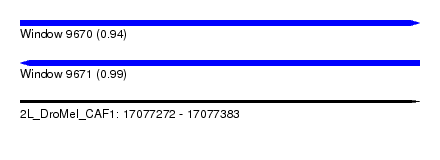
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 17,077,272 – 17,077,383 |
| Length | 111 |
| Max. P | 0.988961 |

| Location | 17,077,272 – 17,077,383 |
|---|---|
| Length | 111 |
| Sequences | 4 |
| Columns | 115 |
| Reading direction | forward |
| Mean pairwise identity | 94.67 |
| Mean single sequence MFE | -30.40 |
| Consensus MFE | -27.64 |
| Energy contribution | -27.26 |
| Covariance contribution | -0.38 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.12 |
| Structure conservation index | 0.91 |
| SVM decision value | 1.29 |
| SVM RNA-class probability | 0.938568 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
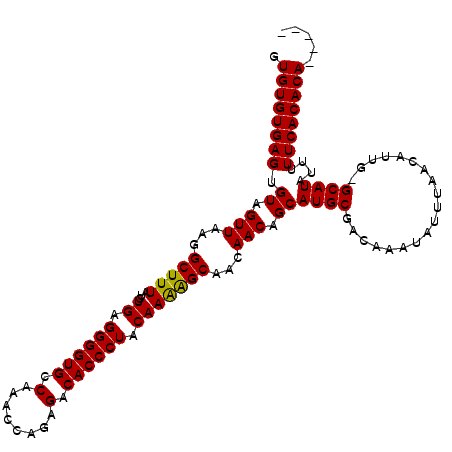
>2L_DroMel_CAF1 17077272 111 + 22407834 GUGUGUGAGUGUAGUUAAGGCUUUAUCUGAGGGGUGCCAAACCAGAGACACCCUACAAAGGCAACAACAGCAUGCGACAAAUAUUUAACAUUG-GCAUAUUUUUCACACACA--- (((((((((.((.(((...(((((...((.((((((.(........).)))))).)))))))...))).))((((..................-))))....))))))))).--- ( -31.77) >DroSec_CAF1 41204 110 + 1 GUGUGUGAGUGUAGUUAAGGCUUUAUCUGAGGGGUGCCAAACCAGAGACACCCUACAAAAGCAACAACAGCAUGCGACAAAUAUUUAACAUUGGGCAUAUUUUUCACACA----- .((((((((.((.(((...(((((...((.((((((.(........).)))))).)))))))...))).))((((..(((..........))).))))....))))))))----- ( -29.40) >DroSim_CAF1 36697 109 + 1 GUGUGUGAGUGUAGUUAAGGCUUUAUCUGAGGGGUGCCAAACCAGAGACACCCUACAAAAGCAACAACAGCAUGCGACAAAUAUUUAACAUUG-GCAUAUUUUUCACACA----- .((((((((.((.(((...(((((...((.((((((.(........).)))))).)))))))...))).))((((..................-))))....))))))))----- ( -27.87) >DroEre_CAF1 40950 114 + 1 GUGUGUGAGUGUAGUUAAGGCUUUAUCUGAGGGGUGCCAAACCAGGGACACCCUGCAAGAGCUACAACAGCAUGCGACAAAUAUUUAACAUUG-GCAUAUUUUUCACACAAACAC .((((((((.((.(((..((((((...((.((((((((.......)).)))))).))))))))..))).))((((..................-))))....))))))))..... ( -32.57) >consensus GUGUGUGAGUGUAGUUAAGGCUUUAUCUGAGGGGUGCCAAACCAGAGACACCCUACAAAAGCAACAACAGCAUGCGACAAAUAUUUAACAUUG_GCAUAUUUUUCACACA_____ .((((((((.((.(((...(((((...((.((((((.(........).)))))).)))))))...))).))((((...................))))....))))))))..... (-27.64 = -27.26 + -0.38)

