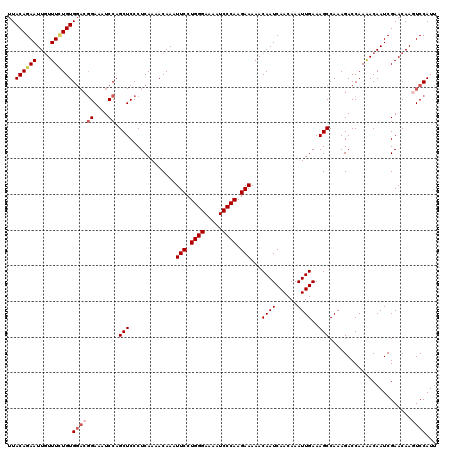
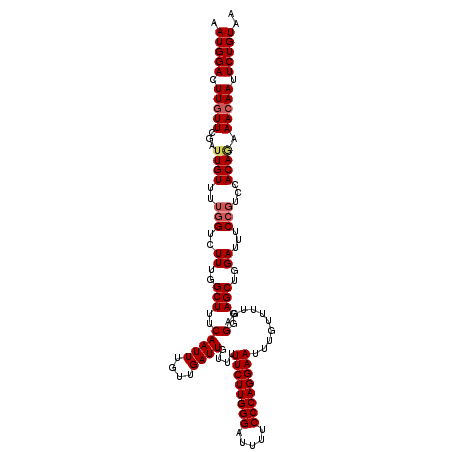
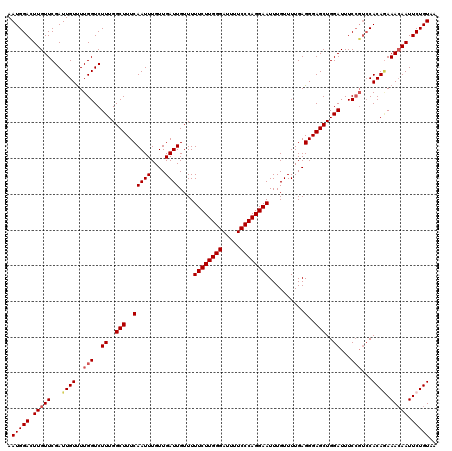
>2L_DroMel_CAF1 17130256 120 - 22407834 AAUGGACUUGUUCGAUUGUUUUGGUCUUUGGCUUUCAAUUUGUUGAUUGUUUUUCUUGGGAUUUUCCCAGGAAUUUGUUUUGAGGGAGCUGGAUUUCCGUCCACACAAAGAAUUCUGUAA .(((((...((((..((((..(((.(...((((..(((((....))))....((((((((.....))))))))..........)..))))((....))).))).)))).))))))))).. ( -30.10) >DroSec_CAF1 98740 120 - 1 AAUGGACUUGUUCGAUUGUUUUGGUCUUUGGCUUUCAAUUUGUUGAUUGUUUUUCUUGGGAUUUUCCCAGGAAUUUGUUUUGAGGGAGCUGGAUUUCCGUCCACAGAAACAAUUCUGUAA ...((((......((((.....))))((..(((..(((((....))))....((((((((.....))))))))..........)..)))..)).....))))((((((....)))))).. ( -34.10) >DroSim_CAF1 110685 120 - 1 AAUGGACUUGUUCGAUUGUUUUGGUCUUUGGCUUUCAAUUUGUUGAUUGUUUUUCUUGGGAUUUUCCCAGGAAUUUGUUUUGAGGGAGCUGGAUUUCCGUCCACAGAAACAAUUCUGUAA ...((((......((((.....))))((..(((..(((((....))))....((((((((.....))))))))..........)..)))..)).....))))((((((....)))))).. ( -34.10) >DroEre_CAF1 96183 120 - 1 AAUGGAUUUGUUCGAUUGUCUUGGUCUUUGGCUUUCAAUUUGUUGAUUGUUUUUCUUGGGAUUUUCCCAGGAAUUUGUUUUGAGGGAGCUGGAUUUCCGUCCACAGAAACAAUUCUGUAA .(((((.(((((...((((..(((..((..(((..(((((....))))....((((((((.....))))))))..........)..)))..))...)))...)))).))))).))))).. ( -32.80) >DroYak_CAF1 96866 116 - 1 AAUGGAUUUGUUCGAUUGUUUUGGUCUUUGGCUUUCAAUUUGUUGAUUGUUUUUCUUGGGAUUUUCCCAGGAAUUUGUUUUGAGGGAGCUGGAUUUC----CACAAAAACAAUUCUGUAA .(((((.(((((...((((...((..((..(((..(((((....))))....((((((((.....))))))))..........)..)))..))...)----))))).))))).))))).. ( -31.00) >consensus AAUGGACUUGUUCGAUUGUUUUGGUCUUUGGCUUUCAAUUUGUUGAUUGUUUUUCUUGGGAUUUUCCCAGGAAUUUGUUUUGAGGGAGCUGGAUUUCCGUCCACAGAAACAAUUCUGUAA .(((((.(((((...((((..(((..((..(((..(((((....))))....((((((((.....))))))))..........)..)))..))...)))...)))).))))).))))).. (-29.74 = -30.42 + 0.68)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:03:01 2006