
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 17,108,251 – 17,108,529 |
| Length | 278 |
| Max. P | 0.698178 |

| Location | 17,108,251 – 17,108,370 |
|---|---|
| Length | 119 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 93.15 |
| Mean single sequence MFE | -29.58 |
| Consensus MFE | -21.40 |
| Energy contribution | -21.64 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.21 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.10 |
| SVM RNA-class probability | 0.585364 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
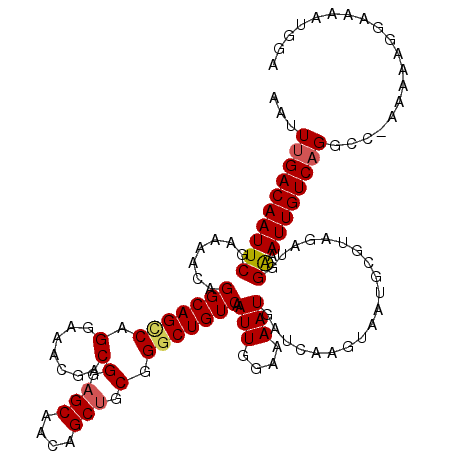
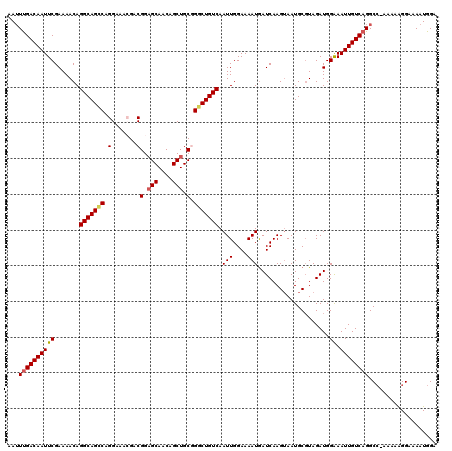
>2L_DroMel_CAF1 17108251 119 - 22407834 AAUUCGACAAUUCGAAAACAGGCAGCCAGGAAACGCCGGCGCAACAGCUGCGGGCUGUCAAUUGGAAAAUGAUCAAGAAAUGCUUAGAUGGAAAUUGUCAGGCC-AAAAAGGAAAAUGGA ..(((((....)))))....(((((((.(....)((.(((......))))).)))))))..((((....(((.(((......((.....))...))))))..))-))............. ( -30.40) >DroSec_CAF1 72315 119 - 1 AAUUUGACAAUUCGAAAACAGGCAGCCAGGAAACGACGGAGCAACAGCUGCAGGCUGUCAAUUGGAAAAUAAUCAAGUAAUGCGUAGAUGGGAAUUGUCAGGCC-AAAAAGGAAAAUGGA ..(((((((((((.......(((((((.(....)...(.(((....))).).))))))).(((....))).....................)))))))))))((-.....))........ ( -33.10) >DroSim_CAF1 85944 119 - 1 AAUUUGACAAUUCGAAAACAGGCAGCCAGGAAACGACGGAGCAACAGCUGCAGGCUGUCAAUUGGAAAAUGAUCAAGUAAUGCGUAGAUGGGAAUUGUCAGGCC-AAAAAGGAAAAUGGA ..(((((((((((.......(((((((.(....)...(.(((....))).).)))))))..........(.(((..(.....)...))).))))))))))))((-.....))........ ( -33.20) >DroEre_CAF1 73595 120 - 1 AAUUUGACAAUUCGAAAACAGGCAGUCAGGAAAAGCCGGAGCAACAGCUGCGGGCUGUCAAUUGGAAAAUGAUCAAGUAAUGCGUAGAUGGAAAUUGUCAGGCCGAAAAGGGAAAAUGGG ..(((((((((((.......(((((((.((.....))(.(((....))).).))))))).(((....)))....................).))))))))))((.....))......... ( -26.50) >DroYak_CAF1 74027 119 - 1 AAAUUGACAAUUCGAAAACAGGCAG-CAGGAAAAGGCGGAGCAACAGCUGCGGGCUGUCAAUUGGAAAAUGAUCAAGUAAUGCGUAGAUGGAAAUUGUCAGGCACAAAAAGGAAACUGGG ...((((((((((.......(((((-(.........((.(((....))).)).)))))).(((....)))....................).)))))))))........((....))... ( -24.70) >consensus AAUUUGACAAUUCGAAAACAGGCAGCCAGGAAACGACGGAGCAACAGCUGCGGGCUGUCAAUUGGAAAAUGAUCAAGUAAUGCGUAGAUGGAAAUUGUCAGGCC_AAAAAGGAAAAUGGA ...((((((((((.......(((((((.(.......)(.(((....))).).))))))).(((....)))....................)).))))))))................... (-21.40 = -21.64 + 0.24)

