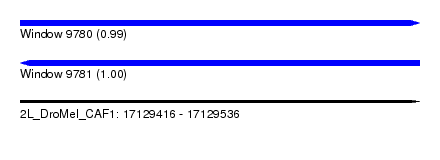

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 17,129,416 – 17,129,536 |
| Length | 120 |
| Max. P | 0.998829 |

| Location | 17,129,416 – 17,129,536 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 97.66 |
| Mean single sequence MFE | -30.34 |
| Consensus MFE | -28.50 |
| Energy contribution | -28.70 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.84 |
| Structure conservation index | 0.94 |
| SVM decision value | 2.24 |
| SVM RNA-class probability | 0.990904 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 17129416 120 + 22407834 GCAGUAUAGGGGGUGCCCCAAAACAACAACUUAAUGUCCAAAUGAAAUAAGUGAAAAUUUCACACCGAAAUUAAUAAAGACAUUAGUGCGUGUGGGCGUGGCUCAUUUGGGGCAUUUAAC ..........(((((((((((((((...(((.(((((((....)......((((.....))))...............)))))))))...)))((((...)))).))))))))))))... ( -32.70) >DroSec_CAF1 97852 119 + 1 GCCGUAUAGG-GGAGCCCCAAAACAACAACUUAAUGUCCAAAUGAAAUAAGUGAAAAUUUCACACCGAAAUUAAUAAAGACAUUAGUGCGUGUGGGCGUGGCUCAUUUGGGGCAUUUAAC .((.....))-...(((((((((((...(((.(((((((....)......((((.....))))...............)))))))))...)))((((...)))).))))))))....... ( -30.90) >DroSim_CAF1 109794 119 + 1 GCAGUAUAGGGGGAGCCCCAAAACAACAACUUAAUGUCCAAAUGAAAUAAGUGAAAAUUUCACACCGAAAUUAAUAAAGACAUUAGUGCGUGUGGGCGUGGCUC-UAUGGGGCAUUUAAC ((..(((((((...((((....(((...(((.(((((((....)......((((.....))))...............)))))))))...)))))))....)))-))))..))....... ( -29.10) >DroEre_CAF1 95277 119 + 1 GCAGUAAAGG-GGAGCCCCAAAACAACAACUUAAUGUCCAAAUGAAAUAAGUGAAAAUUUCACACCGAAAUUAAUAAAGACAUUAGUGCGUGUGGGCGUGGCUCAUUUGGGGCAUUUAAC ..........-...(((((((((((...(((.(((((((....)......((((.....))))...............)))))))))...)))((((...)))).))))))))....... ( -29.50) >DroYak_CAF1 95986 120 + 1 GCAGUAAAGGGGGAGCCCCAAAACAACAACUUAAUGUCCAAAUGAAAUAAGUGAAAAUUUCACACCGAAAUUAAUAAAGACAUUAGUGCGUGUGGGCGUGGCUCAUUUGGGGCAUUUAAC ..............(((((((((((...(((.(((((((....)......((((.....))))...............)))))))))...)))((((...)))).))))))))....... ( -29.50) >consensus GCAGUAUAGGGGGAGCCCCAAAACAACAACUUAAUGUCCAAAUGAAAUAAGUGAAAAUUUCACACCGAAAUUAAUAAAGACAUUAGUGCGUGUGGGCGUGGCUCAUUUGGGGCAUUUAAC ..............(((((((((((...(((.(((((((....)......((((.....))))...............)))))))))...)))((((...)))).))))))))....... (-28.50 = -28.70 + 0.20)