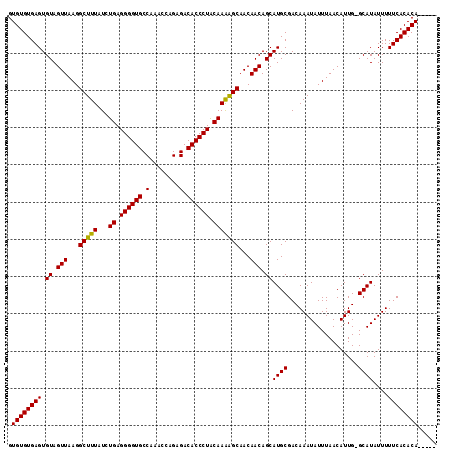
| Location | 17,077,272 – 17,077,383 |
|---|---|
| Length | 111 |
| Sequences | 4 |
| Columns | 115 |
| Reading direction | reverse |
| Mean pairwise identity | 94.67 |
| Mean single sequence MFE | -32.98 |
| Consensus MFE | -30.58 |
| Energy contribution | -31.07 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.76 |
| Structure conservation index | 0.93 |
| SVM decision value | 2.14 |
| SVM RNA-class probability | 0.988961 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
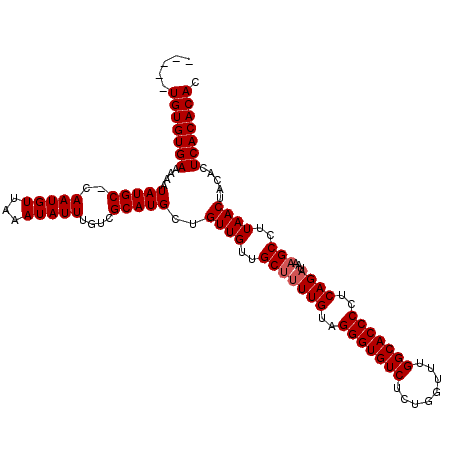
>2L_DroMel_CAF1 17077272 111 - 22407834 ---UGUGUGUGAAAAAUAUGC-CAAUGUUAAAUAUUUGUCGCAUGCUGUUGUUGCCUUUGUAGGGUGUCUCUGGUUUGGCACCCCUCAGAUAAAGCCUUAACUACACUCACACAC ---.((((((((....(((((-(((..........)))..)))))..((((..((.((((..(((((((........)))))))..))))....))..)))).....)))))))) ( -33.00) >DroSec_CAF1 41204 110 - 1 -----UGUGUGAAAAAUAUGCCCAAUGUUAAAUAUUUGUCGCAUGCUGUUGUUGCUUUUGUAGGGUGUCUCUGGUUUGGCACCCCUCAGAUAAAGCCUUAACUACACUCACACAC -----(((((((....(((((.(((..........)))..)))))..((((..(((((((..(((((((........)))))))..))))...)))..)))).....))))))). ( -33.60) >DroSim_CAF1 36697 109 - 1 -----UGUGUGAAAAAUAUGC-CAAUGUUAAAUAUUUGUCGCAUGCUGUUGUUGCUUUUGUAGGGUGUCUCUGGUUUGGCACCCCUCAGAUAAAGCCUUAACUACACUCACACAC -----(((((((....(((((-(((..........)))..)))))..((((..(((((((..(((((((........)))))))..))))...)))..)))).....))))))). ( -32.70) >DroEre_CAF1 40950 114 - 1 GUGUUUGUGUGAAAAAUAUGC-CAAUGUUAAAUAUUUGUCGCAUGCUGUUGUAGCUCUUGCAGGGUGUCCCUGGUUUGGCACCCCUCAGAUAAAGCCUUAACUACACUCACACAC ((((..(((((......((..-..))(((((...(((((((((.((((...))))...))).(((((((........)))))))....))))))...))))))))))..)))).. ( -32.60) >consensus _____UGUGUGAAAAAUAUGC_CAAUGUUAAAUAUUUGUCGCAUGCUGUUGUUGCUUUUGUAGGGUGUCUCUGGUUUGGCACCCCUCAGAUAAAGCCUUAACUACACUCACACAC .....(((((((....(((((..(((((...)))))....)))))..((((..(((((((..(((((((........)))))))..))))...)))..)))).....))))))). (-30.58 = -31.07 + 0.50)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:00:50 2006