| Location | 17,108,370 – 17,108,489 |
|---|---|
| Length | 119 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 98.89 |
| Mean single sequence MFE | -21.70 |
| Consensus MFE | -20.77 |
| Energy contribution | -21.10 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.69 |
| Structure conservation index | 0.96 |
| SVM decision value | 0.29 |
| SVM RNA-class probability | 0.670621 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
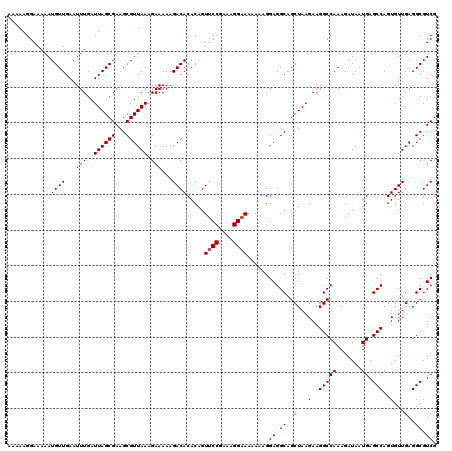
>2L_DroMel_CAF1 17108370 119 + 22407834 AAAAAGGAAAAAUGUUGAAUUUGAUUAGCGAAGCGUUAAAGAAAAAGACACACAGUCCCGAAAGGAAA-AAAGGAGGCAGCUAAGAAGGCCAAAGAUAAUGAGCCAGUGUUGAGGCGUCG ..............(((..(((..(((((...((.((.........(((.....)))((....))...-....)).)).))))).)))..))).......(((((........))).)). ( -20.70) >DroSec_CAF1 72434 120 + 1 AAAAAGGAAAAAUGUUGAAUUUGAUUAGCGAAGCGUUAAAGAAAAAGACACACAGUUCCGAAAGGAAAAAAAGGAGGCAGCUAAGAAGGCCAAAGAUAAUGAGCCAGUGUUGAGGCGUCG ............((((...(((..((((((...)))))).)))...)))).....((((....))))......((.((..(...(..(((((.......)).)))..)...)..)).)). ( -22.20) >DroSim_CAF1 86063 120 + 1 AAAAAGGAAAAAUGUUGAAUUUGAUUAGCGAAGCGUUAAAGAAAAAGACACACAGUUCCGAAAGGAAAAAAAGGAGGCAGCUAAGAAGGCCAAAGAUAAUGAGCCAGUGUUGAGGCGUCG ............((((...(((..((((((...)))))).)))...)))).....((((....))))......((.((..(...(..(((((.......)).)))..)...)..)).)). ( -22.20) >consensus AAAAAGGAAAAAUGUUGAAUUUGAUUAGCGAAGCGUUAAAGAAAAAGACACACAGUUCCGAAAGGAAAAAAAGGAGGCAGCUAAGAAGGCCAAAGAUAAUGAGCCAGUGUUGAGGCGUCG ............((((...(((..((((((...)))))).)))...)))).....((((....))))......((.((..(...(..(((((.......)).)))..)...)..)).)). (-20.77 = -21.10 + 0.33)


| Location | 17,108,410 – 17,108,529 |
|---|---|
| Length | 119 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 95.33 |
| Mean single sequence MFE | -32.90 |
| Consensus MFE | -27.28 |