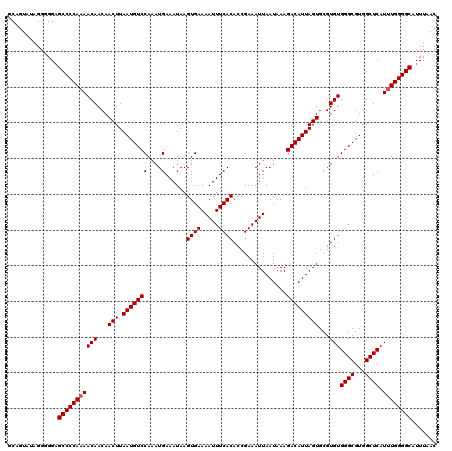
| Location | 17,129,416 – 17,129,536 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 97.66 |
| Mean single sequence MFE | -29.11 |
| Consensus MFE | -28.55 |
| Energy contribution | -28.75 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.98 |
| SVM decision value | 3.24 |
| SVM RNA-class probability | 0.998829 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
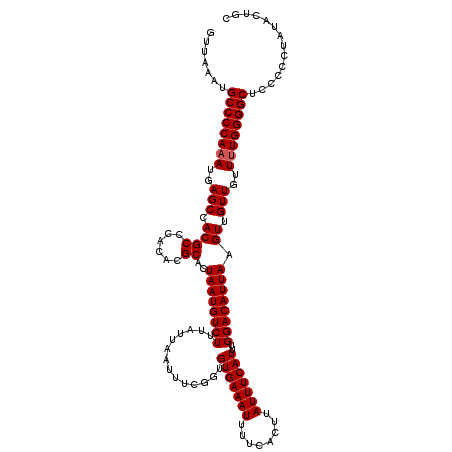
>2L_DroMel_CAF1 17129416 120 - 22407834 GUUAAAUGCCCCAAAUGAGCCACGCCCACACGCACUAAUGUCUUUAUUAAUUUCGGUGUGAAAUUUUCACUUAUUUCAUUUGGACAUUAAGUUGUUGUUUUGGGGCACCCCCUAUACUGC ......(((((((((..(((.((((......))..((((((((..............(((((((........)))))))..)))))))).)).)))..)))))))))............. ( -30.49) >DroSec_CAF1 97852 119 - 1 GUUAAAUGCCCCAAAUGAGCCACGCCCACACGCACUAAUGUCUUUAUUAAUUUCGGUGUGAAAUUUUCACUUAUUUCAUUUGGACAUUAAGUUGUUGUUUUGGGGCUCC-CCUAUACGGC .......((((((((..(((.((((......))..((((((((..............(((((((........)))))))..)))))))).)).)))..))))))))...-.......... ( -29.29) >DroSim_CAF1 109794 119 - 1 GUUAAAUGCCCCAUA-GAGCCACGCCCACACGCACUAAUGUCUUUAUUAAUUUCGGUGUGAAAUUUUCACUUAUUUCAUUUGGACAUUAAGUUGUUGUUUUGGGGCUCCCCCUAUACUGC .......((((((.(-.(((.((((......))..((((((((..............(((((((........)))))))..)))))))).)).))).)..)))))).............. ( -27.19) >DroEre_CAF1 95277 119 - 1 GUUAAAUGCCCCAAAUGAGCCACGCCCACACGCACUAAUGUCUUUAUUAAUUUCGGUGUGAAAUUUUCACUUAUUUCAUUUGGACAUUAAGUUGUUGUUUUGGGGCUCC-CCUUUACUGC .......((((((((..(((.((((......))..((((((((..............(((((((........)))))))..)))))))).)).)))..))))))))...-.......... ( -29.29) >DroYak_CAF1 95986 120 - 1 GUUAAAUGCCCCAAAUGAGCCACGCCCACACGCACUAAUGUCUUUAUUAAUUUCGGUGUGAAAUUUUCACUUAUUUCAUUUGGACAUUAAGUUGUUGUUUUGGGGCUCCCCCUUUACUGC .......((((((((..(((.((((......))..((((((((..............(((((((........)))))))..)))))))).)).)))..)))))))).............. ( -29.29) >consensus GUUAAAUGCCCCAAAUGAGCCACGCCCACACGCACUAAUGUCUUUAUUAAUUUCGGUGUGAAAUUUUCACUUAUUUCAUUUGGACAUUAAGUUGUUGUUUUGGGGCUCCCCCUAUACUGC .......((((((((..(((.((((......))..((((((((..............(((((((........)))))))..)))))))).)).)))..)))))))).............. (-28.55 = -28.75 + 0.20)

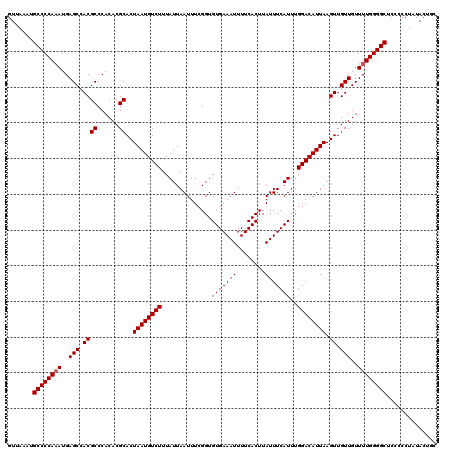
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:02:58 2006