| Energy contribution | -27.76 |
| Covariance contribution | 0.48 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.52 |
| Structure conservation index | 0.83 |
| SVM decision value | 0.35 |
| SVM RNA-class probability | 0.698178 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
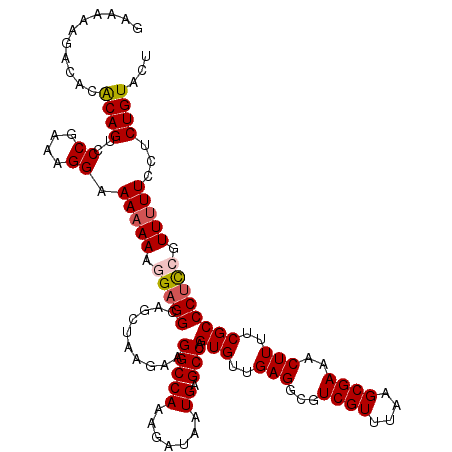
>2L_DroMel_CAF1 17108410 119 + 22407834 GAAAAAGACACACAGUCCCGAAAGGAAA-AAAGGAGGCAGCUAAGAAGGCCAAAGAUAAUGAGCCAGUGUUGAGGCGUCGUUUAAGCGAAACUUUUCGCCCUCCGUUUUUCCUCUGUACU ...........((((.......((((((-((.(((((..........(((((.......)).))).(((..(((...((((....))))..)))..)))))))).))))))))))))... ( -33.41) >DroSec_CAF1 72474 120 + 1 GAAAAAGACACACAGUUCCGAAAGGAAAAAAAGGAGGCAGCUAAGAAGGCCAAAGAUAAUGAGCCAGUGUUGAGGCGUCGUUUAAGCGAAACUUUUCGCCCUCCGUUUUUCCUCUGUACU ((.((((((......((((....)))).....(((((..........(((((.......)).))).(((..(((...((((....))))..)))..))))))))))))))..))...... ( -33.60) >DroSim_CAF1 86103 120 + 1 GAAAAAGACACACAGUUCCGAAAGGAAAAAAAGGAGGCAGCUAAGAAGGCCAAAGAUAAUGAGCCAGUGUUGAGGCGUCGUUUAAGCGAAACUUUUCGCCCUCCGUUUUUCCUCUGUACU ((.((((((......((((....)))).....(((((..........(((((.......)).))).(((..(((...((((....))))..)))..))))))))))))))..))...... ( -33.60) >DroEre_CAF1 73755 118 + 1 GAAAAAGACACGCAGCCCCGAAAGGAAAAAA--GCGGCAACUAAGAAGGCCAAAGAUAAUGAGCCAGUGCUGAGGCGUCGUUUAAGCGAAACUUUUCGCCCUCCGUUUUUCCGCUGUACU ...........(((((.((....))..((((--((((..........(((((.......)).))).(((..(((...((((....))))..)))..)))...))))))))..)))))... ( -32.00) >DroYak_CAF1 74186 118 + 1 GAAAAAGACACACAGUCCCGAAAGGAAAAAA--GAGGCAACUAAGAAGGCCAAAGAUAAUGAGCCAGUGUUGAGGCGUCGUUUAAGCGAAACUUUUCGCCCUUCGUUUUUCCUCUGUGCU ..........(((((.......((((((((.--.((....))..(((((........((((((((........))).)))))...(((((....)))))))))).))))))))))))).. ( -31.91) >consensus GAAAAAGACACACAGUCCCGAAAGGAAAAAAAGGAGGCAGCUAAGAAGGCCAAAGAUAAUGAGCCAGUGUUGAGGCGUCGUUUAAGCGAAACUUUUCGCCCUCCGUUUUUCCUCUGUACU ...........((((..((....)).(((((.(((((..........(((((.......)).))).(((..(((...((((....))))..)))..)))))))).)))))...))))... (-27.28 = -27.76 + 0.48)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:02:06 2006